Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 56(3); 2022 > Article
-
Original Article
Prognostic and clinicopathological significance of Fusobacterium nucleatum in colorectal cancer: a systemic review and meta-analysis -
Younghoon Kim1,2
 , Nam Yun Cho1
, Nam Yun Cho1 , Gyeong Hoon Kang1,2
, Gyeong Hoon Kang1,2
-
Journal of Pathology and Translational Medicine 2022;56(3):144-151.
DOI: https://doi.org/10.4132/jptm.2022.03.13
Published online: May 15, 2022
1Laboratory of Epigenetics, Cancer Research Institute, Seoul National University College of Medicine, Seoul, Korea
2Department of Pathology, Seoul National University College of Medicine, Seoul, Korea
- Corresponding Author: Gyeong Hoon Kang, MD, PhD, Department of Pathology, Seoul National University College of Medicine, 103 Daehak-ro, Jongno-gu, Seoul 03080, Korea Tel: +82-22-740-8272, Fax: +82-2-765-5600, E-mail: ghkang@snu.ac.kr
© 2022 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- Fusobacterium nucleatum has been identified to promote tumor progression in colorectal cancer (CRC). However, association between F. nucleatum and prognostic or clinicopathological features has been diverse among studies, which could be affected by type of biospecimen (formalin-fixed paraffin-embedded or fresh frozen [FF]).
-
Methods
- Articles were systemically reviewed for studies that included the correlation between F. nucleatum and prognosis or clinicopathological features in CRC.
-
Results
- Ten articles, eight studies with survival-related features involving 3,199 patients and nine studies with clinical features involving 2,655 patients, were eligible for the meta-analysis. Overall survival, disease-free survival, and cancer-specific survival were all associated with worse prognosis in F. nucleatum–high patients (p<.05). In subgroup analysis, only studies with FF tissues retained prognostic significance with F. nucleatum. In meta-analysis of clinicopathological variables, F. nucleatum level was associated with location within colon, pT category, MLH1 hypermethylation, microsatellite instability status, and BRAF mutation regardless of type of biospecimen. However, lymph node metastasis and KRAS mutation was only associated with F. nucleatum level in FF-based studies.
-
Conclusions
- In conclusion, type of biospecimen could affect the role of F. nucleatum as a biomarker associated with clinicopathological features and prognosis.
- Literature search strategy
- Articles relevant with the subject were retrieved from electronic databases, PubMed, Embase, and Web of Science until December 3 of 2020. Search terms were, “Fusobacterium nucleatum, Fusobacterium, Fusobacteria, or F. nucleatum” and “colorectal cancer, colon cancer, rectal cancer, colorectal carcinoma, or CRC”. Each article was examined carefully to determine studies with identical patient population. When multiple studies present overlapping patient data, we selected the studies with the largest patient cohort for clinical data and survival data, respectively.
- Inclusion and exclusion criteria
- Studies that were eligible for the inclusion in this meta-analysis were as follows: (1) full length articles published in English that detected Fusobacterium nucleatum from human tumor samples derived from CRC tissue, (2) studies with CRC samples divided into high and low (or high and low/negative) according to F. nucleatum level, (3) studies that included clinicopathological, molecular, or survival-associated data that were correlated with F. nucleatum level. Studies that only contain abstract or posters, studies could not be arranged as dichotomized F. nucleatum level data, studies with insufficient data, studies that used more than one type of biospecimen to detect F. nucleatum level, and studies that detected F. nucleatum level in stools were excluded.
- Data extraction and quality assessment
- Two reviewers (Y.K. and G.H.K.) independently collected data from eligible articles according to the criteria. Newcastle-Ottawa Scale was used to evaluate study quality. A score higher than six out of nine criteria was defined as qualified. Name of author, year of publication, number of total patients, number of F. nucleatum positive patients, type of biospeicmen (FFPE or FF), method of F. nucleatum level detection, survival data, and clinicopathological data associated with F. nucleatum level were collected from each study. Survival data were extracted as hazard ratio [HR] and 95% confidence intervals [CIs] from univariate Cox analysis. When only Kaplan-Meier curves were presented for survival analysis, HR and 95% CIs were estimated by using Engauge Digitizer V9.8 (http://digitizer.sourceforge.net) and spreadsheets provided by Tierney et al. [21]. When a three-tier survival was given (negative, low, and high) we pooled negative and low as reference to calculate HR and 95% CIs for high and rearranged it into two-tier (low/negative and high).
- Statistical analysis
- Meta-analysis was conducted using R software ver. 3.5.3 (https://cran.r-project.org/). Pooled HR and 95% CIs for overall survival (OS), disease-free survival (DFS), and cancer-specific survival (CSS) and pooled odds ratio and 95% CIs for clinicopathological features were analyzed via R package ‘meta’ and ‘dmetar’. Subgroup analysis was performed if multiple types of biospecimen existed within a pooled analysis. Statistic heterogeneity was determined by Cochran’s Q and I2. When heterogeneity was observed (p<.05 or I2>50%) random effect model was used for the analysis instead of fixed effect model. A graphical funnel plot was used to evaluate publication bias.
MATERIALS AND METHODS
- Literature search and study characteristics
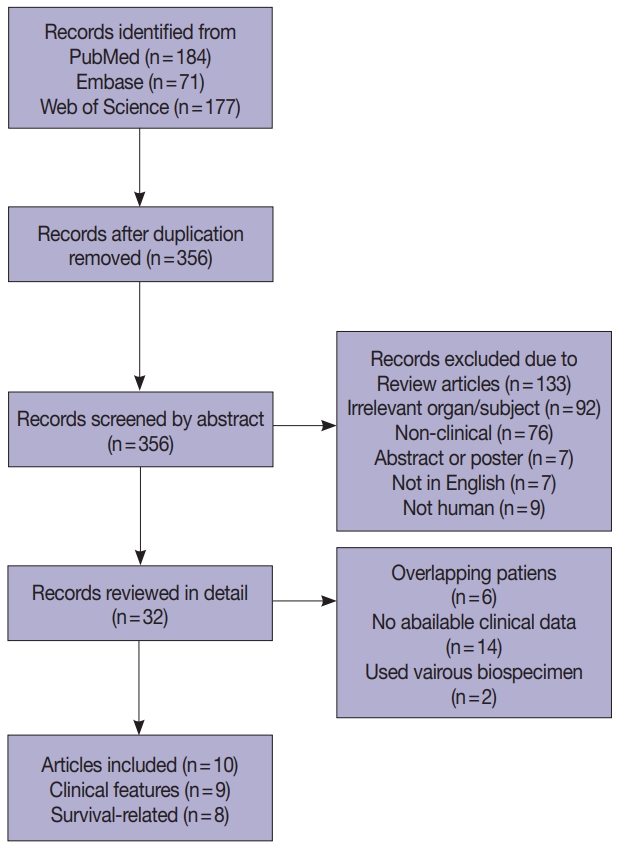
- Four hundred and thirty-two records were collected from PubMed, Embase, and Web of Science (Fig. 1). After removing 76 duplications, 356 records were screened by abstract and 32 records were categorized as being relevant with the subject. These records were reviewed in detail and 10 articles, eight studies with survival-related features involving 3,199 patients [14-18,22-24] and nine studies with clinical features involving 2,655 patients [14-17,22-25], were selected eligible for the meta-analysis (Table 1). Seven studies were included in the meta-analysis for both survival-related and clinical features [14-17,22-24]. Two studies had overlapping patients but were used for meta-analysis of either clinical or survival-related features, respectively [18,25]. Six out of the 10 articles assessed F. nucleatum level with FFPE tissues [6,16-18,24,25] and four articles used FF tissues to detect F. nucleatum [14,15,22,23].
- Meta-analysis between F. nucleatum level and survival
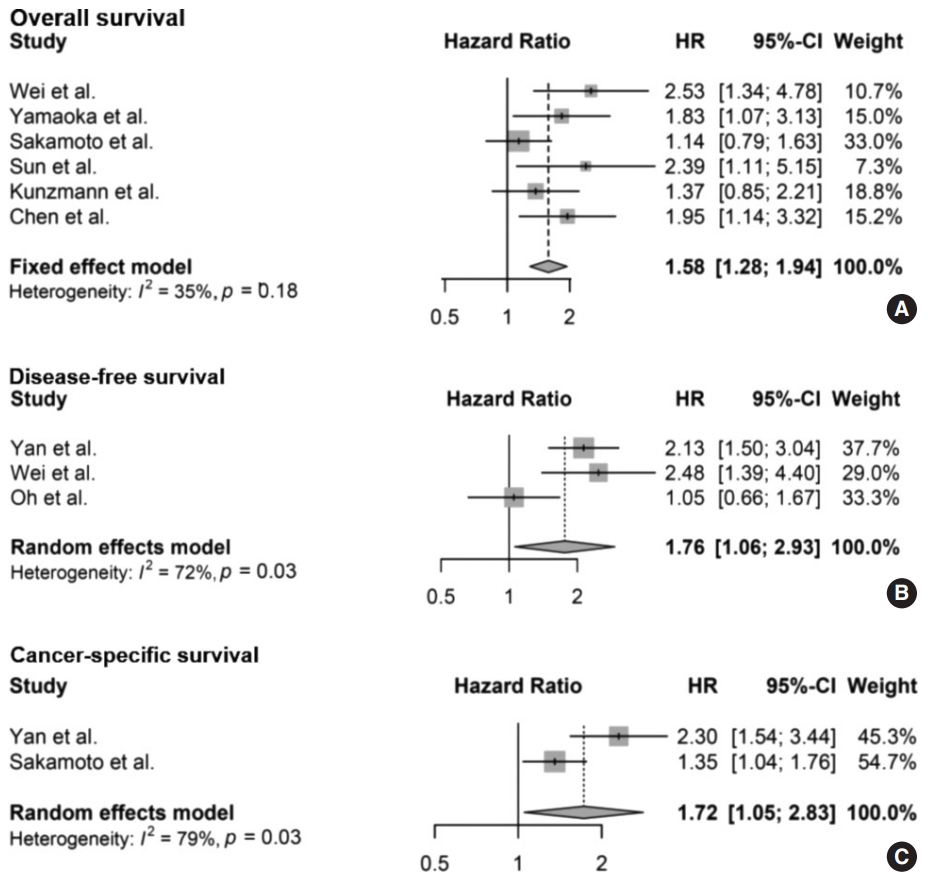
- The meta-analysis between F. nucleatum level and prognostic values of OS (Fig. 2A), DFS (Fig. 2B), and CSS (Fig. 2C) in CRC has shown that high F. nucleatum level is associated with poor OS, DFS, and CSS (OS: HR, 1.58; 95% CI, 1.28 to 1.94; p<.001; DFS: HR, 1.76; 95% CI, 1.06 to 2.93; p=.030; CSS: HR, 1.72; 95% CI, 1.05 to 2.83; p=.031, respectively). In subgroup analysis, studies with FF samples (p<.001), but not those with FFPE samples (p=.168) retained prognostic significance between F. nucleatum level and OS (Table 2, Supplementary Fig. S1). Furthermore, studies with FFPE samples did not show significant correlation between F. nucleatum level and DFS (p=.214). Analyzing studies with stage I though IV CRC did not affect prognostic significance between F. nucleatum level and OS (HR, 1.41; 95% CI, 1.11 to 1.80; p=.005).
- Meta-analysis between F. nucleatum level and clinicopathological features
- Meta-analysis for correlation between F. nucleatum level and clinicopathological features of CRC are assessed in Table 3. High F. nucleatum level was associated with proximal colon location (p<.001), worse tumor differentiation (p=.002), high pT category (p<.001), presence of distal metastasis (p=.007), MLH1 hypermethylation (p<.001), and MSI-high CRC (p=.001) when all studies were considered. In subgroup analysis (Table 4), studies with FFPE samples showed significant correlation between high F. nucleatum level and proximal colon location (p=.001), worse tumor differentiation (p=.015), high pT category (p<.001), MLH1 hypermethylation (p<.001), CpG island methylator phenotype (CIMP)–high status (p<.001), and MSI-high status (p<.001). Studies with FF samples, on the other hand, demonstrated that high F. nucleatum level is significantly correlated with high pT category (p=.001), presence of lymph node metastasis (p=.001), and presence of KRAS mutation (p=.007), which showed borderline significance with high F. nucleatum level in overall studies (p=.052).
- Publication bias
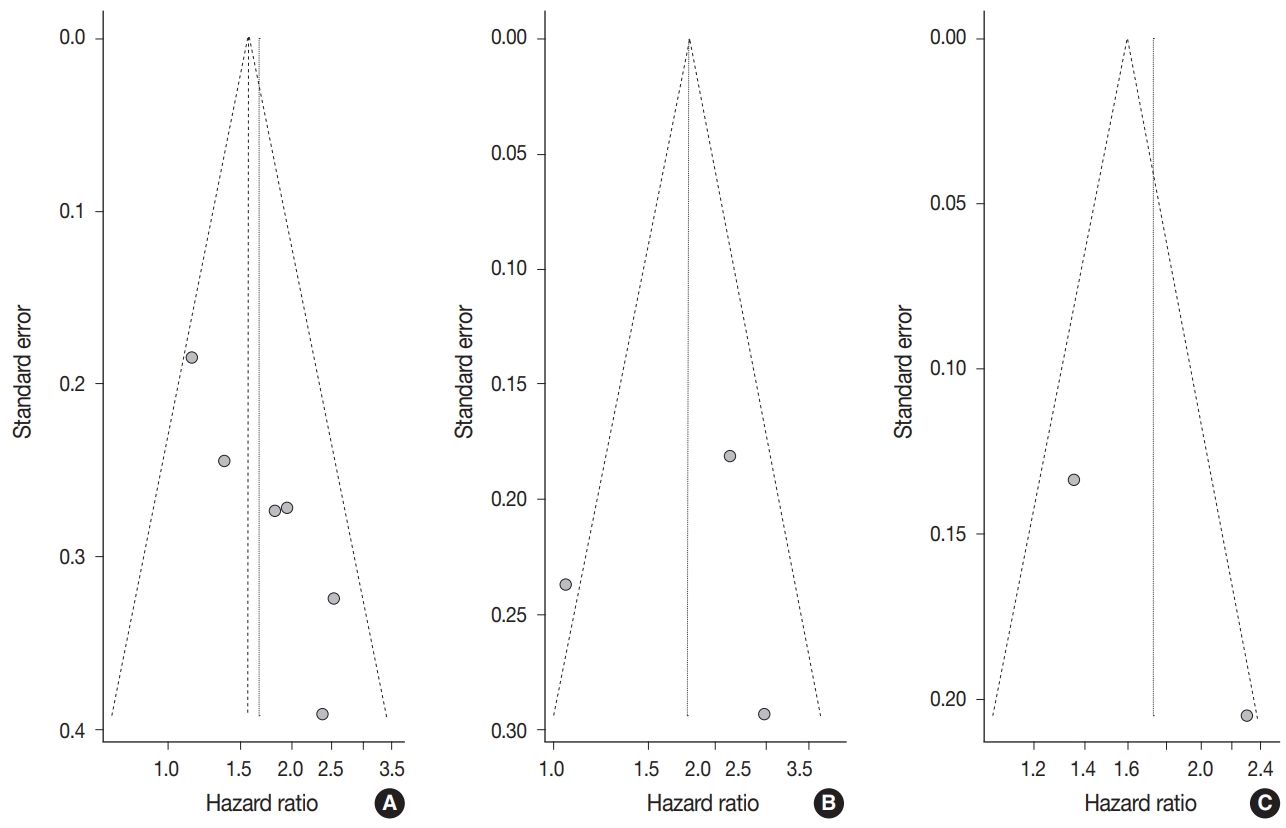
- Begg’s funnel plot was evaluated for presence of potential publication bias in meta-analysis associated with survival (Fig. 3). Visible asymmetry was observed with Egger’s test reporting p-value of .006 for OS but not for DFS (p=.935).
RESULTS
- F. nucleatum has been associated with cancer development and progression of CRC, as well as a potential target of cancer treatment. F. nucleatum is abdudnant in CRC tissues from patients with metastasis than those from non-metastatic CRC patients [26]. Presence F. nucleatum was essential in establishment of CRC patients-derived xenografts in mice [13]. Not only does infection of F. nucleatum has promoted metastasis in CRC cells via autophage but also is found in distant metastasis sites including liver, lung, and lymph nodes, although the mechanism is yet to be unveiled [9,13,26,27]. Moreover, previous studies have suggested that F. nucleatum provoke chemoresistance inhibiting caspase cascade and activating autophagy pathway to prevent apoptosis in CRC cells [11,28].
- It is still unclear if F. nucleatum is a significant driver of tumor initiation and F. nucleatum level detected from CRC tissues and association with prognosis and clinicopathological variables has not been consistent among studies. Herein, we hypothesized that this could be partially caused by different type of biospecimen being used for each study and investigated its affect by a systemic review and meta-analysis with the difference between FFPE and FF into consideration.
- High F. nucleatum level was associated with worse survival in OS, DFS, and CSS in overall meta-analysis. Subgroup analysis revealed that studies with FF show a significant correlation between F. nucleatum level and OS. However, studies with FFPE tissues show that a significant correlation between F. nucleatum and OS or DFS is not observed.
- Association between F. nucleatum level and clinicopathological features are much more diverse. A pooled analysis of FFPE and FF studies together shows that high F. nucleatum level samples are significantly enriched in proximal colon, but on subgroup analysis, only studies using FFPE samples retain statistical significance for the same variable. Similar findings could be observed for correlation between F. nucleatum level and distant metastasis. On the contrary, pT category, MLH1 hypermethylation, and MSI status remained its positive correlation with F. nucleatum level regardless of the biospecimen being used. Lymph node metastasis and KRAS mutation were only significant in FF tissues and MLH1 hypermethylation was only significant in FFPE tissues but not in its amalgamated state. Interestingly, the aforementioned features are all considered as hallmark characteristics of MSI-high CRC [29]. These findings emphasize that regardless of type of specimen, high F. nucleatum level is strongly indicative of MSI-high status.
- There are several putative explanations for discrepancy of results between biospecimen. Mean patient number for studies with FFPE tissue (515.4 patients per study) is more than three times larger than the mean patient number for studies with FF tissues (155.5 patients per study). On the contrary, F. nucleatum positivity is significantly higher in studies with FF tissues compared with those of FFPE tissue (41.8% and 24.4%. respectively) which may be a result of a lower threshold in studies with FF tissues to compensate for smaller sample size. This may allude to the fact that features with relatively low incidence such as KRAS mutation, could have a higher chance of being categorized as high F. nucleatum level sample in studies with FF compared with studies with FFPE.
- As an alternative explanation, lower F. nucleatum positivity in FFPE tissues may reflect the inherited nature of FFPE samples rather than inflation of threshold in studies with FF tissues. In a study that measured F. nucleatum level in paired FFPE and FF tissues of CRC, FF tissues had higher detection rate of overall F. nucleatum resulting in difference in survival and clinicopathological features from FFPE-measured samples [20]. Furthermore, concordance between F. nucleatum positive cases in FF and those in FFPE samples were low. Discrepancy between DNA results of FFPE and FF samples has also been reported in other paired tumor tissues [30]. This was in part due to inter-strand crosslink between complementary DNA strands which could be aggravated by prolonged fixation [31]. Other potential factors of low F. nucleatum detection in FFPE samples includes fragmentation and degeneration of DNA, paraffin embedding, and storage process [25,32].
- Recently, two articles analyzed the correlations between F. nucleatum level and prognosis in CRC with meta-analysis [33,34]. Both articles showed a significant correlation between F. nucleatum level and OS, but only one study showed a significant correlation between F. nucleatum level and DFS. Herein, we suggest that type of biospecimen that was included in each meta-analysis could have affected the discrepancy between the articles.
- Limitation of our study is that we cannot identify pathological and molecular discrepancies between FF and FFPE tissues such as correlation between F. nucleatum level and T cell density, CIMP, and various immunohistochemical results due to lack of studies with FF that obtain related data. Moreover, although metaanalysis provides evidence that discordance between FF and FFPE samples are present in detecting F. nucleatum, we could not narrow down the cause of the misalignment between the two biospecimen.
- In conclusion, F. nucleatum level is significantly associated with pathological and molecular features including pT category, MLH1 hypermethylation, and MSI status regardless of biospecimen, but other survival, clinical, and molecular features, such as location of tumor within the colon, distant metastasis, and KRAS mutation, could be affected by type of biospecimen in CRC. Therefore, although a strong association between MSI and F. nucleatum is always evident, selection of biospecimen could affect the role of F. nucleatum as a biomarker in CRC to predict and monitor tumor progression and prognosis as well as evaluate treatment effect.
DISCUSSION
Supplementary Information
Ethics Statement
Not applicable.
Availability of Data and Material
All data generated or analyzed during the study are included in this published article (and its supplementary information files).
Code Availability
Not applicable.
Author Contributions
Conceptualization: YK. Methodology: YK, GHK. Resources: NYC. Writing—original draft: YK. Writing—review & editing: GHK. Approval of final manuscript: all authors.
Conflicts of Interest
The authors declare that they have no potential conflicts of interest.
Funding Statement
This work was financially supported by a grant from the National Research Foundation (NRF) funded by the Korean Ministry of Science and ICT (2019R1F1A1061227) and a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: HI14C1277).
| Authors | Year | No. of patients | No. of positives | Biospecimen | Method | Survival | Clinical | Stage |
|---|---|---|---|---|---|---|---|---|
| Yan et al. [16] | 2017 | 280 | 187 | FFPE | qPCR | O | O | III/IV |
| Wei et al. [14] | 2016 | 180 | 90 | FF | qPCR | O | O | I–III |
| Yamaoka et al. [15] | 2018 | 100 | 50 | FF | ddPCR | O | O | I–IV |
| Samkamoto et al. [18] | 2018 | 1,069 | 67 | FFPE | qPCR | O | I–IV | |
| Sun et al. [22] | 2016 | 152 | 59 | FF | qPCR | O | O | I–IV |
| Kunzmann et al. [23] | 2019 | 190 | 61 | FF | qPCR | O | O | I–IV |
| Oh et al. [17] | 2019 | 593 | 204 | FFPE | qPCR | O | O | II/III |
| Chen et al. [24] | 2019 | 91 | 25 | FFPE | qPCR | O | O | II/III |
| Ito et al. [6] | 2015 | 511 | 143 | FFPE | qPCR | O | I–IV | |
| Mima et al. [25] | 2015 | 1,102 | 69 | FFPE | qPCR | O | I–IV |
| Survival | Specimen | Study | HR | 95% CI | p-value |
|---|---|---|---|---|---|
| Overall survival | FFPE | 2 | 1.43 | 0.85–2.42 | .168 |
| FF | 4 | 1.84 | 1.36–2.48 | < .001 | |
| Disease-free survival | FFPE | 2 | 1.52 | 0.76–3.04 | .214 |
- 1. Wu Y, Wu J, Chen T, et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis in mice via a Toll-like receptor 4/p21-activated kinase 1 cascade. Dig Dis Sci 2018; 63: 1210-8. ArticlePubMedPDF
- 2. Kostic AD, Chun E, Robertson L, et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 2013; 14: 207-15. ArticlePubMedPMC
- 3. Marchesi JR, Adams DH, Fava F, et al. The gut microbiota and host health: a new clinical frontier. Gut 2016; 65: 330-9. ArticlePubMedPMC
- 4. Kostic AD, Gevers D, Pedamallu CS, et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res 2012; 22: 292-8. ArticlePubMedPMC
- 5. Castellarin M, Warren RL, Freeman JD, et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res 2012; 22: 299-306. ArticlePubMedPMC
- 6. Ito M, Kanno S, Nosho K, et al. Association of Fusobacterium nucleatum with clinical and molecular features in colorectal serrated pathway. Int J Cancer 2015; 137: 1258-68. ArticlePubMed
- 7. McCoy AN, Araujo-Perez F, Azcarate-Peril A, Yeh JJ, Sandler RS, Keku TO. Fusobacterium is associated with colorectal adenomas. PLoS One 2013; 8: e53653.ArticlePubMedPMC
- 8. Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/beta-catenin signaling via its FadA adhesin. Cell Host Microbe 2013; 14: 195-206. PubMedPMC
- 9. Abed J, Emgard JE, Zamir G, et al. Fap2 mediates Fusobacterium nucleatum colorectal adenocarcinoma enrichment by binding to tumor-expressed Gal-GalNAc. Cell Host Microbe 2016; 20: 215-25. ArticlePubMedPMC
- 10. Yang Y, Weng W, Peng J, et al. Fusobacterium nucleatum increases proliferation of colorectal cancer cells and tumor development in mice by activating Toll-like receptor 4 signaling to nuclear factorkappaB, and up-regulating expression of microRNA-21. Gastroenterology 2017; 152: 851-66. ArticlePubMedPMC
- 11. Yu T, Guo F, Yu Y, et al. Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell 2017; 170: 548-63. ArticlePubMedPMC
- 12. Mandip KC, Steer CJ. Novel mechanisms of chemoresistance by Fusobacterium nucleatum involve not so novel pathways of microRNAs and autophagy. Transl Cancer Res 2018; 7(Suppl 1): S10-5. Article
- 13. Bullman S, Pedamallu CS, Sicinska E, et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 2017; 358: 1443-8. ArticlePubMedPMC
- 14. Wei Z, Cao S, Liu S, et al. Could gut microbiota serve as prognostic biomarker associated with colorectal cancer patients’ survival? A pilot study on relevant mechanism. Oncotarget 2016; 7: 46158-72. ArticlePubMedPMC
- 15. Yamaoka Y, Suehiro Y, Hashimoto S, et al. Fusobacterium nucleatum as a prognostic marker of colorectal cancer in a Japanese population. J Gastroenterol 2018; 53: 517-24. ArticlePubMedPDF
- 16. Yan X, Liu L, Li H, Qin H, Sun Z. Clinical significance of Fusobacterium nucleatum, epithelial-mesenchymal transition, and cancer stem cell markers in stage III/IV colorectal cancer patients. Onco Targets Ther 2017; 10: 5031-46. PubMedPMC
- 17. Oh HJ, Kim JH, Bae JM, Kim HJ, Cho NY, Kang GH. Prognostic impact of Fusobacterium nucleatum depends on combined tumor location and microsatellite instability status in stage II/III colorectal cancers treated with adjuvant chemotherapy. J Pathol Transl Med 2019; 53: 40-9. ArticlePubMedPMCPDF
- 18. Sakamoto Y, Mima K, Daitoku N, et al. Fusobacterium nucleatum in colorectal cancer liver metastasis and patient prognosis. Cancer Sci 2018; 109: 1360.
- 19. Chernyavskiy P, Kennerley VM, Jemal A, Little MP, Rosenberg PS. Heterogeneity of colon and rectum cancer incidence across 612 SEER counties, 2000-2014. Int J Cancer 2019; 144: 1786-95. ArticlePubMedPMCPDF
- 20. de Carvalho AC, de Mattos Pereira L, Datorre JG, et al. Microbiota profile and impact of Fusobacterium nucleatum in colorectal cancer patients of Barretos Cancer Hospital. Front Oncol 2019; 9: 813.PubMedPMC
- 21. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into metaanalysis. Trials 2007; 8: 16.ArticlePubMedPMCPDF
- 22. Sun Y, An QM, Tian XY, et al. Fusobacterium nucleatum infection is correlated with tumor metastasis and postoperative survival of colorectal cancer patients in China. Transl Cancer Res 2016; 5: 579-88. Article
- 23. Kunzmann AT, Proenca MA, Jordao HW, et al. Fusobacterium nucleatum tumor DNA levels are associated with survival in colorectal cancer patients. Eur J Clin Microbiol Infect Dis 2019; 38: 1891-9. ArticlePubMedPMCPDF
- 24. Chen Y, Lu Y, Ke Y, Li Y. Prognostic impact of the Fusobacterium nucleatum status in colorectal cancers. Medicine (Baltimore) 2019; 98: e17221.ArticlePubMedPMC
- 25. Mima K, Sukawa Y, Nishihara R, et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol 2015; 1: 653-61. ArticlePubMedPMC
- 26. Chen Y, Chen Y, Zhang J, et al. Fusobacterium nucleatum promotes metastasis in colorectal cancer by activating autophagy signaling via the upregulation of CARD3 expression. Theranostics 2020; 10: 323-39. ArticlePubMedPMC
- 27. Wang S, Liu Y, Li J, et al. Fusobacterium nucleatum acts as a procarcinogenic bacterium in colorectal cancer: from association to causality. Front Cell Dev Biol 2021; 9: 710165.ArticlePubMedPMC
- 28. Zhang S, Yang Y, Weng W, et al. Fusobacterium nucleatum promotes chemoresistance to 5-fluorouracil by upregulation of BIRC3 expression in colorectal cancer. J Exp Clin Cancer Res 2019; 38: 14.ArticlePubMedPMCPDF
- 29. Bae JM, Kim JH, Kang GH. Molecular subtypes of colorectal cancer and their clinicopathologic features, with an emphasis on the serrated neoplasia pathway. Arch Pathol Lab Med 2016; 140: 406-12. ArticlePubMedPDF
- 30. Wen X, Jeong S, Kim Y, et al. Improved results of LINE-1 methylation analysis in formalin-fixed, paraffin-embedded tissues with the application of a heating step during the DNA extraction process. Clin Epigenetics 2017; 9: 1.ArticlePubMedPMCPDF
- 31. Freifelder D, Davison PF. Physicochemical studies on the reaction between formaldehyde and DNA. Biophys J 1963; 3: 49-63. ArticlePubMedPMC
- 32. Imrit K, Goldfischer M, Wang J, et al. Identification of bacteria in formalin-fixed, paraffin-embedded heart valve tissue via 16S rRNA gene nucleotide sequencing. J Clin Microbiol 2006; 44: 2609-11. ArticlePubMedPMCPDF
- 33. Gethings-Behncke C, Coleman HG, Jordao HWT, et al. Fusobacterium nucleatum in the colorectum and its association with cancer risk and survival: a systematic review and meta-analysis. Cancer Epidemiol Biomarkers Prev 2020; 29: 539-48. ArticlePubMedPDF
- 34. Huangfu SC, Zhang WB, Zhang HR, et al. Clinicopathological and prognostic significance of Fusobacterium nucleatum infection in colorectal cancer: a meta-analysis. J Cancer 2021; 12: 1583-91. ArticlePubMedPMC
REFERENCES
Figure & Data
References
Citations

- GUT DYSBIOSIS IN THE ADENOMA-CARCINOMA SEQUENCE: A COMPREHENSIVE REVIEW OF CLINICAL AND PRECLINICAL EVIDENCE
Paulina Biedroń, Anna Skrzypek, Maciej Hutkowski, Mikołaj Zbrożek, Zuzanna Chwostek, Bartłomiej Kosiarski, Patrycja Markowicz, Hanna Maruchniak, Wiktoria Marzec, Krzysztof Biłyk
International Journal of Innovative Technologies in Social Science.2026;[Epub] CrossRef - Associations of epidemiologic risk factors with Fusobacterium nucleatum and bacterial alpha diversity in the colorectal tumor-associated microbiota
Courtney M. Hill, Rachel C. Malen, Adriana M. Reedy, Orsalem Kahsai, Keith Curtis, Ningxin Ma, Timothy W. Randolph, Jing Ma, Claire E. Thomas, Shuji Ogino, John D. Potter, Daniel D. Buchanan, Polly A. Newcomb, Meredith A. J. Hullar, Amanda I. Phipps
Cancer Causes & Control.2026;[Epub] CrossRef - Gut microbiome in colorectal cancer: recent advances and clinical implications
Jun Yong Han, Min Jung Kim, Ji Won Park, Seung-Yong Jeong
Annals of Coloproctology.2026; 42(1): 72. CrossRef - Spermine Oxidase Serves as a Key Functional Node in Microbial Dysbiosis–Induced Breast Carcinogenesis
Deeptashree Nandi, Sheetal Parida, Deepak Verma, Jackson R. Foley, Tracy Murray Stewart, Preethi Korangath, Sowjanya Thatikonda, Sumit Siddharth, Qitong Wu, Mingyang Yi, William Bishai, Robert Ivkov, Cynthia L. Sears, Robert A. Casero, Dipali Sharma
Cancer Research.2026; 86(8): 1920. CrossRef - Unraveling the Role of Fusobacterium nucleatum in Colorectal Cancer: Molecular Mechanisms and Pathogenic Insights
Linda Galasso, Fabrizio Termite, Irene Mignini, Giorgio Esposto, Raffaele Borriello, Federica Vitale, Alberto Nicoletti, Mattia Paratore, Maria Elena Ainora, Antonio Gasbarrini, Maria Assunta Zocco
Cancers.2025; 17(3): 368. CrossRef - Intratumoural pks Escherichia coli is associated with risk of metachronous colorectal cancer and adenoma development in people with Lynch syndrome
Yen Lin Chu, Peter Georgeson, Mark Clendenning, Khalid Mahmood, Romy Walker, Julia Como, Sharelle Joseland, Susan G. Preston, Toni Rice, Brigid M. Lynch, Roger L. Milne, Melissa C. Southey, Graham G. Giles, Amanda I. Phipps, John L. Hopper, Aung K. Win, C
eBioMedicine.2025; 114: 105661. CrossRef - Fusobacterium nucleatum Enrichment in Colorectal Tumor Tissue: Associations With Tumor Characteristics and Survival Outcomes
Amanda I. Phipps, Courtney M. Hill, Genevieve Lin, Rachel C. Malen, Adriana M. Reedy, Orsalem Kahsai, Hamza Ammar, Keith Curtis, Ningxin Ma, Timothy W. Randolph, Jing Ma, Shuji Ogino, Polly A. Newcomb, Meredith AJ. Hullar
Gastro Hep Advances.2025; 4(6): 100644. CrossRef - Enhancing fibroblast–epithelial cell communications: Serpine2 as a key molecule in Fusobacterium nucleatum–promoted colon cancer
Xueke Li, Simin Luo, Yifang Jiang, Qiong Ma, Fengming You, Qixuan Kuang, Xi Fu, Chuan Zheng
Frontiers in Immunology.2025;[Epub] CrossRef - The presence and relative abundance of salivary Fusobacterium nucleatum are not associated with colorectal cancer: a systematic review and meta-analysis
Ellay Gutmacher, Bálint Zsombor Sárai, Petrana Martineková, Szilvia Kiss-Dala, Gergely Agócs, Péter Hegyi, Andrea Bródy, Ákos Zsembery
Scientific Reports.2025;[Epub] CrossRef - The role of oral microbiota in digestive system diseases: current advances and perspectives
Yaqi Li, Yiping Xin, Wenlu Zong, Xiaoyu Li
Journal of Oral Microbiology.2025;[Epub] CrossRef - Fusobacterium Nucleatum in Colorectal Cancer: Relationship Among Immune Modulation, Potential Biomarkers and Therapeutic Implications
Dalila Incognito, Giuliana Ciappina, Claudia Gelsomino, Antonio Picone, Pierluigi Consolo, Alessandra Scano, Tindara Franchina, Nicola Maurea, Vincenzo Quagliariello, Salvatore Berretta, Alessandro Ottaiano, Massimiliano Berretta
International Journal of Molecular Sciences.2025; 26(19): 9710. CrossRef - Intratumoral presence of the genotoxic gut bacteria pks+ E. coli, Enterotoxigenic Bacteroides fragilis, and Fusobacterium nucleatum and their association with clinicopathological and molecular features of colorectal cancer
Jihoon E. Joo, Yen Lin Chu, Peter Georgeson, Romy Walker, Khalid Mahmood, Mark Clendenning, Aaron L. Meyers, Julia Como, Sharelle Joseland, Susan G. Preston, Natalie Diepenhorst, Julie Toner, Danielle J. Ingle, Norelle L. Sherry, Andrew Metz, Brigid M. Ly
British Journal of Cancer.2024; 130(5): 728. CrossRef - The role of Fusobacterium nucleatum in cancer and its implications for clinical applications
Wanyi Luo, Juxi Han, Xian Peng, Xuedong Zhou, Tao Gong, Xin Zheng
Molecular Oral Microbiology.2024; 39(6): 417. CrossRef -
Fusobacterium nucleatum Load Correlates with KRAS Mutation and Sessile Serrated Pathogenesis in Colorectal Adenocarcinoma
Koki Takeda, Minoru Koi, Yoshiki Okita, Sija Sajibu, Temitope O. Keku, John M. Carethers
Cancer Research Communications.2023; 3(9): 1940. CrossRef - Tumour Colonisation of Parvimonas micra Is Associated with Decreased Survival in Colorectal Cancer Patients
Thyra Löwenmark, Anna Löfgren-Burström, Carl Zingmark, Ingrid Ljuslinder, Michael Dahlberg, Sofia Edin, Richard Palmqvist
Cancers.2022; 14(23): 5937. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
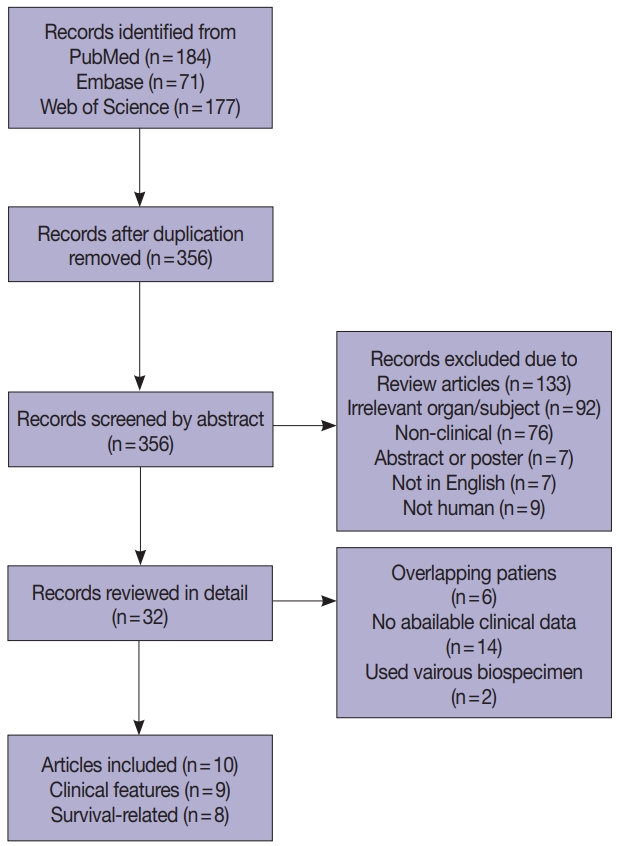
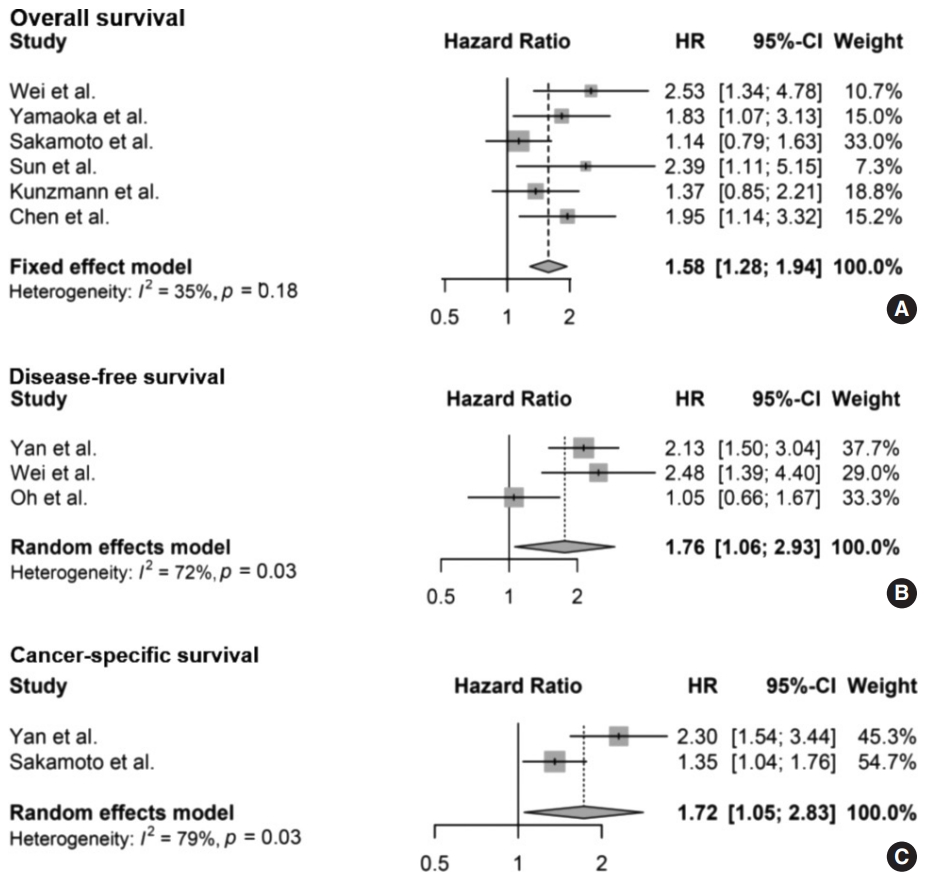
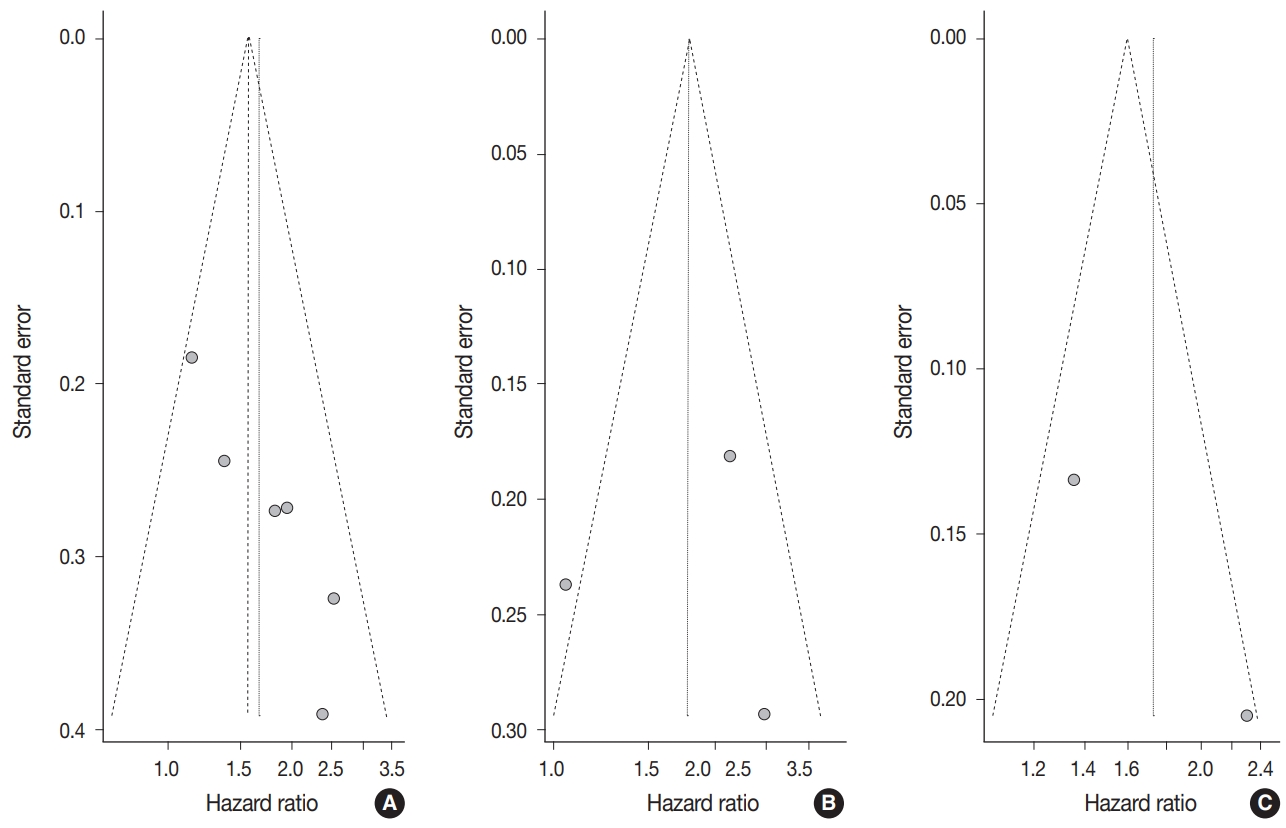
Fig. 1.
Fig. 2.
Fig. 3.
| Authors | Year | No. of patients | No. of positives | Biospecimen | Method | Survival | Clinical | Stage |
|---|---|---|---|---|---|---|---|---|
| Yan et al. [16] | 2017 | 280 | 187 | FFPE | qPCR | O | O | III/IV |
| Wei et al. [14] | 2016 | 180 | 90 | FF | qPCR | O | O | I–III |
| Yamaoka et al. [15] | 2018 | 100 | 50 | FF | ddPCR | O | O | I–IV |
| Samkamoto et al. [18] | 2018 | 1,069 | 67 | FFPE | qPCR | O | I–IV | |
| Sun et al. [22] | 2016 | 152 | 59 | FF | qPCR | O | O | I–IV |
| Kunzmann et al. [23] | 2019 | 190 | 61 | FF | qPCR | O | O | I–IV |
| Oh et al. [17] | 2019 | 593 | 204 | FFPE | qPCR | O | O | II/III |
| Chen et al. [24] | 2019 | 91 | 25 | FFPE | qPCR | O | O | II/III |
| Ito et al. [6] | 2015 | 511 | 143 | FFPE | qPCR | O | I–IV | |
| Mima et al. [25] | 2015 | 1,102 | 69 | FFPE | qPCR | O | I–IV |
| Survival | Specimen | Study | HR | 95% CI | p-value |
|---|---|---|---|---|---|
| Overall survival | FFPE | 2 | 1.43 | 0.85–2.42 | .168 |
| FF | 4 | 1.84 | 1.36–2.48 | < .001 | |
| Disease-free survival | FFPE | 2 | 1.52 | 0.76–3.04 | .214 |
| Clinical feature | Study | OR | 95% CI | p-value |
|---|---|---|---|---|
| Age | 5 | 0.993 | 0.787–1.254 | .954 |
| Sex (female/male) | 9 | 1.087 | 0.910–1.299 | .356 |
| Proximal/Distal colon | 4 | 1.642 | 1.252–2.153 | < .001 |
| Rectum/Colon | 5 | 1.184 | 0.899–1.560 | .229 |
| Tumor size | 2 | 0.897 | 0.592–1.360 | .609 |
| Differentiation | 7 | 1.547 | 1.166–2.052 | .002 |
| Vascular invasion | 2 | 0.627 | 0.337–1.167 | .141 |
| Perineural invasion (FFPE only) | 2 | 1.022 | 0.722–1.448 | .900 |
| pT category | 5 | 2.253 | 1.633–3.108 | < .001 |
| Lymph node metastasis | 5 | 1.308 | 0.979–1.748 | .069 |
| Distant metastasis | 4 | 1.912 | 1.196–3.058 | .007 |
| Stage | 6 | 1.110 | 0.885–1.394 | .367 |
| CIMP status (FFPE only) | 3 | 2.612 | 1.849–3.690 | < .001 |
| MLH1 hypermethylation | 3 | 2.852 | 1.987–4.095 | < .001 |
| MSI status | 5 | 2.982 | 2.175–4.086 | < .001 |
| KRAS mutation | 5 | 1.248 | 0.998–1.561 | .052 |
| BRAF mutation | 4 | 1.869 | 1.273–2.745 | .001 |
| PIK3CA mutation | 3 | 1.440 | 0.965–2.147 | .074 |
| Clinical feature | Specimen | Study | OR | 95% CI | p-value |
|---|---|---|---|---|---|
| Age | FFPE | 2 | 1.158 | 0.871–1.539 | .312 |
| FF | 3 | 0.720 | 0.478–1.086 | .117 | |
| Sex | FFPE | 5 | 1.096 | 0.891–1.349 | .386 |
| FF | 4 | 1.063 | 0.753–1.501 | .727 | |
| Proximal/Distal colon | FFPE | 2 | 1.676 | 1.220–2.303 | .001 |
| FF | 2 | 1.554 | 0.925–2.612 | .096 | |
| Rectum/Colon | FFPE | 2 | 1.259 | 0.841–1.884 | .263 |
| FF | 3 | 1.120 | 0.767–1.636 | .557 | |
| Differentiation | FFPE | 4 | 1.513 | 1.083–2.112 | .015 |
| FF | 3 | 1.632 | 0.958–2.778 | .071 | |
| pT category | FFPE | 2 | 2.187 | 1.437–3.329 | < .001 |
| FF | 3 | 2.354 | 1.428–3.880 | .001 | |
| Lymph node metastasis | FFPE | 2 | 0.832 | 0.549–1.262 | .387 |
| FF | 3 | 2.009 | 1.337–3.017 | .001 | |
| Distant metastasis | FF | 3 | 1.669 | 0.863–3.228 | .128 |
| Stage | FFPE | 2 | 0.917 | 0.673–1.251 | .586 |
| FF | 4 | 1.389 | 0.993–1.943 | .055 | |
| MLH1 hypermethylation | FFPE | 2 | 3.210 | 2.208–4.667 | < .001 |
| MSI status | FFPE | 4 | 2.757 | 1.987–3.824 | < .001 |
| KRAS mutation | FFPE | 3 | 1.125 | 0.881–1.436 | .345 |
| FF | 2 | 2.220 | 1.246–3.954 | .007 | |
| BRAF mutation | FFPE | 3 | 1.882 | 1.260–2.811 | .002 |
| PIK3CA mutation | FFPE | 2 | 1.464 | 0.960–2.233 | .076 |
FFPE, formalin-fixed paraffin-embedded; qPCR, quantitative polymerase chain reaction; FF, fresh frozen; ddPCR, droplet digital polymerase chain reaction.
HR, hazard ratio; CI, confidence interval; FFPE, formalin-fixed paraffin-embedded; FF, fresh frozen.
OR, odds ratio; CI, confidence interval; FFPE, formalin-fixed paraffin-embedded; CIMP, CpG island methylator phenotype; MSI, microsatellite instability.
OR, odds ratio; CI, confidence interval; FFPE, formalin-fixed paraffin-embedded; FF, fresh frozen; MSI, microsatellite instability.

 E-submission
E-submission