Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 49(2); 2015 > Article
-
Original Article
Overexpression of C-reactive Protein as a Poor Prognostic Marker of Resectable Hepatocellular Carcinomas - Jin Ho Shin, Chong Jai Kim, Eun Jeong Jeon, Chang Ohk Sung, Hwa Jeong Shin, Jene Choi, Eunsil Yu
-
Journal of Pathology and Translational Medicine 2015;49(2):105-111.
DOI: https://doi.org/10.4132/jptm.2015.01.19
Published online: March 12, 2015
Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
- Corresponding Author: Chong Jai Kim, M.D. Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, 88 Olympic-ro 43-gil, Songpa-gu, Seoul 138-736, Korea Tel: +82-2-3010-4516 Fax: +82-2-472-7898 E-mail: ckim@amc.seoul.kr
• Received: November 20, 2014 • Revised: January 9, 2015 • Accepted: January 19, 2015
© 2015 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted noncommercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background:
- C-reactive protein (CRP) is an acute phase reactant synthesized in the liver. CRP immunoreactivity is a feature of inflammatory hepatocellular adenomas with a higher risk of malignant transformation. A high serum CRP level denotes poor prognosis in hepatocellular carcinoma (HCC) patients. This study was conducted to determine whether CRP is produced in HCC and to assess the clinicopathologic significance of CRP expression in cancer cells.
-
Methods:
- CRP immunoreactivity was examined in treatment-naïve HCCs (n=224) using tissue microarrays and was correlated with clinicopathologic parameters. The expression of CRP mRNA and protein was also assessed in 12 HCC cases by quantitative real-time polymerase chain reaction and immunoblotting. Hep3B and SNU-449 HCC cell lines were used for the analysis of CRP mRNA regulation by interleukin 6 (IL-6).
-
Results:
- CRP was expressed in 133 of 224 HCCs (59.4%) with a variable degree of immunoreactivity (grade 1 in 25.9%; grade 2 in 20.1%; grade 3 in 13.4%). There was an inverse relationship between grade 3 CRP immunoreactivity and cancer-specific survival (p=.0047), while no associations were found with other parameters, including recurrence-free survival. The CRP mRNA expression level was significantly higher in CRP immunopositive cases than in immunonegative cases (p<.05). CRP mRNA expression was increased in Hep3B cells, but was not detected in SNU-449 cells even after IL-6 treatment.
-
Conclusions:
- We report the expression of CRP in HCC for the first time. CRP expression was associated with poor cancer-specific survival in patients with resectable HCC.
- Patients and tissue samples
- A total of 224 cases of treatment-naïve HCCs (n=224) were retrieved from the files of the Department of Pathology, Asan Medical Center, Seoul, Korea. All cases were surgically resectable (R0). Early recurrence was defined as a recurrence of the tumor within 2 years after surgery. All patients provided written informed consent, and this study was approved by the Institutional Review Board of Asan Medical Center, Seoul, Korea.
- Tissue microarray and immunohistochemistry
- Tissue microarrays were prepared using representative formalin-fixed, paraffin-embedded blocks of HCC cases. Two 2-mmthick tissue cores were obtained from the donor blocks and transferred onto the recipient blocks after reviewing hematoxylin and eosin–stained slides. Four-micrometer-thick tissue microarray sections were immunostained using a rabbit polyclonal antiCRP antibody (1:1,000, AbCam, Cambridge, UK). The sections were transferred onto silanized slides, and heat-induced epitope retrieval was performed by treating the slides with Cell Conditioning 1 buffer for 32 minutes in a BenchMark XT automatic immunostainer (Ventana Medical Systems, Tucson, AZ, USA). The signals were detected using the OptiView DAB IHC Detection Kit (Ventana Medical Systems). The immunoreactivity was evaluated by a pathologist (C.J.K.) blinded to clinical information, using a 4-tier grading system: negative, grade 0; positive in less than 10% of tumor cells, grade 1; positive in less than 50% of tumor cells, grade 2; diffusely positive, grade 3.
- Cell culture
- Human HCC cell lines, Hep3B and SNU-449, were used for the analysis of CRP mRNA regulation by IL-6. Hep3B cells were cultured in Dulbecco’s modified Eagle medium (Life Technologies, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. SNU-449 cells were cultured in RPMI 1640 medium (GE Healthcare Life Sciences, Pittsburgh, PA, USA) supplemented with 10% FBS and 1% penicillin/streptomycin. For IL-6 treatment, 5×105 cells were seeded into 100-mm dishes and were treated with 50 ng/mL of IL-6 (Cell Sciences, Canton, MA, USA) on the following day for 6 hours.
- Quantitative real-time polymerase chain reaction
- Total RNA was isolated from liver tissues (n=12) and cultured Hep3B and SNU-449 cells using Trizol and the miRNeasy Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The extracted RNA (1 µg) was reversetranscribed using the Reverse Transcription System (Promega, Madison, WI, USA). Quantitative real-time polymerase chain reaction (qRT-PCR) analyses of CRP mRNA expression were performed using a TaqMan Gene Expression Assay (Hs00357041_ m1, Applied Biosystems, Carlsbad, CA, USA) and an ABI PRISM 7000 Sequence Detection System (Applied Biosystems). The human RPLPO (Large Ribosomal Protein) was used as an endogenous control for normalization.
- Immunoblotting
- Immunoblotting was done in 12 cases. Liver tissues were pulverized in liquid nitrogen, and total protein lysates were obtained using RIPA lysis buffer. Thirty micrograms of protein were electrophoresed in 12% sodium dodecyl sulfate–polyacrylamide gel and transferred onto polyvinylidene fluoride membranes (GE Healthcare Life Sciences). The membranes were blocked with 5% bovine serum albumin in Tris-buffered saline with 0.1% Tween 20 (TBS-T) and incubated at 4˚C overnight with mouse monoclonal primary antibodies against CRP (1:1,000, AbCam, Cambridge, MA, USA) and vinculin (1:1,000; SigmaAldrich, St. Louis, MO, USA), respectively. The blots were subsequently incubated at room temperature with horseradish peroxidase–conjugated secondary antibody for 1 hour (Cell Signaling Technology, Danvers, MA, USA). The signals were detected using a SuperSignal West Dura Chemiluminescent Substrate (Thermo Scientific, Waltham, MA, USA).
- Statistical analyses
- The analyses of continuous variables and proportions were done using Pearson’s chi-square test and Fisher exact test. The survival analysis was done using the Kaplan-Meier method. Independent variables and groups were compared using the MannWhitney U test. SPSS ver. 18.0 software (SPSS Inc., Chicago, IL, USA) was used for all statistical analyses.
MATERIALS AND METHODS
- CRP expression in HCC
- The demographics of the study population are summarized in Table 1. CRP immunoreactivity was observed as diffuse cytoplasmic immunostaining. Overall, 59.4% (133/224) of HCC cases were immunopositive for CRP, with grades 1, 2, and 3 immunoreactivity in 25.9% (58/224), 20.1% (45/224), and 13.4% (30/224) of the cases, respectively (Fig. 1). When we analyzed the relationship between multiple clinicopathologic parameters and CRP immunoreactivity, we found a significant difference in cancer-specific survival between CRP-negative or non-diffuse immunopositive cases (grades 0, 1, and 2) and grade 3 immunopositive cases (Fig. 2A). Grade 3 CRP immunopositive cases showed significantly shorter survival compared to CRP-negative or non-diffuse immunopositive cases (median of 50.7 months and range of 6-58 months vs. median of 58.5 months and range of 5–88 years; p=.0047). However, no significant correlation was found between the degree of CRP immunoreactivity and other clinicopathologic parameters including recurrence-free survival (Fig. 2B).
- CRP mRNA expression in HCC
- We further analyzed CRP protein and mRNA expression in HCCs (n=12) to see whether CRP expression is regulated at the transcriptional level. Immunoblotting for CRP confirmed variable expression of CRP in HCC cases (Fig. 3A). CRP signals were readily detected in six cases, but not in the remaining six cases. qRT-PCR analysis demonstrated a higher CRP mRNA expression in CRP immunopositive cases than in immunonegative cases (p=.046; ΔCt median of 3.62, range of 2.8–5.98 vs ΔCt median of 8.56, range of 1.07–15) (Fig. 3B).
- After confirming the relationship between CRP protein and mRNA expression, we tested if CRP mRNA transcription was dependent on pro-inflammatory mediators. The treatment of Hep3B and SNU-449 cells with IL-6 at the concentration of 50 ng/mL for 6 hours induced a 17.5-fold increase in CRP mRNA expression in Hep3B cells, but CRP mRNA expression was not detected in SNU-449 cells even after IL-6 treatment (Fig. 4).
RESULTS
- The primary findings of this study are (1) CRP expression is relatively common in HCCs, with variable immunoreactivity in nearly 60% of the cases tested, (2) there is an inverse relationship between diffuse and strong CRP immunoreactivity and cancer-specific patient survival (p=.0047), (3) CRP-positive cases in immunoblotting showed significantly higher CRP mRNA expression, suggesting that increased CRP production is a consequence of transcriptional activation of CRP, and (4) CRP mRNA transcription is induced by IL-6 in Hep3B cells, but not in SNU- 449 cells, suggesting that CRP expression marks distinct molecular phenotypes among HCCs.
- Hepatocellular carcinogenesis represents a classic model of viral etiology associated with chronic inflammation [23]. The elevation of inflammatory biomarkers such as CRP, IL-6, C-peptide, and adiponectin is associated with a higher risk of HCC [24]. Several investigations have addressed the clinical significance of serum CRP. Serum CRP has been consistently shown to be a key component of inflammation-based prognostication of HCC. Mori et al. [25] proposed that a preoperative scoring system based on the preoperative serum concentration of CRP and alpha-fetoprotein has a prognostic value in patients with HCC after hepatectomy. Although the contribution of CRP produced by tumor cells to the elevation of serum CRP cannot be directly assessed, it is very probable that CRP of tumor origin is released into the systemic circulation. Although we expected certain differences in clinicopathologic characteristics between CRP immunopositive and immunonegative HCCs, a clear difference was found only in the cancer-specific survival of strong CRP immunopositive cases.
- One of the major potential drives for CRP overexpression is IL-6, and a role of IL-6 in hepatocellular carcinogenesis has been strongly suggested. When compared to healthy controls, cirrhotic patients and HCC patients had serum IL-6 concentrations that were 4-fold and 25-fold higher, respectively [26]. IL-6 signaling mediated via IL-6 receptors activates STAT3, mitogen-activated protein kinase, and phosphatidylinositol 3-kinase pathways. IL-6–mediated STAT3 activation is known to be a biological link between chronic inflammation and carcinogenesis [27], and IL-6 confers anti-apoptotic effects to cells. Blocking STAT3 activation using STAT3 siRNA or small molecular STAT3 inhibitor LLL 12 has been shown to abrogate the anti-apoptotic effects of IL-6 against doxorubicin-induced apoptosis in SNU- 449 HCC cells, which express a higher level of endogenous IL-6 compared to Hep3B cells [28]. We thought that CRP would be induced in both Hep3B and SNU-449 cells by IL-6 treatment. However, CRP mRNA expression after IL-6 treatment was significantly different between SNU-449 cells and Hep3B cells. Therefore, the observations in HCC tissue samples and the cell lines in vitro indicate that the degree of CRP expression involves several mechanisms, which need further elucidation.
- A major drawback of this study is that serum CRP or IL-6 levels could not be determined due to a lack of blood samples. Future analyses of the relationship between the CRP-positive HCC phenotype and systemic inflammatory profile will provide a more comprehensive understanding of the biology of HCC. We also could not determine the underlying biochemical mechanisms involved in the differential IL-6-induced responses between Hep3B and SNU-449 cells in terms of induction of CRP mRNA expression. Considering the complexities of IL-6 signaling and CRP transcription, more in vitro studies are necessary.
- In summary, we report the expression of CRP in HCC for the first time, and provide evidence to support the significance of CRP in HCC based on the clear difference in cancer-specific survival between CRP-positive and -negative cases. Overall findings strongly suggest that CRP is a marker for future molecular phenotyping of HCC.
DISCUSSION
Acknowledgments
Fig. 1.Cytoplasmic C-reactive protein (CRP) immunoreactivity in hepatocellular carcinoma cases. Immunoreactivity is analyzed using a 4-tier grading system: grade 0 (A), grade 1 (B), grade 2 (C), and grade 3 (D).
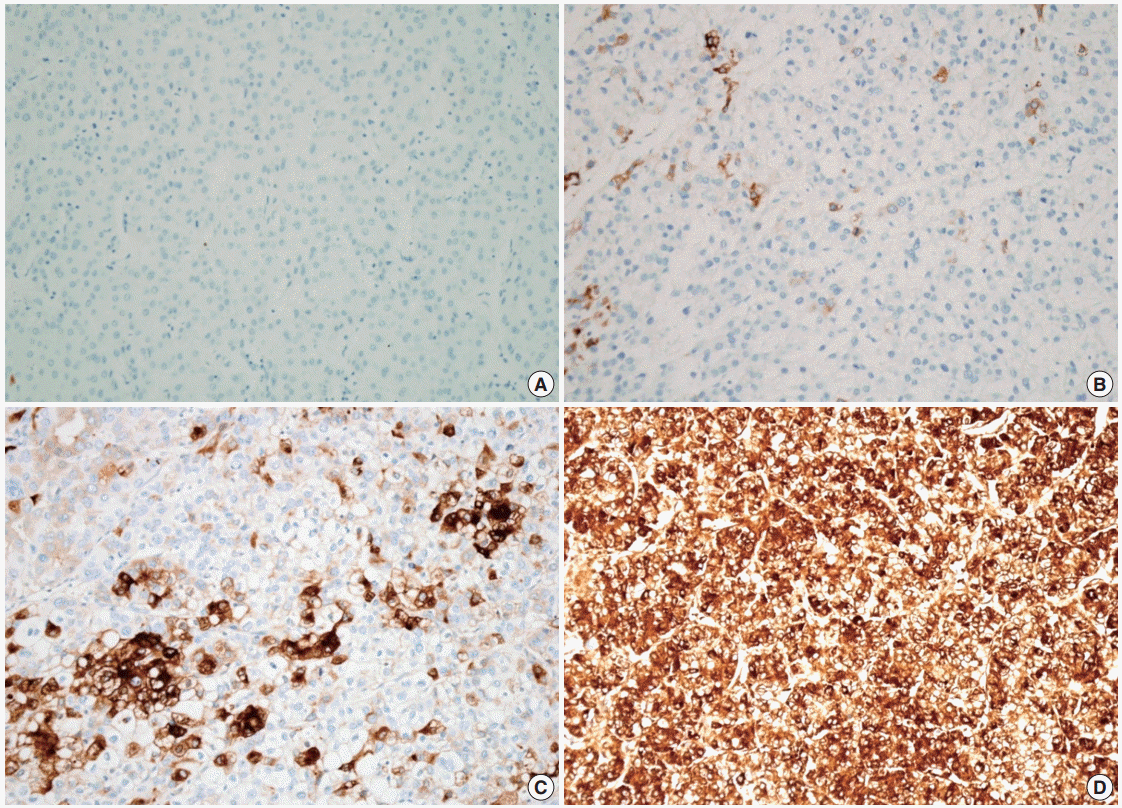

Fig. 2.Clinical significance of C-reactive protein (CRP) immunoreactivity. (A) There is a significant difference in cancer-specific survival between patients with CRP grade 3 hepatocellular carcinomas (HCCs) and those with CRP grade 0, 1, and 2 HCCs. (B) There is no difference in recurrence-free survival between the two groups.
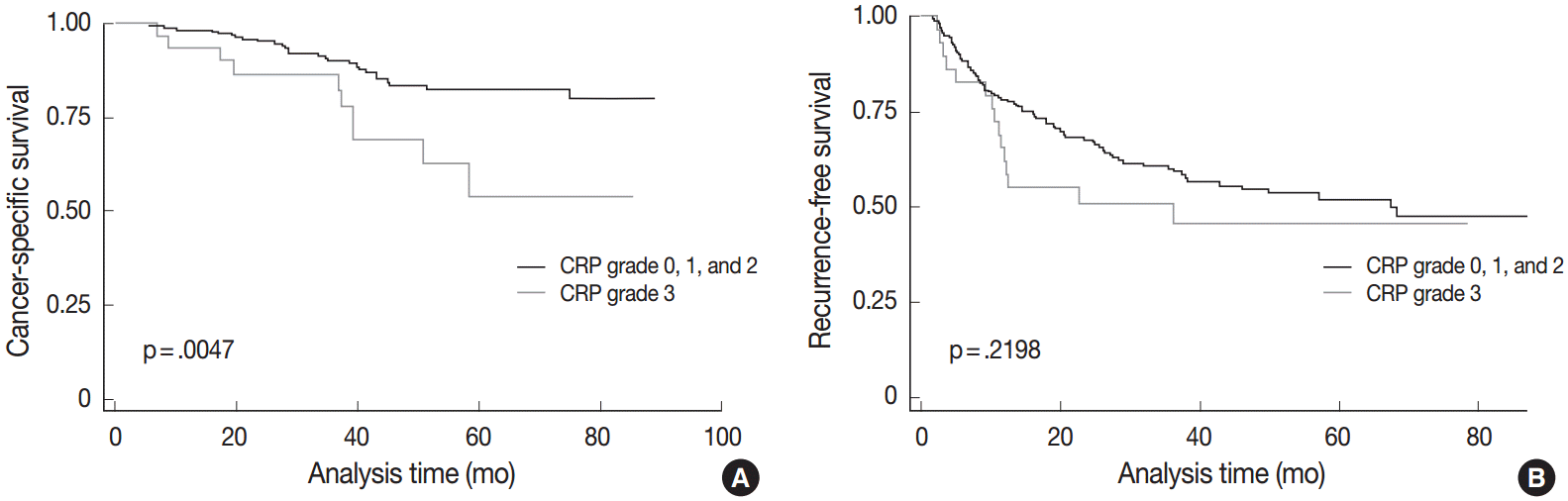

Fig. 3.Correlation between C-reactive protein (CRP) mRNA and protein expression in hepatocellular carcinomas (HCCs). (A) Immunoblotting for CRP protein shows variable expression in HCCs (N, non-neoplastic liver; T, HCC). CRP protein expression is not found in the tumor of case 4, while CRP bands are readily detectable in both non-neoplastic and HCC samples of cases 1, 2, and 3. (B) Quantitative real-time polymerase chain reaction results are shown in box plots of ΔCt for CRP mRNA expression (Ct_CRP–Ct_RPLPO).
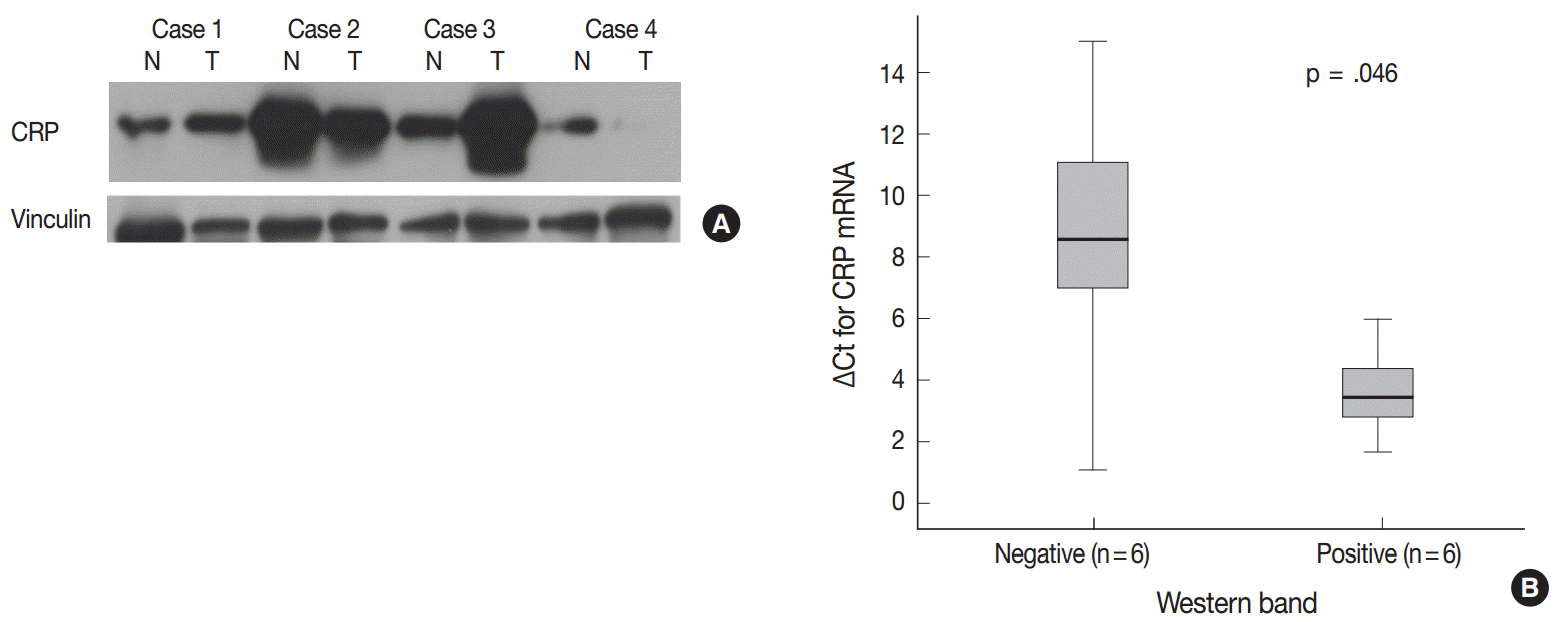

Fig. 4.Induction of C-reactive protein (CRP) mRNA in Hep3B and SNU-449 hepatocellular carcinoma cell lines after interleukin 6 (IL-6) treatment. (A) There are no significant changes in cellular morphology in either cell line after IL-6 treatment. (B) There is a 17.5-fold increase in CRP mRNA expression in IL-6–treated Hep3B cells (50 ng/mL for 6 hours), while CRP mRNA expression is not detected in SNU-449 cells. The yaxis represents fold-changes in CRP mRNA expression following IL-6 treatment.


Table 1.Clinical parameters and their relationship with CRP immunoreactivity
- 1. Lazzarotto C, Ronsoni MF, Fayad L, et al. Acute phase proteins for the diagnosis of bacterial infection and prediction of mortality in acute complications of cirrhosis. Ann Hepatol 2013; 12: 599-607. ArticlePubMed
- 2. Kushner I, Feldmann G. Control of the acute phase response: demonstration of C-reactive protein synthesis and secretion by hepatocytes during acute inflammation in the rabbit. J Exp Med 1978; 148: 466-77. ArticlePubMedPMCPDF
- 3. Volanakis JE, Kaplan MH. Specificity of C-reactive protein for choline phosphate residues of pneumococcal C-polysaccharide. Proc Soc Exp Biol Med 1971; 136: 612-4. ArticlePubMed
- 4. Castell JV, Gómez-Lechón MJ, David M, et al. Interleukin-6 is the major regulator of acute phase protein synthesis in adult human hepatocytes. FEBS Lett 1989; 242: 237-9. ArticlePubMedPDF
- 5. Castell JV, Gómez-Lechón MJ, David M, Fabra R, Trullenque R, Heinrich PC. Acute-phase response of human hepatocytes: regulation of acute-phase protein synthesis by interleukin-6. Hepatology 1990; 12: 1179-86. ArticlePubMed
- 6. Yoshizaki K. Pathogenic role of IL-6 combined with TNF-alpha or IL-1 in the induction of acute phase proteins SAA and CRP in chronic inflammatory diseases. Adv Exp Med Biol 2011; 691: 141-50. PubMed
- 7. de Man P, Jodal U, Svanborg C. Dependence among host response parameters used to diagnose urinary tract infection. J Infect Dis 1991; 163: 331-5. ArticlePubMed
- 8. Yin G, Hu G, Cang X, et al. C-reactive protein: rethinking its role in evaluating the severity of hyperlipidemic acute pancreatitis. Pancreas 2014; 43: 1323-8. PubMed
- 9. Lee CH, Chang JS, Syu SH, et al. IL-1beta promotes malignant transformation and tumor aggressiveness in oral cancer. J Cell Physiol 2015; 230: 875-84. PubMed
- 10. Sipos F, Fűri I, Constantinovits M, Tulassay Z, Műzes G. Contribution of TLR signaling to the pathogenesis of colitis-associated cancer in inflammatory bowel disease. World J Gastroenterol 2014; 20: 12713-21. ArticlePubMedPMC
- 11. Saxena M, Yeretssian G. NOD-like receptors: master regulators of inflammation and cancer. Front Immunol 2014; 5: 327.ArticlePubMedPMC
- 12. Castaño-Rodríguez N, Kaakoush NO, Mitchell HM. Pattern-recognition receptors and gastric cancer. Front Immunol 2014; 5: 336.PubMedPMC
- 13. Szkandera J, Stotz M, Absenger G, et al. Validation of C-reactive protein levels as a prognostic indicator for survival in a large cohort of pancreatic cancer patients. Br J Cancer 2014; 110: 183-8. ArticlePubMedPMCPDF
- 14. Yi JH, Wang D, Li ZY, Hu J, Niu XF, Liu XL. C-reactive protein as a prognostic factor for human osteosarcoma: a meta-analysis and literature review. PLoS One 2014; 9: e94632.ArticlePubMedPMC
- 15. Zheng Z, Zhou L, Gao S, Yang Z, Yao J, Zheng S. Prognostic role of C-reactive protein in hepatocellular carcinoma: a systematic review and meta-analysis. Int J Med Sci 2013; 10: 653-64. ArticlePubMedPMC
- 16. Nishikawa H, Arimoto A, Wakasa T, Kita R, Kimura T, Osaki Y. Pretreatment C-reactive protein as a prognostic factor for recurrence after surgical resection of hepatocellular carcinoma. Anticancer Res 2013; 33: 1181-8. PubMed
- 17. Bioulac-Sage P, Rebouissou S, Thomas C, et al. Hepatocellular adenoma subtype classification using molecular markers and immunohistochemistry. Hepatology 2007; 46: 740-8. ArticlePubMed
- 18. Han J, van den Heuvel MC, Kusano H, de Jong KP, Gouw AS. How normal is the liver in which the inflammatory type hepatocellular adenoma develops? Int J Hepatol 2012; 2012: 805621.ArticlePubMedPMCPDF
- 19. Bioulac-Sage P, Laumonier H, Cubel G, Rossi JZ, Balabaud C. Hepatic resection for inflammatory hepatocellular adenomas: pathological identification of micronodules expressing inflammatory proteins. Liver Int 2010; 30: 149-54. ArticlePubMed
- 20. Fonseca S, Hoton D, Dardenne S, et al. Histological and immunohistochemical revision of hepatocellular adenomas: a learning experience. Int J Hepatol 2013; 2013: 398308.ArticlePubMedPMCPDF
- 21. Sempoux C, Chang C, Gouw A, et al. Benign hepatocellular nodules: what have we learned using the patho-molecular classification. Clin Res Hepatol Gastroenterol 2013; 37: 322-7. ArticlePubMed
- 22. Kakar S, Grenert JP, Paradis V, Pote N, Jakate S, Ferrell LD. Hepatocellular carcinoma arising in adenoma: similar immunohistochemical and cytogenetic features in adenoma and hepatocellular carcinoma portions of the tumor. Mod Pathol 2014; 27: 1499-509. ArticlePubMedPMCPDF
- 23. Tarocchi M, Polvani S, Marroncini G, Galli A. Molecular mechanism of hepatitis B virus-induced hepatocarcinogenesis. World J Gastroenterol 2014; 20: 11630-40. ArticlePubMedPMC
- 24. Aleksandrova K, Boeing H, Nöthlings U, et al. Inflammatory and metabolic biomarkers and risk of liver and biliary tract cancer. Hepatology 2014; 60: 858-71. PubMedPMC
- 25. Mori S, Kita J, Kato M, Shimoda M, Kubota K. Usefulness of a new inflammation-based scoring system for prognostication of patients with hepatocellular carcinoma after hepatectomy. Am J Surg 2015; 209: 187-93. ArticlePubMed
- 26. Porta C, De Amici M, Quaglini S, et al. Circulating interleukin-6 as a tumor marker for hepatocellular carcinoma. Ann Oncol 2008; 19: 353-8. ArticlePubMed
- 27. Hodge DR, Hurt EM, Farrar WL. The role of IL-6 and STAT3 in inflammation and cancer. Eur J Cancer 2005; 41: 2502-12. ArticlePubMed
- 28. Liu Y, Li PK, Li C, Lin J. Inhibition of STAT3 signaling blocks the anti-apoptotic activity of IL-6 in human liver cancer cells. J Biol Chem 2010; 285: 27429-39. ArticlePubMedPMC
REFERENCES
Figure & Data
References
Citations
Citations to this article as recorded by 

- Influencing Factors for Postembolization Fever in Patients Undergoing Transarterial Chemoembolization Based on Machine Learning
Won-Du Chang, Myoung Soo Kim
CIN: Computers, Informatics, Nursing.2026;[Epub] CrossRef - Diagnostic and prognostic relevance of inflammatory markers in surgically treated thymic epithelial tumors: An international multicenter study
Evelyn Megyesfalvi, Aron Ghimessy, Jonas Bauer, Orsolya Pipek, Kevin Saghi, Aron Gellert, Janos Fillinger, Ozlem Okumus, Vivien Teglas, Erna Ganofszky, Krisztina Bogos, Ferenc Renyi-Vamos, Zsolt Megyesfalvi, Clemens Aigner, Balazs Hegedus, Balazs Dome, Be
Lung Cancer.2025; 200: 108111. CrossRef - The Role of IL‐6, IL‐10 and CRP in Gastrointestinal Cancers
Reyhaneh Yarmohammadi, Kazem Najafi, Mina Noroozbeygi, Kimia Didehvar, Aria Rastin, Faezeh Ataei, Mohammad Reza Atashzar, Sahar Shomeil Shushtari
Cell Biology International.2025; 49(9): 1061. CrossRef - Peritumoral portal enhancement during transarterial chemoembolization: a potential prognostic factor for patients with hepatocellular carcinoma
Sofi Sennefelt Nyman, Angeliki Dimopoulou Creusen, Ulf Johnsson, Fredrik Rorsman, Johan Vessby, Charlotte Ebeling Barbier
Acta Radiologica.2022; 63(10): 1323. CrossRef - The quest for precision oncology with immune checkpoint inhibitors for hepatocellular carcinoma
Giuseppe Cabibbo, Amit G. Singal
Journal of Hepatology.2022; 76(2): 262. CrossRef - HNF-1β is a More Sensitive and Specific Marker Than C-Reactive Protein for Identifying Biliary Differentiation in Primary Hepatic Carcinomas
Pallavi A. Patil, Tamar Taddei, Dhanpat Jain, Xuchen Zhang
Archives of Pathology & Laboratory Medicine.2022; 146(2): 220. CrossRef - Malignant transformation of hepatocellular adenoma
Céline Julien, Brigitte Le Bail, Laurence Chiche, Charles Balabaud, Paulette Bioulac-Sage
JHEP Reports.2022; 4(3): 100430. CrossRef - Effect of Treatment with Colchicine after Acute Coronary Syndrome on Major Cardiovascular Events: A Systematic Review and Meta-Analysis of Clinical Trials
Erfan Razavi, Akam Ramezani, Asma Kazemi, Armin Attar, Baohui Xu
Cardiovascular Therapeutics.2022; 2022: 1. CrossRef - Steatotic and Steatohepatitic Hepatocellular Carcinomas
Umut Aykutlu, Asuman Argon, Mehmet Orman, Sezgin Ulukaya, Murat Zeytunlu, Zeki Karasu, Fulya Günşar, Deniz Nart, Ulus Akarca, Funda Yilmaz
American Journal of Surgical Pathology.2021; 45(9): 1252. CrossRef - Cytochrome P450 4A11 expression in tumor cells: A favorable prognostic factor for hepatocellular carcinoma patients
Hyuk Soo Eun, Sang Yeon Cho, Byung Seok Lee, Sup Kim, In‐Sang Song, Kwangsik Chun, Cheong‐Hae Oh, Min‐Kyung Yeo, Seok Hyun Kim, Kyung‐Hee Kim
Journal of Gastroenterology and Hepatology.2019; 34(1): 224. CrossRef - Investigating Trk Protein Expression between Oropharyngeal and Non-oropharyngeal Squamous Cell Carcinoma: Clinical Implications and Possible Roles of Human Papillomavirus Infection
Yoon Ah Cho, Ji Myung Chung, Hyunmi Ryu, Eun Kyung Kim, Byoung Chul Cho, Sun Och Yoon
Cancer Research and Treatment.2019; 51(3): 1052. CrossRef - Increased systemic zonula occludens 1 associated with inflammation and independent biomarker in patients with hepatocellular carcinoma
Amit Kumar Ram, Biju Pottakat, Balasubramaniyan Vairappan
BMC Cancer.2018;[Epub] CrossRef - C-reactive Protein Overexpression in the Background Liver of Hepatitis B Virus–Associated Hepatocellular Carcinoma Is a Prognostic Biomarker
Jin Ho Shin, Eunsil Yu, Eun Na Kim, Chong Jai Kim
Journal of Pathology and Translational Medicine.2018; 52(5): 267. CrossRef - Efficacy of vasopressin V2 receptor antagonist tolvaptan in treatment of hepatic edema
Yasunari Hiramine, Hirofumi Uto, Yasushi Imamura, Takuya Hiwaki, Takeshi Kure, Sho Ijuin, Kohei Oda, Seiichi Mawatari, Kotaro Kumagai, Koki Tokunaga, Hirofumi Higashi, Ichiro Kanetsuki, Osamu Kubozono, Shigeho Maenohara, Akio Ido
Hepatology Research.2017; 47(6): 542. CrossRef - Elevated CRP levels predict poor outcome and tumor recurrence in patients with thymic epithelial tumors: A pro- and retrospective analysis
Stefan Janik, Christine Bekos, Philipp Hacker, Thomas Raunegger, Bahil Ghanim, Elisa Einwallner, Lucian Beer, Walter Klepetko, Leonhard Müllauer, Hendrik J. Ankersmit, Bernhard Moser
Oncotarget.2017; 8(29): 47090. CrossRef - Pretreatment neutrophil-to-lymphocyte ratio predicts prognosis in patients with metastatic renal cell carcinoma receiving targeted therapy
Gui-Ming Zhang, Yao Zhu, Wei-Jie Gu, Hai-Liang Zhang, Guo-Hai Shi, Ding-Wei Ye
International Journal of Clinical Oncology.2016; 21(2): 373. CrossRef - Current Proceedings in the Molecular Dissection of Hepatocellular Adenomas: Review and Hands-on Guide for Diagnosis
Diane Goltz, Hans-Peter Fischer
International Journal of Molecular Sciences.2015; 16(9): 20994. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
Overexpression of C-reactive Protein as a Poor Prognostic Marker of Resectable Hepatocellular Carcinomas
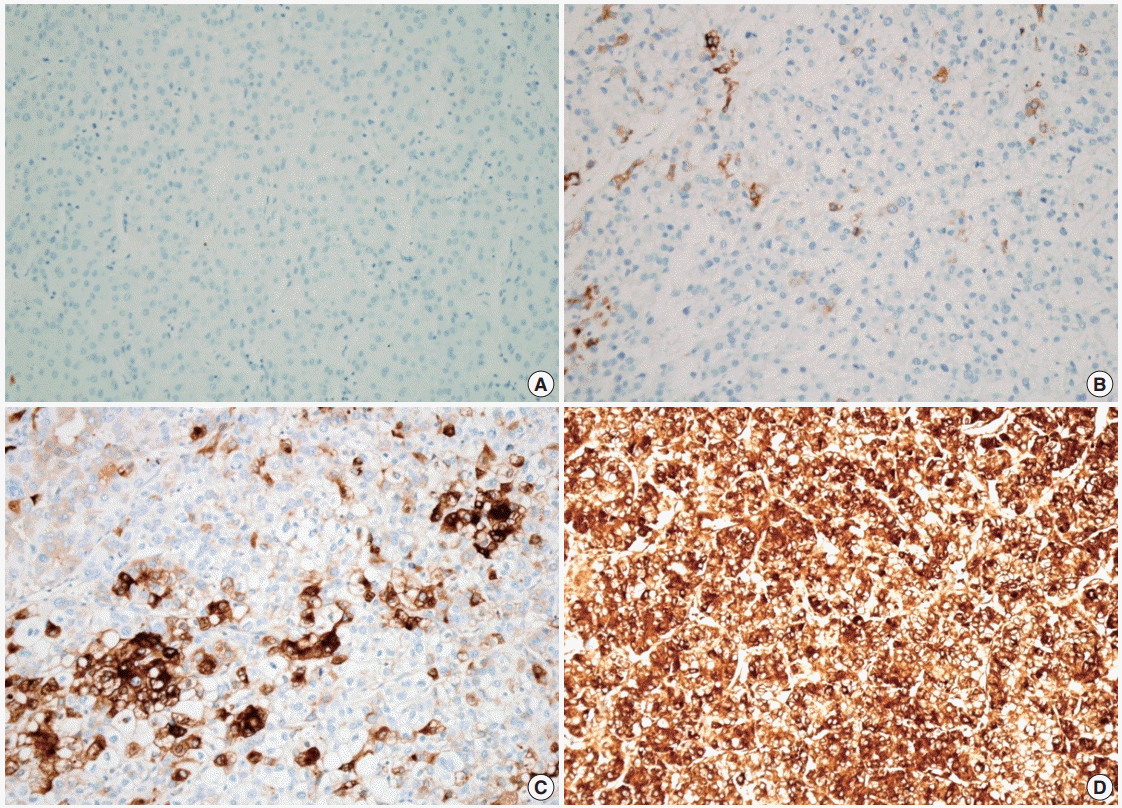
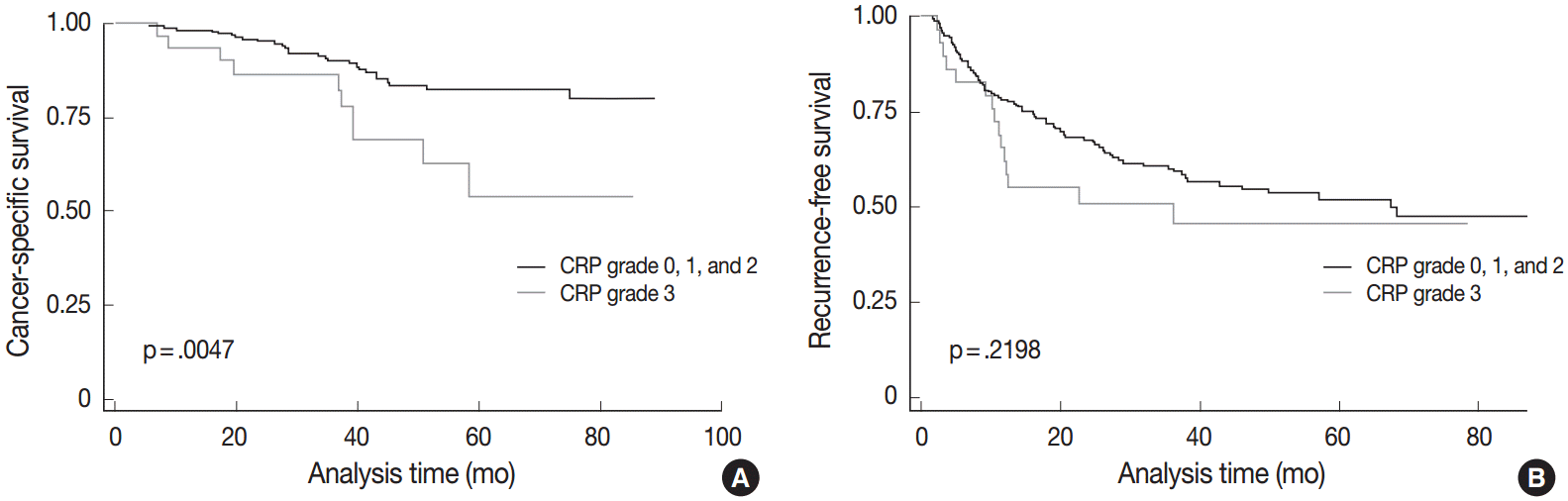
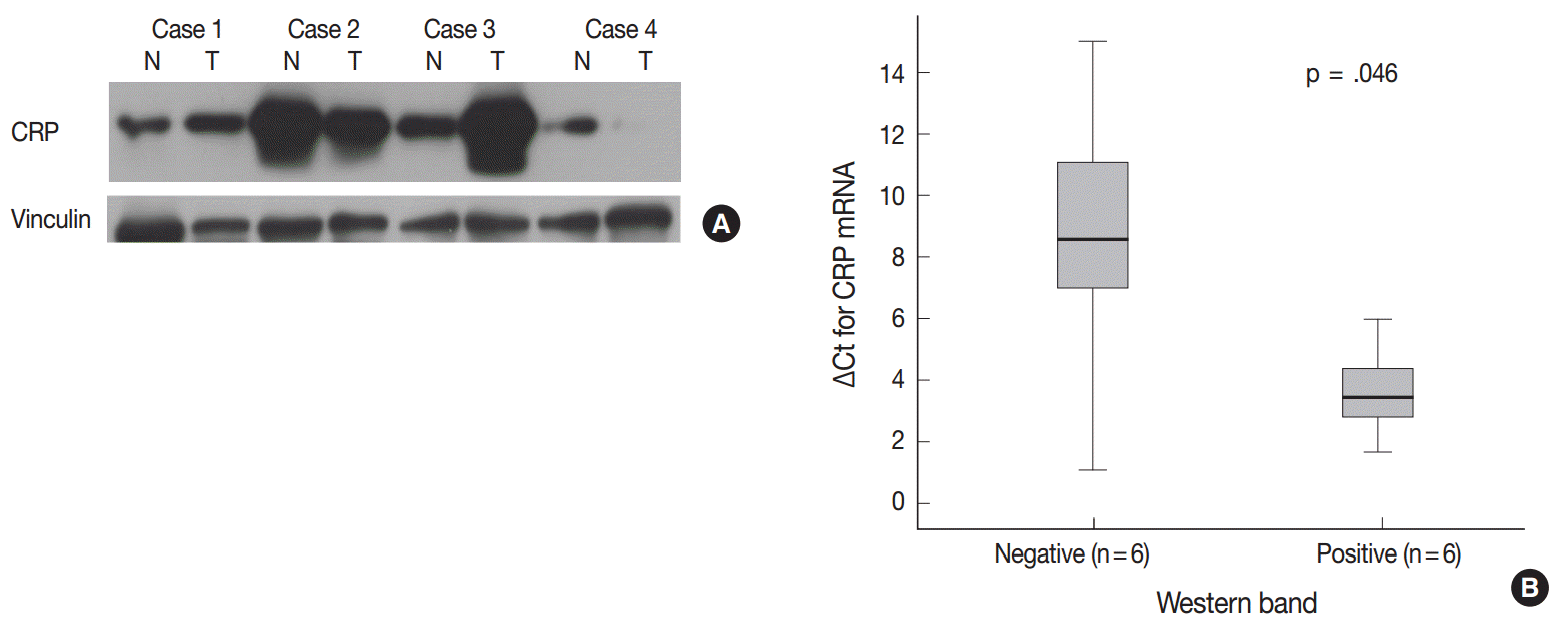

Fig. 1. Cytoplasmic C-reactive protein (CRP) immunoreactivity in hepatocellular carcinoma cases. Immunoreactivity is analyzed using a 4-tier grading system: grade 0 (A), grade 1 (B), grade 2 (C), and grade 3 (D).
Fig. 2. Clinical significance of C-reactive protein (CRP) immunoreactivity. (A) There is a significant difference in cancer-specific survival between patients with CRP grade 3 hepatocellular carcinomas (HCCs) and those with CRP grade 0, 1, and 2 HCCs. (B) There is no difference in recurrence-free survival between the two groups.
Fig. 3. Correlation between C-reactive protein (CRP) mRNA and protein expression in hepatocellular carcinomas (HCCs). (A) Immunoblotting for CRP protein shows variable expression in HCCs (N, non-neoplastic liver; T, HCC). CRP protein expression is not found in the tumor of case 4, while CRP bands are readily detectable in both non-neoplastic and HCC samples of cases 1, 2, and 3. (B) Quantitative real-time polymerase chain reaction results are shown in box plots of ΔCt for CRP mRNA expression (Ct_CRP–Ct_RPLPO).
Fig. 4. Induction of C-reactive protein (CRP) mRNA in Hep3B and SNU-449 hepatocellular carcinoma cell lines after interleukin 6 (IL-6) treatment. (A) There are no significant changes in cellular morphology in either cell line after IL-6 treatment. (B) There is a 17.5-fold increase in CRP mRNA expression in IL-6–treated Hep3B cells (50 ng/mL for 6 hours), while CRP mRNA expression is not detected in SNU-449 cells. The yaxis represents fold-changes in CRP mRNA expression following IL-6 treatment.
Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Overexpression of C-reactive Protein as a Poor Prognostic Marker of Resectable Hepatocellular Carcinomas
| Clinical parameter | CRP grade 0, 1, 2 | CRP grade 3 | p-value |
|---|---|---|---|
| Gender | .019 | ||
| Male | 150 (67.0) | 20 (8.9) | |
| Female | 44 (19.6) | 10 (4.5) | |
| Age (yr) | .054 | ||
| < 60 | 123 (54.9) | 22 (9.8) | |
| ≥ 60 | 71 (31.7) | 8 (3.6) | |
| Tumor size (cm) | .110 | ||
| < 5 | 138 (61.6) | 19 (8.5) | |
| ≥ 5 | 56 (25.0) | 11 (4.9) | |
| Serum AFP (ng/mL) | .125 | ||
| < 400 | 141 (62.9) | 24 (10.7) | |
| ≥ 400 | 53 (23.6) | 6 (2.7) | |
| BCLC stage | < .001 | ||
| A | 185 (82.6) | 26 (11.6) | |
| B | 9 (4.0) | 4 (1.8) | |
| Etiology | .096 | ||
| HBV | 139 (62.1) | 23 (10.3) | |
| HCV | 20 (8.9) | 1 (0.4) | |
| NBNC | 35 (15.6) | 6 (2.7) | |
| Fibrosis stage (Batts-Ludwig) | .454 | ||
| Stage 1, 2 | 39 (17.4) | 7 (3.1) | |
| Stage 3, 4 | 155 (69.2) | 23 (10.3) | |
| Microvascular invasion | .983 | ||
| Not identified | 136 (60.7) | 21 (9.4) | |
| Present | 58 (25.9) | 9 (4.0) | |
| Tumor number | < .001 | ||
| <3 | 185 (82.6) | 26 (11.6) | |
| ≥3 | 9 (4.0) | 4 (1.8) | |
| Edmondson-Steiner grade (worst) | .814 | ||
| Grade 1, 2 | 67 (29.9) | 10 (4.5) | |
| Grade 3, 4 | 127 (56.7) | 20 (8.9) | |
| Edmondson-Steiner grade (most) | .241 | ||
| Grade 1, 2 | 128 (57.1) | 18 (8.0) | |
| Grade 3, 4 | 66 (29.5) | 12 (5.4) | |
| Capsular invasion | .492 | ||
| Absent | 156 (69.6) | 25 (11.2) | |
| Present | 38 (17.0) | 5 (2.2) | |
| Early recurrence | .098 | ||
| Absent | 114 (50.9) | 15 (6.7) | |
| Occurs | 80 (35.7) | 15 (6.7) |
Table 1. Clinical parameters and their relationship with CRP immunoreactivity
Values are presented as number (%). CRP, C-reactive protein; AFP, alpha-fetoprotein; BCLC, Barcelona Clinic Liver Cancer; HBV, hepatitis B virus; HCB, hepatitis C virus; NBNC, non-B, non-C hepatocellular carcinoma.

 E-submission
E-submission






