Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 53(1); 2019 > Article
-
Review
Artificial Intelligence in Pathology -
Hye Yoon Chang
 , Chan Kwon Jung1
, Chan Kwon Jung1 , Junwoo Isaac Woo
, Junwoo Isaac Woo , Sanghun Lee
, Sanghun Lee , Joonyoung Cho
, Joonyoung Cho , Sun Woo Kim
, Sun Woo Kim , Tae-Yeong Kwak
, Tae-Yeong Kwak
-
Journal of Pathology and Translational Medicine 2019;53(1):1-12.
DOI: https://doi.org/10.4132/jptm.2018.12.16
Published online: December 28, 2018
Deep Bio Inc., Seoul, Korea
1Department of Hospital Pathology, College of Medicine, The Catholic University of Korea, Seoul, Korea
- Corresponding Author Tae-Yeong Kwak, PhD Deep Bio Inc., 1201 Hanwha Bizmetro, 242 Digital-ro, Guro-gu, Seoul 08394, Korea Tel: +82-70-7703-4746 Fax: +82-2-2621-2223 E-mail: tykwak@deepbio.co.kr
© 2019 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Figure & Data
References
Citations

- Interpretable Machine Learning Approaches for Identification of Acute Aortic Dissection in Chest Pain Patients
Shuangshuang Li, Kaiwen Zhao, Wen Li, Qingsheng Lu, Jian Zhou, Jia He
Annals of Vascular Surgery.2026; 122: 895. CrossRef - An automatic, rapid and accurate method for the annotation of tumor components on whole slide images
Hong Tang, Xiaodong Wang, Xiaolin Zhang, Xiaojun Wu, Xinyue Tang, Yaqiong Ma, Ying Chen, Guanzhen Yu
Journal of Histotechnology.2026; 49(1): 13. CrossRef - Pengaruh Personalisasi Artificial Intelligence dan Pengalaman Pelanggan terhadap Loyalitas Pelanggan
Mohammad Bastian Dwiki Rahmat, Darianto Darianto, Amanatul Khoiriyah
Jurnal Simki Economic.2026; 9(1): 168. CrossRef - Adaptive diagnostic reasoning framework for pathology with multimodal large language models
Yunqi Hong, Kuei-Chun Kao, Liam Edwards, Nein-Tzu Liu, Chung-Yen Huang, Alex Oliveira-Kowaleski, Cho-Jui Hsieh, Neil Y. C. Lin
Communications Medicine.2026;[Epub] CrossRef - Development and validation of a CNN model for grading oral epithelial dysplasia
Chetan Belaldavar, Punnya V. Angadi, Uma Mudenagudi, Ujwala Patil, Ramesh Ashok Tabib
Revista Española de Patología.2026; 59(2): 100869. CrossRef - Review of the Changing Roles of Clinical Laboratory Scientists and Strategies for Curricular Innovation in the Era of Artificial Intelligence
Hee Sung KIM
Korean Journal of Clinical Laboratory Science.2026; 58(1): 1. CrossRef - Exploring the status of artificial intelligence for healthcare research in Africa: a bibliometric and thematic analysis
Tabu S. Kondo, Salim A. Diwani, Ally S. Nyamawe, Mohamed M. Mjahidi
AI and Ethics.2025; 5(1): 117. CrossRef - Prioritize Threat Alerts Based on False Positives Qualifiers Provided by Multiple AI Models Using Evolutionary Computation and Reinforcement Learning
Anup Sharma, V. G. Kiran Kumar, Asmita Poojari
Journal of The Institution of Engineers (India): Series B.2025; 106(4): 1305. CrossRef - Artificial intelligence versus human analysis: Interpreting data in elderly fat reduction study
Piotr Sporek, Mariusz Konieczny
Advances in Integrative Medicine.2025; 12(1): 13. CrossRef - Artificial intelligence in healthcare applications targeting cancer diagnosis—part I: data structure, preprocessing and data organization
Anna Luíza Damaceno Araújo, Marcelo Sperandio, Giovanna Calabrese, Sarah S. Faria, Diego Armando Cardona Cardenas, Manoela Domingues Martins, Cristina Saldivia-Siracusa, Daniela Giraldo-Roldán, Caique Mariano Pedroso, Pablo Agustin Vargas, Marcio Ajudarte
Oral Surgery, Oral Medicine, Oral Pathology and Oral Radiology.2025; 140(1): 79. CrossRef - Artificial intelligence–driven digital pathology in urological cancers: current trends and future directions
Inyoung Paik, Geongyu Lee, Joonho Lee, Tae-Yeong Kwak, Hong Koo Ha
Prostate International.2025; 13(4): 181. CrossRef - Optimizing deep learning for accurate blood cell classification: A study on stain normalization and fine-tuning techniques
Mohammed Tareq Mutar, Jaffar Nouri Alalsaidissa, Mustafa Majid Hameed, Ali Almothaffar
Iraqi Journal of Hematology.2025; 14(1): 60. CrossRef - Structural imbalance of medical resources amid population mobility and digital empowerment: a study of national and port-developed provinces in China
Haiwei Fu, Junjie Lu
Frontiers in Public Health.2025;[Epub] CrossRef - Exploring the evolution of artificial intelligence in pathology: a bibliometric and network analysis
Burcu Sanal Yılmaz
Journal of Medicine and Palliative Care.2025; 6(3): 224. CrossRef - ШТУЧНИЙ ІНТЕЛЕКТ У СУЧАСНІЙ СТОМАТОЛОГІЇ
О. І. Бульбук, О. В. Бульбук, О. В. Шутак, Ю. І. Сухоребський
Art of Medicine.2025; : 101. CrossRef - Natural language processing in veterinary pathology: A review
Lev Stimmer, Raoul V. Kuiper, Laura Polledo, Lorenzo Ressel, Josep M. Monné Rodriguez, Inês B. Veiga, Jonathan Williams, Vanessa Herder
Veterinary Pathology.2025; 62(6): 829. CrossRef - Impact of Magnification, Image Type, and Number on Convolutional Neural Network Performance in Differentiating Canine Large Cell Lymphoma From Non‐Lymphoma via Lymph Node Cytology
Christina Pacholec, Hehuang Xie, Julianne Curnin, Amy Lin, Kurt Zimmerman
Veterinary Clinical Pathology.2025;[Epub] CrossRef - Pathology image-based predictive model for individual survival time of early-stage lung adenocarcinoma patients
Vi Thi-Tuong Vo, Hyung-Jeong Yang, Taebum Lee, Soo-Hyung Kim
Scientific Reports.2025;[Epub] CrossRef - Exploring Artificial Intelligence's Potential to Enhance Conventional Anticancer Drug Development
Sorin‐Ștefan Bobolea, Miruna‐Ioana Hinoveanu, Andreea Dimitriu, Miruna‐Andrada Brașoveanu, Cristian‐Nicolae Iliescu, Cristina‐Elena Dinu‐Pîrvu, Mihaela Violeta Ghica, Valentina Anuța, Lăcrămioara Popa, Răzvan Mihai Prisada
Drug Development Research.2025;[Epub] CrossRef - Predicability of PD-L1 expression in cancer cells based solely on H&E-stained sections
Gavino Faa, Matteo Fraschini, Pina Ziranu, Andrea Pretta, Giuseppe Porcu, Luca Saba, Mario Scartozzi, Nazar Shokun, Massimo Rugge
Journal of Pathology Informatics.2025; 19: 100524. CrossRef - Artificial Intelligence in Medicine
Umur Karan, Osman Elbek
Thoracic Research and Practice.2025;[Epub] CrossRef - Whole Slide Imaging Technology and Its Applications: Current and Emerging Perspectives
Ekta Jain, Ankush Patel, Anil V. Parwani, Saba Shafi, Zoya Brar, Shivani Sharma, Sambit K. Mohanty
International Journal of Surgical Pathology.2024; 32(3): 433. CrossRef - ChatGPT as an aid for pathological diagnosis of cancer
Shaivy Malik, Sufian Zaheer
Pathology - Research and Practice.2024; 253: 154989. CrossRef - Computational pathology: A survey review and the way forward
Mahdi S. Hosseini, Babak Ehteshami Bejnordi, Vincent Quoc-Huy Trinh, Lyndon Chan, Danial Hasan, Xingwen Li, Stephen Yang, Taehyo Kim, Haochen Zhang, Theodore Wu, Kajanan Chinniah, Sina Maghsoudlou, Ryan Zhang, Jiadai Zhu, Samir Khaki, Andrei Buin, Fatemeh
Journal of Pathology Informatics.2024; 15: 100357. CrossRef - Applications of artificial intelligence in the field of oral and maxillofacial pathology: a systematic review and meta-analysis
Nishath Sayed Abdul, Ganiga Channaiah Shivakumar, Sunila Bukanakere Sangappa, Marco Di Blasio, Salvatore Crimi, Marco Cicciù, Giuseppe Minervini
BMC Oral Health.2024;[Epub] CrossRef - Machine-learning models are superior to severity scoring systems for the prediction of the mortality of critically ill patients in a tertiary medical center
Ruey-Hsing Chou, Benny Wei-Yun Hsu, Chun-Lin Yu, Tai-Yuan Chen, Shuo-Ming Ou, Kuo-Hua Lee, Vincent S. Tseng, Po-Hsun Huang, Der-Cherng Tarng
Journal of the Chinese Medical Association.2024; 87(4): 369. CrossRef - The Evaluation of Artificial Intelligence Technology for the Differentiation of Fresh Human Blood Cells From Other Species Blood in the Investigation of Crime Scenes
Syed Sajid Hussain Shah, Ekramy Elmorsy, Rashad Qasem Ali Othman, Asmara Syed, Syed Umar Armaghan, Syed Usama Khalid Bokhari, Mahmoud E Elmorsy, Abdulhakim Bawadekji
Cureus.2024;[Epub] CrossRef - A Comparison of Diagnostic and Immunohistochemical Workup and Literature Review Capabilities of Online Artificial Intelligence Assistance Models in Pathology
Johnika Dougan, Netra Patel, Svetoslav Bardarov
Cureus.2024;[Epub] CrossRef - ChatENT: Augmented Large Language Model for Expert Knowledge Retrieval in Otolaryngology–Head and Neck Surgery
Cai Long, Deepak Subburam, Kayle Lowe, André dos Santos, Jessica Zhang, Sang Hwang, Neil Saduka, Yoav Horev, Tao Su, David W.J. Côté, Erin D. Wright
Otolaryngology–Head and Neck Surgery.2024; 171(4): 1042. CrossRef - Artificial intelligence in forensic medicine and related sciences – selected issues = Sztuczna inteligencja w medycynie sądowej i naukach pokrewnych – wybrane zagadnienia
Michał Szeremeta, Julia Janica, Anna Niemcunowicz-Janica
Archives of Forensic Medicine and Criminology.2024; 74(1): 64. CrossRef - Unveiling the landscape of pathomics in personalized immunotherapy for lung cancer: a bibliometric analysis
Lei Yuan, Zhiming Shen, Yibo Shan, Jianwei Zhu, Qi Wang, Yi Lu, Hongcan Shi
Frontiers in Oncology.2024;[Epub] CrossRef - PathEX: Make good choice for whole slide image extraction
Xinda Yang, Ranze Zhang, Yuan Yang, Yu Zhang, Kai Chen, Alberto Marchisio
PLOS ONE.2024; 19(8): e0304702. CrossRef - Automatic point detection on cephalograms using convolutional neural networks: A two-step method
Miki HORI, Makoto JINCHO, Tadasuke HORI, Hironao SEKINE, Akiko KATO, Ken MIYAZAWA, Tatsushi KAWAI
Dental Materials Journal.2024; 43(5): 701. CrossRef - The use of generative artificial intelligence (AI) in teaching and assessment of postgraduate students in pathology and microbiology
Dipmala Das, Asitava Deb Roy, Subhayan Dasgupta, Rohon Das Roy
Indian Journal of Microbiology Research.2024; 11(3): 140. CrossRef - Inteligencia artificial: desafíos éticos y futuros
Jhadson Silva Leonel, Camila Ferreira Silva Leonel, Jonas Byk, Silvania da Conceição Furtado
Revista Bioética.2024;[Epub] CrossRef - Artificial intelligence: ethical and future challenges
Jhadson Silva Leonel, Camila Ferreira Silva Leonel, Jonas Byk, Silvania da Conceição Furtado
Revista Bioética.2024;[Epub] CrossRef - Inteligência artificial: desafios éticos e futuros
Jhadson Silva Leonel, Camila Ferreira Silva Leonel, Jonas Byk, Silvania da Conceição Furtado
Revista Bioética.2024;[Epub] CrossRef - The Constrained-Disorder Principle Assists in Overcoming Significant Challenges in Digital Health: Moving from “Nice to Have” to Mandatory Systems
Noa Hurvitz, Yaron Ilan
Clinics and Practice.2023; 13(4): 994. CrossRef - Building a nonclinical pathology laboratory of the future for pharmaceutical research excellence
D.G. Rudmann, L. Bertrand, A. Zuraw, J. Deiters, M. Staup, Y. Rivenson, J. Kuklyte
Drug Discovery Today.2023; 28(10): 103747. CrossRef - Automated image analysis of keratin 7 staining can predict disease outcome in primary sclerosing cholangitis
Nelli Sjöblom, Sonja Boyd, Anniina Manninen, Sami Blom, Anna Knuuttila, Martti Färkkilä, Johanna Arola
Hepatology Research.2023; 53(4): 322. CrossRef - Application of convolutional neural network for analyzing hepatic fibrosis in mice
Hyun-Ji Kim, Eun Bok Baek, Ji-Hee Hwang, Minyoung Lim, Won Hoon Jung, Myung Ae Bae, Hwa-Young Son, Jae-Woo Cho
Journal of Toxicologic Pathology.2023; 36(1): 21. CrossRef - Machine Learning Techniques for Prognosis Estimation and Knowledge Discovery From Lab Test Results With Application to the COVID-19 Emergency
Alfonso Emilio Gerevini, Roberto Maroldi, Matteo Olivato, Luca Putelli, Ivan Serina
IEEE Access.2023; 11: 83905. CrossRef - Artificial intelligence in dentistry—A review
Hao Ding, Jiamin Wu, Wuyuan Zhao, Jukka P. Matinlinna, Michael F. Burrow, James K. H. Tsoi
Frontiers in Dental Medicine.2023;[Epub] CrossRef - Dental Age Estimation Using the Demirjian Method: Statistical Analysis Using Neural Networks
Byung-Yoon Roh, Jong-Seok Lee, Sang-Beom Lim, Hye-Won Ryu, Su-Jeong Jeon, Ju-Heon Lee, Yo-Seob Seo, Ji-Won Ryu, Jong-Mo Ahn
Korean Journal of Legal Medicine.2023; 47(1): 1. CrossRef - The use of artificial intelligence in health care. Problems of identification of patients' conditions in the processes of detailing the diagnosis
Mintser O
Artificial Intelligence.2023; 28(AI.2023.28): 8. CrossRef - The Effectiveness of Data Augmentation for Mature White Blood Cell Image Classification in Deep Learning — Selection of an Optimal Technique for Hematological Morphology Recognition —
Hiroyuki NOZAKA, Kosuke KAMATA, Kazufumi YAMAGATA
IEICE Transactions on Information and Systems.2023; E106.D(5): 707. CrossRef - Rectal Cancer Stages T2 and T3 Identification Based on Asymptotic Hybrid Feature Maps
Shujing Sun, Jiale Wu, Jian Yao, Yang Cheng, Xin Zhang, Zhihua Lu, Pengjiang Qian
Computer Modeling in Engineering & Sciences.2023; 137(1): 923. CrossRef - How to use AI in pathology
Peter Schüffler, Katja Steiger, Wilko Weichert
Genes, Chromosomes and Cancer.2023; 62(9): 564. CrossRef - Cutting-Edge Technologies for Digital Therapeutics: A Review and Architecture Proposals for Future Directions
Joo Hun Yoo, Harim Jeong, Tai-Myoung Chung
Applied Sciences.2023; 13(12): 6929. CrossRef - A convolutional neural network STIFMap reveals associations between stromal stiffness and EMT in breast cancer
Connor Stashko, Mary-Kate Hayward, Jason J. Northey, Neil Pearson, Alastair J. Ironside, Johnathon N. Lakins, Roger Oria, Marie-Anne Goyette, Lakyn Mayo, Hege G. Russnes, E. Shelley Hwang, Matthew L. Kutys, Kornelia Polyak, Valerie M. Weaver
Nature Communications.2023;[Epub] CrossRef - Artificial Intelligence-Based PTEN Loss Assessment as an Early Predictor of Prostate Cancer Metastasis After Surgery: A Multicenter Retrospective Study
Palak Patel, Stephanie Harmon, Rachael Iseman, Olga Ludkowski, Heidi Auman, Sarah Hawley, Lisa F. Newcomb, Daniel W. Lin, Peter S. Nelson, Ziding Feng, Hilary D. Boyer, Maria S. Tretiakova, Larry D. True, Funda Vakar-Lopez, Peter R. Carroll, Matthew R. Co
Modern Pathology.2023; 36(10): 100241. CrossRef - Minimum resolution requirements of digital pathology images for accurate classification
Lydia Neary-Zajiczek, Linas Beresna, Benjamin Razavi, Vijay Pawar, Michael Shaw, Danail Stoyanov
Medical Image Analysis.2023; 89: 102891. CrossRef - Artificial Intelligence in the Pathology of Gastric Cancer
Sangjoon Choi, Seokhwi Kim
Journal of Gastric Cancer.2023; 23(3): 410. CrossRef - Endoscopic Ultrasound-Based Artificial Intelligence Diagnosis of Pancreatic Cystic Neoplasms
Jin-Seok Park, Seok Jeong
The Korean Journal of Pancreas and Biliary Tract.2023; 28(3): 53. CrossRef - Framework for Classifying Explainable Artificial Intelligence (XAI) Algorithms in Clinical Medicine
Thomas Gniadek, Jason Kang, Talent Theparee, Jacob Krive
Online Journal of Public Health Informatics.2023; 15: e50934. CrossRef - A Literature Review of the Future of Oral Medicine and Radiology, Oral Pathology, and Oral Surgery in the Hands of Technology
Ishita Singhal, Geetpriya Kaur, Dirk Neefs, Aparna Pathak
Cureus.2023;[Epub] CrossRef - AI-Powered Biomolecular-Specific and Label-Free Multispectral Imaging Rapidly Detects Malignant Neoplasm in Surgically Excised Breast Tissue Specimens
Rishikesh Pandey, David Fournier, Gary Root, Machele Riccio, Aditya Shirvalkar, Gianfranco Zamora, Noel Daigneault, Michael Sapack, Minghao Zhong, Malini Harigopal
Archives of Pathology & Laboratory Medicine.2023; 147(11): 1298. CrossRef - Artificial intelligence for patient scheduling in the real-world health care setting: A metanarrative review
Dacre R.T. Knight, Christopher A. Aakre, Christopher V. Anstine, Bala Munipalli, Parisa Biazar, Ghada Mitri, Jose Raul Valery, Tara Brigham, Shehzad K. Niazi, Adam I. Perlman, John D. Halamka, Abd Moain Abu Dabrh
Health Policy and Technology.2023; 12(4): 100824. CrossRef - Towards Autonomous Healthcare: Integrating Artificial Intelligence (AI) for Personalized Medicine and Disease Prediction
Nitin Rane, Saurabh Choudhary, Jayesh Rane
SSRN Electronic Journal.2023;[Epub] CrossRef - Medical imaging and multimodal artificial intelligence models for streamlining and enhancing cancer care: opportunities and challenges
Kevin Pierre, Manas Gupta, Abheek Raviprasad, Seyedeh Mehrsa Sadat Razavi, Anjali Patel, Keith Peters, Bruno Hochhegger, Anthony Mancuso, Reza Forghani
Expert Review of Anticancer Therapy.2023; 23(12): 1265. CrossRef - Automated differential diagnostics of respiratory diseases using an electronic stethoscope
Diana Arhypenko, Denis Panaskin, Dmytro Babko
Polish Journal of Medical Physics and Engineering.2023; 29(4): 208. CrossRef - Application of machine learning in identification of pathogenic microbes
Lakshmi Venkata S Kutikuppala, Kanishk K Adhit, Reewen George D Silva
Digital Medicine.2023;[Epub] CrossRef - The Beginning of a New Era
C Nandini, Shaik Basha, Aarchi Agarawal, R Parikh Neelampari, Krishna P Miyapuram, R Jadeja Nileshwariba
Advances in Human Biology.2023; 13(1): 4. CrossRef - Artificial Intelligence in Respiratory Medicine
K Kalaiyarasan, R Sridhar
Journal of Association of Pulmonologist of Tamil Nadu.2023; 6(2): 53. CrossRef - Automated abstraction of myocardial perfusion imaging reports using natural language processing
Parija Sharedalal, Ajay Singh, Neal Shah, Diwakar Jain
Journal of Nuclear Cardiology.2022; 29(3): 1188. CrossRef - Polyploid giant cancer cell characterization: New frontiers in predicting response to chemotherapy in breast cancer
Geetanjali Saini, Shriya Joshi, Chakravarthy Garlapati, Hongxiao Li, Jun Kong, Jayashree Krishnamurthy, Michelle D. Reid, Ritu Aneja
Seminars in Cancer Biology.2022; 81: 220. CrossRef - A Comprehensive Review of Markov Random Field and Conditional Random Field Approaches in Pathology Image Analysis
Yixin Li, Chen Li, Xiaoyan Li, Kai Wang, Md Mamunur Rahaman, Changhao Sun, Hao Chen, Xinran Wu, Hong Zhang, Qian Wang
Archives of Computational Methods in Engineering.2022; 29(1): 609. CrossRef - Artificial intelligence in oncology: From bench to clinic
Jamal Elkhader, Olivier Elemento
Seminars in Cancer Biology.2022; 84: 113. CrossRef - Yeast‐like organisms phagocytosed by circulating neutrophils: Evidence of disseminated histoplasmosis
Yue Zhao, Jenna McCracken, Endi Wang
International Journal of Laboratory Hematology.2022; 44(1): 51. CrossRef - Whole-slide imaging, tissue image analysis, and artificial intelligence in veterinary pathology: An updated introduction and review
Aleksandra Zuraw, Famke Aeffner
Veterinary Pathology.2022; 59(1): 6. CrossRef - A comprehensive review of computer-aided whole-slide image analysis: from datasets to feature extraction, segmentation, classification and detection approaches
Xintong Li, Chen Li, Md Mamunur Rahaman, Hongzan Sun, Xiaoqi Li, Jian Wu, Yudong Yao, Marcin Grzegorzek
Artificial Intelligence Review.2022; 55(6): 4809. CrossRef - Liquid Biopsy and Artificial Intelligence as Tools to Detect Signatures of Colorectal Malignancies: A Modern Approach in Patient’s Stratification
Octav Ginghina, Ariana Hudita, Marius Zamfir, Andrada Spanu, Mara Mardare, Irina Bondoc, Laura Buburuzan, Sergiu Emil Georgescu, Marieta Costache, Carolina Negrei, Cornelia Nitipir, Bianca Galateanu
Frontiers in Oncology.2022;[Epub] CrossRef - Automated bone marrow cytology using deep learning to generate a histogram of cell types
Rohollah Moosavi Tayebi, Youqing Mu, Taher Dehkharghanian, Catherine Ross, Monalisa Sur, Ronan Foley, Hamid R. Tizhoosh, Clinton J. V. Campbell
Communications Medicine.2022;[Epub] CrossRef - Risultati di esami di laboratorio per intelligenza artificiale e "machine learning"
Marco PRADELLA
La Rivista Italiana della Medicina di Laboratorio.2022;[Epub] CrossRef - The Deception of Certainty: how Non-Interpretable Machine Learning Outcomes Challenge the Epistemic Authority of Physicians. A deliberative-relational Approach
Florian Funer
Medicine, Health Care and Philosophy.2022; 25(2): 167. CrossRef - Deep discriminative learning model with calibrated attention map for the automated diagnosis of diffuse large B-cell lymphoma
Sautami Basu, Ravinder Agarwal, Vishal Srivastava
Biomedical Signal Processing and Control.2022; 76: 103728. CrossRef - Question and Answer Techniques for Financial Audits in Universities Based on Deep Learning
Qiang Li, Hangjun Che
Mathematical Problems in Engineering.2022; 2022: 1. CrossRef - Noninvasive Screening Tool for Hyperkalemia Using a Single-Lead Electrocardiogram and Deep Learning: Development and Usability Study
Erdenebayar Urtnasan, Jung Hun Lee, Byungjin Moon, Hee Young Lee, Kyuhee Lee, Hyun Youk
JMIR Medical Informatics.2022; 10(6): e34724. CrossRef - Impact of artificial intelligence on pathologists’ decisions: an experiment
Julien Meyer, April Khademi, Bernard Têtu, Wencui Han, Pria Nippak, David Remisch
Journal of the American Medical Informatics Association.2022; 29(10): 1688. CrossRef - Rapid Screening Using Pathomorphologic Interpretation to Detect BRAFV600E Mutation and Microsatellite Instability in Colorectal Cancer
Satoshi Fujii, Daisuke Kotani, Masahiro Hattori, Masato Nishihara, Toshihide Shikanai, Junji Hashimoto, Yuki Hama, Takuya Nishino, Mizuto Suzuki, Ayatoshi Yoshidumi, Makoto Ueno, Yoshito Komatsu, Toshiki Masuishi, Hiroki Hara, Taito Esaki, Yoshiaki Nakamu
Clinical Cancer Research.2022; 28(12): 2623. CrossRef - Using Deep Learning to Predict Final HER2 Status in Invasive Breast Cancers That are Equivocal (2+) by Immunohistochemistry
Sean A. Rasmussen, Valerie J. Taylor, Alexi P. Surette, Penny J. Barnes, Gillian C. Bethune
Applied Immunohistochemistry & Molecular Morphology.2022; 30(10): 668. CrossRef - Deep Neural Network for the Prediction of KRAS Genotype in Rectal Cancer
Waleed M Ghareeb, Eman Draz, Khaled Madbouly, Ahmed H Hussein, Mohammed Faisal, Wagdi Elkashef, Mona Hany Emile, Marcus Edelhamre, Seon Hahn Kim, Sameh Hany Emile
Journal of the American College of Surgeons.2022; 235(3): 482. CrossRef - Next Generation Digital Pathology: Emerging Trends and Measurement Challenges for Molecular Pathology
Alex Dexter, Dimitrios Tsikritsis, Natalie A. Belsey, Spencer A. Thomas, Jenny Venton, Josephine Bunch, Marina Romanchikova
Journal of Molecular Pathology.2022; 3(3): 168. CrossRef - Animation Design of Multisensor Data Fusion Based on Optimized AVOD Algorithm
Li Ding, Guobing Wei, Kai Zhang, Gengxin Sun
Journal of Sensors.2022; 2022: 1. CrossRef - Study on Machine Translation Teaching Model Based on Translation Parallel Corpus and Exploitation for Multimedia Asian Information Processing
Yan Gong
ACM Transactions on Asian and Low-Resource Language Information Processing.2022;[Epub] CrossRef - Analysis and Estimation of Pathological Data and Findings with Deep Learning Methods
Ahmet Anıl ŞAKIR, Ali Hakan IŞIK, Özlem ÖZMEN, Volkan İPEK
Veterinary Journal of Mehmet Akif Ersoy University.2022; 7(3): 175. CrossRef - Artificial Intelligence in Pathology: Friend or Enemy?
Selim Sevim, Ezgi Dicle Serbes, Murat Bahadır, Mustafa Said Kartal, Serpil Dizbay Sak
Journal of Ankara University Faculty of Medicine.2022; 75(1): 13. CrossRef - Assessment of knowledge, attitude, and practice regarding artificial intelligence in histopathology: A cross-sectional study among oral pathologists in India
M. Indu, Vidya Gurram Shankar, Latha Mary Cherian, Revathi Krishna, Sabu Paul, Pradeesh Sathyan
Saudi Journal of Oral Sciences.2022; 9(3): 157. CrossRef - Evaluation Challenges in the Validation of B7-H3 as Oral Tongue Cancer Prognosticator
Meri Sieviläinen, Anna Maria Wirsing, Aini Hyytiäinen, Rabeia Almahmoudi, Priscila Rodrigues, Inger-Heidi Bjerkli, Pirjo Åström, Sanna Toppila-Salmi, Timo Paavonen, Ricardo D. Coletta, Elin Hadler-Olsen, Tuula Salo, Ahmed Al-Samadi
Head and Neck Pathology.2021; 15(2): 469. CrossRef - Amsterdam International Consensus Meeting: tumor response scoring in the pathology assessment of resected pancreatic cancer after neoadjuvant therapy
Boris V. Janssen, Faik Tutucu, Stijn van Roessel, Volkan Adsay, Olca Basturk, Fiona Campbell, Claudio Doglioni, Irene Esposito, Roger Feakins, Noriyoshi Fukushima, Anthony J. Gill, Ralph H. Hruban, Jeffrey Kaplan, Bas Groot Koerkamp, Seung-Mo Hong, Alyssa
Modern Pathology.2021; 34(1): 4. CrossRef - Fabrication of ultra-thin 2D covalent organic framework nanosheets and their application in functional electronic devices
Weikang Wang, Weiwei Zhao, Haotian Xu, Shujuan Liu, Wei Huang, Qiang Zhao
Coordination Chemistry Reviews.2021; 429: 213616. CrossRef - Generalizability of Deep Learning System for the Pathologic Diagnosis of Various Cancers
Hyun-Jong Jang, In Hye Song, Sung Hak Lee
Applied Sciences.2021; 11(2): 808. CrossRef - Integrated digital pathology at scale: A solution for clinical diagnostics and cancer research at a large academic medical center
Peter J Schüffler, Luke Geneslaw, D Vijay K Yarlagadda, Matthew G Hanna, Jennifer Samboy, Evangelos Stamelos, Chad Vanderbilt, John Philip, Marc-Henri Jean, Lorraine Corsale, Allyne Manzo, Neeraj H G Paramasivam, John S Ziegler, Jianjiong Gao, Juan C Peri
Journal of the American Medical Informatics Association.2021; 28(9): 1874. CrossRef - Translational Applications of Artificial Intelligence and Machine Learning for Diagnostic Pathology in Lymphoid Neoplasms: A Comprehensive and Evolutive Analysis
Julia Moran-Sanchez, Antonio Santisteban-Espejo, Miguel Angel Martin-Piedra, Jose Perez-Requena, Marcial Garcia-Rojo
Biomolecules.2021; 11(6): 793. CrossRef - Development and operation of a digital platform for sharing pathology image data
Yunsook Kang, Yoo Jung Kim, Seongkeun Park, Gun Ro, Choyeon Hong, Hyungjoon Jang, Sungduk Cho, Won Jae Hong, Dong Un Kang, Jonghoon Chun, Kyoungbun Lee, Gyeong Hoon Kang, Kyoung Chul Moon, Gheeyoung Choe, Kyu Sang Lee, Jeong Hwan Park, Won-Ki Jeong, Se Yo
BMC Medical Informatics and Decision Making.2021;[Epub] CrossRef - Sliding window based deep ensemble system for breast cancer classification
Amin Alqudah, Ali Mohammad Alqudah
Journal of Medical Engineering & Technology.2021; 45(4): 313. CrossRef - Artificial intelligence and computational pathology
Miao Cui, David Y. Zhang
Laboratory Investigation.2021; 101(4): 412. CrossRef - Effects of Image Quantity and Image Source Variation on Machine Learning Histology Differential Diagnosis Models
Elham Vali-Betts, Kevin J. Krause, Alanna Dubrovsky, Kristin Olson, John Paul Graff, Anupam Mitra, Ananya Datta-Mitra, Kenneth Beck, Aristotelis Tsirigos, Cynthia Loomis, Antonio Galvao Neto, Esther Adler, Hooman H. Rashidi
Journal of Pathology Informatics.2021; 12(1): 5. CrossRef - Feasibility of deep learning‐based fully automated classification of microsatellite instability in tissue slides of colorectal cancer
Sung Hak Lee, In Hye Song, Hyun‐Jong Jang
International Journal of Cancer.2021; 149(3): 728. CrossRef - Artificial intelligence in healthcare
Yamini D Shah, Shailvi M Soni, Manish P Patel
Indian Journal of Pharmacy and Pharmacology.2021; 8(2): 102. CrossRef - Proof of Concept for a Deep Learning Algorithm for Identification and Quantification of Key Microscopic Features in the Murine Model of DSS-Induced Colitis
Agathe Bédard, Thomas Westerling-Bui, Aleksandra Zuraw
Toxicologic Pathology.2021; 49(4): 897. CrossRef - An empirical analysis of machine learning frameworks for digital pathology in medical science
S.K.B. Sangeetha, R Dhaya, Dhruv T Shah, R Dharanidharan, K. Praneeth Sai Reddy
Journal of Physics: Conference Series.2021; 1767(1): 012031. CrossRef - Application of Single-Cell Approaches to Study Myeloproliferative Neoplasm Biology
Daniel Royston, Adam J. Mead, Bethan Psaila
Hematology/Oncology Clinics of North America.2021; 35(2): 279. CrossRef - Idiosyncratic Drug-Induced Liver Injury (DILI) and Herb-Induced Liver Injury (HILI): Diagnostic Algorithm Based on the Quantitative Roussel Uclaf Causality Assessment Method (RUCAM)
Rolf Teschke, Gaby Danan
Diagnostics.2021; 11(3): 458. CrossRef - Searching Images for Consensus
Hamid R. Tizhoosh, Phedias Diamandis, Clinton J.V. Campbell, Amir Safarpoor, Shivam Kalra, Danial Maleki, Abtin Riasatian, Morteza Babaie
The American Journal of Pathology.2021; 191(10): 1702. CrossRef - Automated Classification and Segmentation in Colorectal Images Based on Self‐Paced Transfer Network
Yao Yao, Shuiping Gou, Ru Tian, Xiangrong Zhang, Shuixiang He, Zhiguo Zhou
BioMed Research International.2021;[Epub] CrossRef - Artificial intelligence and sleep: Advancing sleep medicine
Nathaniel F. Watson, Christopher R. Fernandez
Sleep Medicine Reviews.2021; 59: 101512. CrossRef - Prospective Of Artificial Intelligence: Emerging Trends In Modern Biosciences Research
Pradeep Kumar, Ajit Kumar Singh Yadav, Abhishek Singh
IOP Conference Series: Materials Science and Engineering.2021; 1020(1): 012008. CrossRef - Use and Control of Artificial Intelligence in Patients Across the Medical Workflow: Single-Center Questionnaire Study of Patient Perspectives
Simon Lennartz, Thomas Dratsch, David Zopfs, Thorsten Persigehl, David Maintz, Nils Große Hokamp, Daniel Pinto dos Santos
Journal of Medical Internet Research.2021; 23(2): e24221. CrossRef - HEAL: an automated deep learning framework for cancer histopathology image analysis
Yanan Wang, Nicolas Coudray, Yun Zhao, Fuyi Li, Changyuan Hu, Yao-Zhong Zhang, Seiya Imoto, Aristotelis Tsirigos, Geoffrey I Webb, Roger J Daly, Jiangning Song, Zhiyong Lu
Bioinformatics.2021; 37(22): 4291. CrossRef - A Review of Applications of Artificial Intelligence in Gastroenterology
Khalid Nawab, Ravi Athwani, Awais Naeem, Muhammad Hamayun, Momna Wazir
Cureus.2021;[Epub] CrossRef - Evaluating Cancer-Related Biomarkers Based on Pathological Images: A Systematic Review
Xiaoliang Xie, Xulin Wang, Yuebin Liang, Jingya Yang, Yan Wu, Li Li, Xin Sun, Pingping Bing, Binsheng He, Geng Tian, Xiaoli Shi
Frontiers in Oncology.2021;[Epub] CrossRef - Deep learning-based histopathological segmentation for whole slide images of colorectal cancer in a compressed domain
Hyeongsub Kim, Hongjoon Yoon, Nishant Thakur, Gyoyeon Hwang, Eun Jung Lee, Chulhong Kim, Yosep Chong
Scientific Reports.2021;[Epub] CrossRef - Deep Learning on Oral Squamous Cell Carcinoma Ex Vivo Fluorescent Confocal Microscopy Data: A Feasibility Study
Veronika Shavlokhova, Sameena Sandhu, Christa Flechtenmacher, Istvan Koveshazi, Florian Neumeier, Víctor Padrón-Laso, Žan Jonke, Babak Saravi, Michael Vollmer, Andreas Vollmer, Jürgen Hoffmann, Michael Engel, Oliver Ristow, Christian Freudlsperger
Journal of Clinical Medicine.2021; 10(22): 5326. CrossRef - A Pathologist-Annotated Dataset for Validating Artificial Intelligence: A Project Description and Pilot Study
Sarah N. Dudgeon, Si Wen, Matthew G. Hanna, Rajarsi Gupta, Mohamed Amgad, Manasi Sheth, Hetal Marble, Richard Huang, Markus D. Herrmann, Clifford H. Szu, Darick Tong, Bruce Werness, Evan Szu, Denis Larsimont, Anant Madabhushi, Evangelos Hytopoulos, Weijie
Journal of Pathology Informatics.2021; 12(1): 45. CrossRef - Artificial Intelligence in Medicine: A Multinational Multi-Center Survey on the Medical and Dental Students' Perception
Sotirios Bisdas, Constantin-Cristian Topriceanu, Zosia Zakrzewska, Alexandra-Valentina Irimia, Loizos Shakallis, Jithu Subhash, Maria-Madalina Casapu, Jose Leon-Rojas, Daniel Pinto dos Santos, Dilys Miriam Andrews, Claudia Zeicu, Ahmad Mohammad Bouhuwaish
Frontiers in Public Health.2021;[Epub] CrossRef - Digital/Computational Technology for Molecular Cytology Testing: A Short Technical Note with Literature Review
Robert Y. Osamura, Naruaki Matsui, Masato Kawashima, Hiroyasu Saiga, Maki Ogura, Tomoharu Kiyuna
Acta Cytologica.2021; 65(4): 342. CrossRef - Advances in Digital Pathology: From Artificial Intelligence to Label-Free Imaging
Frederik Großerueschkamp, Hendrik Jütte, Klaus Gerwert, Andrea Tannapfel
Visceral Medicine.2021; 37(6): 482. CrossRef - Feasibility of fully automated classification of whole slide images based on deep learning
Kyung-Ok Cho, Sung Hak Lee, Hyun-Jong Jang
The Korean Journal of Physiology & Pharmacology.2020; 24(1): 89. CrossRef - Same same but different: A Web‐based deep learning application revealed classifying features for the histopathologic distinction of cortical malformations
Joshua Kubach, Angelika Muhlebner‐Fahrngruber, Figen Soylemezoglu, Hajime Miyata, Pitt Niehusmann, Mrinalini Honavar, Fabio Rogerio, Se‐Hoon Kim, Eleonora Aronica, Rita Garbelli, Samuel Vilz, Alexander Popp, Stefan Walcher, Christoph Neuner, Michael Schol
Epilepsia.2020; 61(3): 421. CrossRef - Segmentation and Classification in Digital Pathology for Glioma Research: Challenges and Deep Learning Approaches
Tahsin Kurc, Spyridon Bakas, Xuhua Ren, Aditya Bagari, Alexandre Momeni, Yue Huang, Lichi Zhang, Ashish Kumar, Marc Thibault, Qi Qi, Qian Wang, Avinash Kori, Olivier Gevaert, Yunlong Zhang, Dinggang Shen, Mahendra Khened, Xinghao Ding, Ganapathy Krishnamu
Frontiers in Neuroscience.2020;[Epub] CrossRef - Artificial intelligence as the next step towards precision pathology
B. Acs, M. Rantalainen, J. Hartman
Journal of Internal Medicine.2020; 288(1): 62. CrossRef - Introduction to digital pathology and computer-aided pathology
Soojeong Nam, Yosep Chong, Chan Kwon Jung, Tae-Yeong Kwak, Ji Youl Lee, Jihwan Park, Mi Jung Rho, Heounjeong Go
Journal of Pathology and Translational Medicine.2020; 54(2): 125. CrossRef - Artificial intelligence with multi-functional machine learning platform development for better healthcare and precision medicine
Zeeshan Ahmed, Khalid Mohamed, Saman Zeeshan, XinQi Dong
Database.2020;[Epub] CrossRef - Scoring pleurisy in slaughtered pigs using convolutional neural networks
Abigail R. Trachtman, Luca Bergamini, Andrea Palazzi, Angelo Porrello, Andrea Capobianco Dondona, Ercole Del Negro, Andrea Paolini, Giorgio Vignola, Simone Calderara, Giuseppe Marruchella
Veterinary Research.2020;[Epub] CrossRef - Current Status of Computational Intelligence Applications in Dermatological Clinical Practice
Carmen Rodríguez-Cerdeira, José Luís González-Cespón, Roberto Arenas
The Open Dermatology Journal.2020; 14(1): 6. CrossRef - New unified insights on deep learning in radiological and pathological images: Beyond quantitative performances to qualitative interpretation
Yoichi Hayashi
Informatics in Medicine Unlocked.2020; 19: 100329. CrossRef - Artificial Intelligence in Cardiology: Present and Future
Francisco Lopez-Jimenez, Zachi Attia, Adelaide M. Arruda-Olson, Rickey Carter, Panithaya Chareonthaitawee, Hayan Jouni, Suraj Kapa, Amir Lerman, Christina Luong, Jose R. Medina-Inojosa, Peter A. Noseworthy, Patricia A. Pellikka, Margaret M. Redfield, Vero
Mayo Clinic Proceedings.2020; 95(5): 1015. CrossRef - Artificial intelligence in oncology
Hideyuki Shimizu, Keiichi I. Nakayama
Cancer Science.2020; 111(5): 1452. CrossRef - Artificial intelligence and the future of global health
Nina Schwalbe, Brian Wahl
The Lancet.2020; 395(10236): 1579. CrossRef - The future of pathology is digital
J.D. Pallua, A. Brunner, B. Zelger, M. Schirmer, J. Haybaeck
Pathology - Research and Practice.2020; 216(9): 153040. CrossRef - Weakly-supervised learning for lung carcinoma classification using deep learning
Fahdi Kanavati, Gouji Toyokawa, Seiya Momosaki, Michael Rambeau, Yuka Kozuma, Fumihiro Shoji, Koji Yamazaki, Sadanori Takeo, Osamu Iizuka, Masayuki Tsuneki
Scientific Reports.2020;[Epub] CrossRef - The use of artificial intelligence, machine learning and deep learning in oncologic histopathology
Ahmed S. Sultan, Mohamed A. Elgharib, Tiffany Tavares, Maryam Jessri, John R. Basile
Journal of Oral Pathology & Medicine.2020; 49(9): 849. CrossRef - Convergence of Digital Pathology and Artificial Intelligence Tools in Anatomic Pathology Practice: Current Landscape and Future Directions
Anil V. Parwani, Mahul B. Amin
Advances in Anatomic Pathology.2020; 27(4): 221. CrossRef - Advances in tissue-based imaging: impact on oncology research and clinical practice
Arman Rahman, Chowdhury Jahangir, Seodhna M. Lynch, Nebras Alattar, Claudia Aura, Niamh Russell, Fiona Lanigan, William M. Gallagher
Expert Review of Molecular Diagnostics.2020; 20(10): 1027. CrossRef - Current Trends of Artificial Intelligence for Colorectal Cancer Pathology Image Analysis: A Systematic Review
Nishant Thakur, Hongjun Yoon, Yosep Chong
Cancers.2020; 12(7): 1884. CrossRef - Explainable Machine Learning Model for Predicting GI Bleed Mortality in the Intensive Care Unit
Farah Deshmukh, Shamel S. Merchant
American Journal of Gastroenterology.2020; 115(10): 1657. CrossRef - Prediction of clinically actionable genetic alterations from colorectal cancer histopathology images using deep learning
Hyun-Jong Jang, Ahwon Lee, J Kang, In Hye Song, Sung Hak Lee
World Journal of Gastroenterology.2020; 26(40): 6207. CrossRef - Application of system analysis methods for modeling the development of hand-arm vibration syndrome: problems and approaches to solution
M P Diakovich, M V Krivov
Journal of Physics: Conference Series.2020; 1661(1): 012029. CrossRef - Histo-ELISA technique for quantification and localization of tissue components
Zhongmin Li, Silvia Goebel, Andreas Reimann, Martin Ungerer
Scientific Reports.2020;[Epub] CrossRef - Role of artificial intelligence in diagnostic oral pathology-A modern approach
Ayinampudi Bhargavi Krishna, Azra Tanveer, Pancha Venkat Bhagirath, Ashalata Gannepalli
Journal of Oral and Maxillofacial Pathology.2020; 24(1): 152. CrossRef - Applications of deep learning for the analysis of medical data
Hyun-Jong Jang, Kyung-Ok Cho
Archives of Pharmacal Research.2019; 42(6): 492. CrossRef - PROMISE CLIP Project: A Retrospective, Multicenter Study for Prostate Cancer that Integrates Clinical, Imaging and Pathology Data
Jihwan Park, Mi Jung Rho, Yong Hyun Park, Chan Kwon Jung, Yosep Chong, Choung-Soo Kim, Heounjeong Go, Seong Soo Jeon, Minyong Kang, Hak Jong Lee, Sung Il Hwang, Ji Youl Lee
Applied Sciences.2019; 9(15): 2982. CrossRef - Key challenges for delivering clinical impact with artificial intelligence
Christopher J. Kelly, Alan Karthikesalingam, Mustafa Suleyman, Greg Corrado, Dominic King
BMC Medicine.2019;[Epub] CrossRef - Deep Learning for Whole Slide Image Analysis: An Overview
Neofytos Dimitriou, Ognjen Arandjelović, Peter D. Caie
Frontiers in Medicine.2019;[Epub] CrossRef - Barriers to Artificial Intelligence Adoption in Healthcare Management: A Systematic Review
Mir Mohammed Assadullah
SSRN Electronic Journal .2019;[Epub] CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
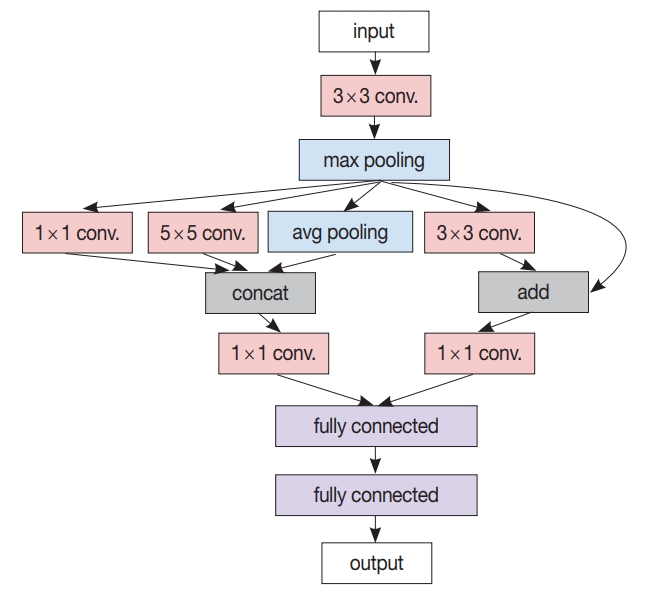
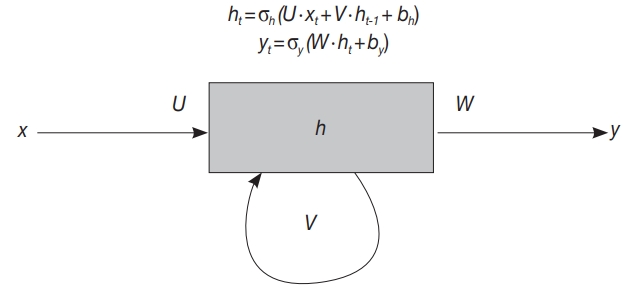

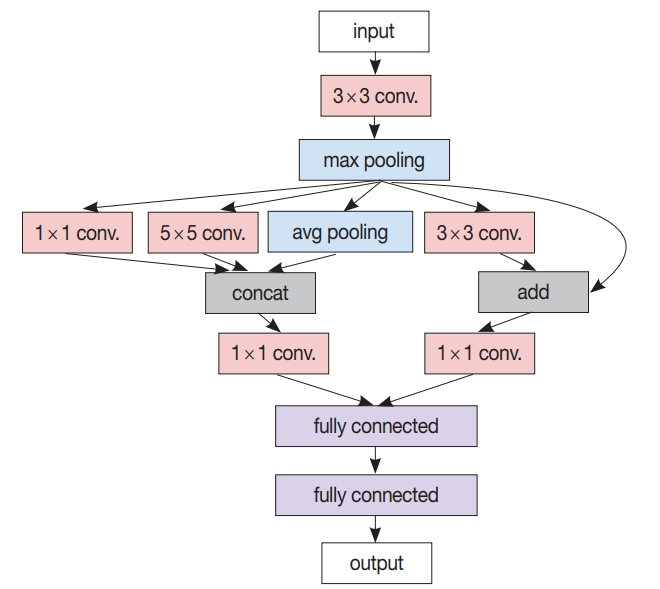
Fig. 1.
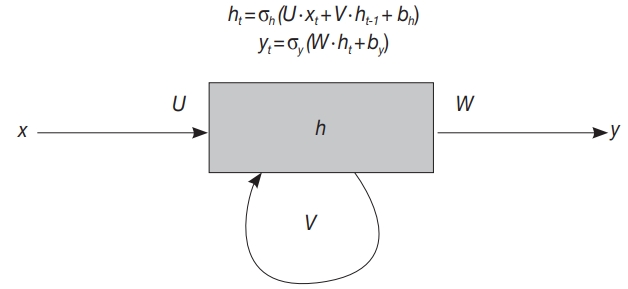
Fig. 2.
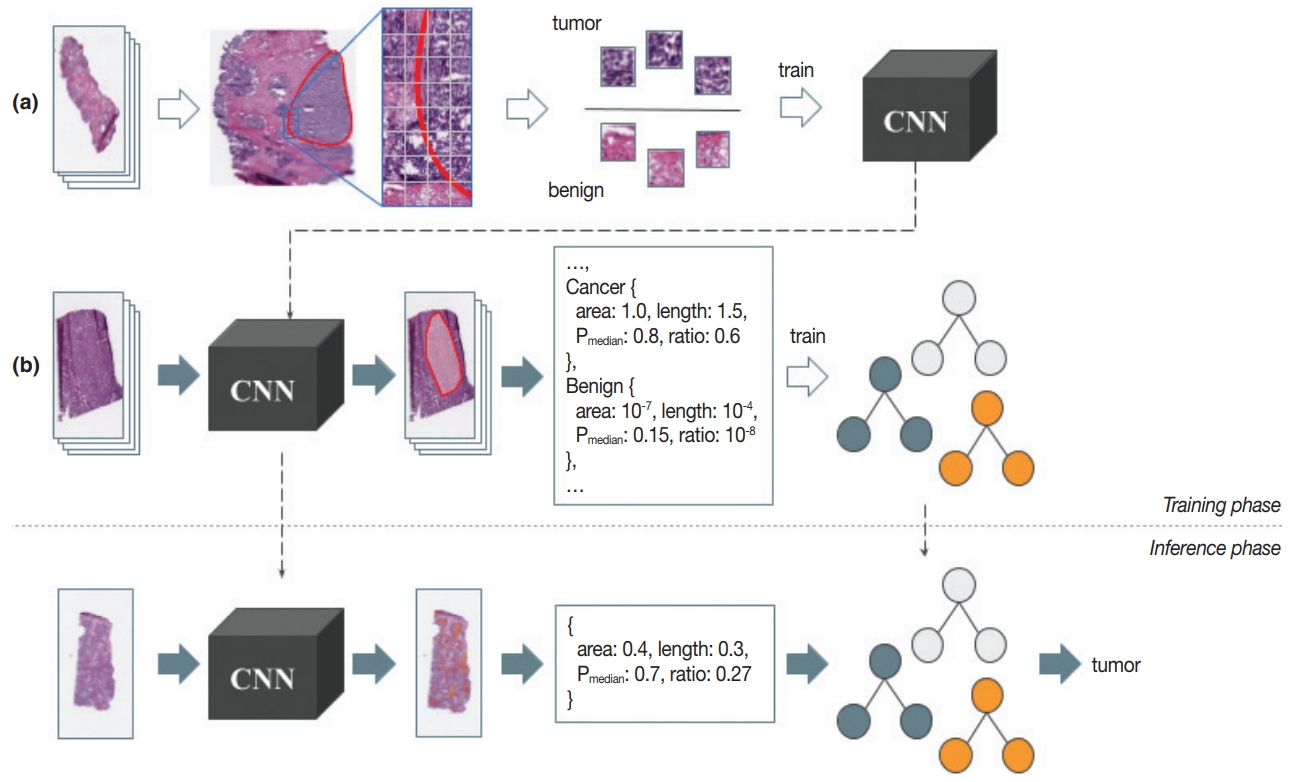
Fig. 3.
| Term | Abbreviation | Explanation |
|---|---|---|
| Artificial intelligence | AI | Intelligence represented by artificial things |
| Machine learning | ML | Data-driven statistical learning approach to AI |
| Deep learning | DL | Deep neural network based ML |
| Convolutional neural network | CNN | Neural network suitable for data with locality, e.g. image |
| Recurrent neural network | RNN | Neural network suitable for data with order dependency, e.g. sentence |
| Long short-term memory | LSTM | Recurrent neuron suitable for learning long-term dependency |
| Support vector machine | SVM | ML method that separates with regard to the trained hyperplane |
| k-nearest neighbor (search) | k-NN | ML method that classifies based on the classes of k similar training data |
| Conditional random field | CRF | ML method suitable for data with spatial/temporal dependency |
| Markov decision process | MDP | Modeling framework for a series of decisions and resulting outcomes |
| Multiple instance learning | MIL | ML approach suitable for labeled sets (whole slides) of unlabeled instances (lesions) |
| Region-of-interest | ROI | Image region containing things of predefined interest, e.g. nuclei, stroma, etc. |
| Area under receiver operating characteristic curve | AUC | Performance measure based on the area under the receiver operating characteristic curve, varying from 0.5 (lowest) to 1.0 (highest) |
| Author (year) | Disease | Data | Task | Model | Augmentation | Performance |
|---|---|---|---|---|---|---|
| Garud et al. (2017) [46] | Breast cancer | FNA cytology/175 (images) | Decision Benign/cancer | CNN | None | Image level decision acc. 89.7% |
| Li and Ping (2018) [47] | Lymph node metastasis | CAMELYON16/400 (WSIs) | Decision Yes/no | CNN + CRF | Color jitter, rotation, etc. | Patch level decision acc. 93.8% |
| Rannen Triki et al. (2018) [48] | Breast cancer | Frozen section OCT/4,921 (frames) | Decision Benign/cancer | CNN | None | Patch level decision acc. 94.96% |
| Ehteshami Bejnordi et al. (2018) [49] | Breast cancer | BREAST Stamp/2,387 (WSIs) | Decision Benign/cancer | CNN + CNN | None | WSI level decision AUC 0.962 |
| Litjens et al. (2016) [50] | Lymph node metastasis | Lymph node specimen/271 (samples) | Decision Yes/no | CNN | None | Sample level decision AUC 0.90 |
| Cires¸ an et al. (2013) [51] | Breast cancer | MITOS/300 mitosis in 50 images | Mitosis detection | CNN | Rotation, flip, etc. | Detection F1-score 0.782 |
| Teramoto et al. (2017) [52] | Lung cancer | FNA cytology/298 (images) | Classification | CNN | Rotation, flip, etc. | Overall classification acc. 71.1% |
| Adeno-Squamous cell | ||||||
| Small cell | ||||||
| Yu et al. (2016) [53] | Lung cancer | TCGA-LUAD/1,074 | Decision Benign/cancer | SVM | None | Patch level decision AUC 0.85 |
| TCGA-LUSC/1,111 | Survival analysis | |||||
| Stanford TMA/294 (samples) | ||||||
| Coudray et al. (2018) [54] | Lung cancer | TCGA lung cancer/1,635 (samples) | Classification | CNN | None | Overall classification AUC 0.97 |
| Adeno-Squamous cell | STK11 mutation decision AUC 0.85 | |||||
| Benign | ||||||
| Multi-task decision | ||||||
| Gene mutation | ||||||
| Campanella et al. (2018) [55] | Prostate cancer | Needle biopsy/12,160 (samples) | Decision Benign/cancer | CNN (MIL) | None | Sample level decision AUC 0.979 |
| Arvaniti et al. (2018) [56] | Prostate cancer | TMA/886 (samples) | Classification Gleason score | CNN +scoring rule | Rotation, flip, color jitter | Model-pathologist Cohen’s kappa 0.71 |
| Zhou et al. (2017) [57] | Prostate cancer | TCGA-PRAD/368 (cases) | Decision 3 + 4/4 + 3 | CNN | None | Sample level decision acc. 75% |
| Nagpal et al. (2018) [58] | Prostate cancer | TCGA-PRAD + others/train 1,226, test 331 (slides) | Classification Gleason group | CNN + k-NN | None | Overall classification acc. 70% |
| Survival analysis | C-index 0.697 | |||||
| Litjens et al. (2016) [50] | Prostate cancer | Needle biopsy / 225 (WSIs) | Decision Benign/cancer | CNN | None | Slide level decision AUC 0.99 |
| Ertosun and Rubin (2015) [59] | Brain cancer | TCGA-GBM & LGG (unknown size) | Classification | CNN + CNN | Color transform to H&E | GBM/LGG decision acc. 96% |
| GBM | LGG grade decision acc. 71% | |||||
| LGG grade 2 | ||||||
| LGG grade 3 | ||||||
| Mobadersany et al. (2018) [60] | Brain cancer | TCGA-GBM & LGG/1,061 (samples) | Survival analysis | CNN | Rotation, normalization | C-index 0.754 |
| Wu et al. (2018) [61] | Ovarian cancer | Biopsy/7,392 (images) | Classification Subtypes | CNN | Rotation, image enhancement | Overall classification acc. 78.2% |
| Zhang et al. (2017) [62] | Cervix cancer | HEMLBC/1,978 Herlev/917 (images) | Decision Benign/cancer | CNN | Rotation, translation, | Image level decision AUC 0.99 |
| Xu et al. (2017) [63] | Sickle cell disease | Red-blood cell/7,206 (patches) | Classification Cell types | CNN | Rotation, flip, translation, etc. | Cell level classification acc. 87.5% |
| Meier et al. (2018) [64] | Gastric cancer | TMA/469 (samples) CD8/Ki67 IHC | Survival analysis | CNN | None | Stratification by risk successful (p < .01) |
| Xie et al. (2016) [65] | - | Synthetic fluorescence microscopy cell/200 (images) | Cell counting | CNN | None | Mean absolute error < 2% |
| Tuominen et al. (2010) [66] | - | IHC stained breast cancer slides/100 | Cell counting | Comp. vision | None | Correlation coefficient 0.98 |
CNN, convolutional neural network; MIL, multiple instance learning; SVM, support vector machine; AUC, area under receiver operating characteristic curve; k-NN, k-nearest neighbor; WSI, whole slide image; CRF, Conditional random field; TCGA, The Cancer Genome Atlas; TMA, tissue microarray; IHC, immunohistochemistry; GBM, glioblastoma multiforme; LGG, lower grade glioma.

 E-submission
E-submission