Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 48(3); 2014 > Article
-
Original Article
Evaluation of Protein Expression in Housekeeping Genes across Multiple Tissues in Rats - Hye Jeong Kim, Jong In Na, Byung Woo Min, Joo Young Na, Kyung Hwa Lee1, Jae Hyuk Lee1, Young Jik Lee, Hyung Seok Kim, Jong Tae Park
-
Korean Journal of Pathology 2014;48(3):193-200.
DOI: https://doi.org/10.4132/KoreanJPathol.2014.48.3.193
Published online: June 26, 2014
Department of Forensic Medicine, Chonnam National University Medical School, Gwangju, Korea.
1Department of Pathology, Chonnam National University Medical School, Gwangju, Korea.
- Corresponding Author: Jong Tae Park, M.D. Department of Forensic Medicine, Chonnam National University Medical School, 160 Baekseo-ro, Dong-gu, Gwangju 501-746, Korea. Tel: +82-62-220-4090, Fax: +82-62-223-4250, jtpark@jnu.ac.kr
• Received: January 23, 2014 • Revised: March 28, 2014 • Accepted: April 21, 2014
© 2014 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- Housekeeping genes, which show constant protein expression patterns between different tissue types, are very important in molecular biological studies as an internal control for protein research.
-
Methods
- The protein expression profiles of seven housekeeping genes (HPRT1, PPIA, GYS1, TBP, YWHAZ, GAPDH and ACTB) in various rat tissues (cerebrum, cerebellum, cardiac ventricle and atrium, psoas muscle, femoral muscle, liver, spleen, kidney, and aorta) were analyzed by Western blot and compared by coefficient of variation (CV).
-
Results
- HPRT1 was stably expressed (CV≤10%) in six tissues (cerebrum, cerebellum, ventricle, femoral muscle, spleen, and kidney), PPIA was stably expressed in five tissues (cerebrum, cerebellum, ventricle, spleen and kidney), YWHAZ was stably expressed in three tissues (cerebrum, cerebellum, and kidney), and GAPDH was stably expressed in four tissues (cerebrum, ventricle, psoas muscle, and kidney). In comparison, GYS1, TBP, and ACTB were found to have CV values over 10% in all tissues. Of the seven genes examined, four (HPRT1, PPIA, YWHAZ, and GAPDH) were found to be stably expressed across multiple organs, with low CV values (≤10%).
-
Conclusions
- These results will provide fundamental information regarding internal controls for protein expression studies and can be used for analysis of postmortem protein degradation patterns in forensic medicine.
- Animals
- A total of eight, eight-week-old male Sprague-Dawley rats were used in this study. The rats were sacrificed by decapitation under anesthesia. Then the tissues from the cerebrum, cerebellum, left ventricular free wall, whole atrial tissue, psoas muscle, femoral muscle, liver, spleen, kidney, and aorta were immediately collected and divided into several small pieces. Tissue samples were put into 1.5 mL tubes and stored at -80℃. This study was approved by the institutional Animal Ethics Committee of Chonnam National University Medical School.
- Protein extraction and Western blot analysis
- PRO-PREP Protein Extraction Solution (iNtRON Biotechnology, Seongnam, Korea) was added to the 1.5 mL tubes containing frozen samples according to the manufacturer's protocol. The tissues were chopped broken into small pieces using microscissors on ice for 30 minutes. The samples were centrifuged at 14,000×g and protein-containing supernatant was moved into a new tube. The protein concentration was measured using a bicinchoninic acid protein assay (Thermo Scientific, Rockford, IL, USA). Twenty microliters samples containing 10 µg total protein were separated by sodium dodecyl sulphate-polyacrylamide gel electrophoresis and subsequently transferred to a nitrocellulose membrane (PALL, Port Washington, NY, USA) by electrophoresis.
- After treatment with 5% nonfat milk in Tris-buffered saline and Tween 20 (TBST) for one hour at room temperature, the membranes were incubated with primary antibodies (Table 1) overnight at 4℃. The membranes were then washed three times in TBST and incubated with horseradish peroxidase-conjugated secondary antibodies (1:10,000) for one hour. Finally, the signals were developed using an enhanced chemiluminescence kit (Millipore, Billerica, MA, USA) and analyzed using a LAS3000 (Fuji Film, Tokyo, Japan).
- Quantitation and comparison of signal intensity
- Points of interest (POIs) from Western blot bands were marked and quantitated using FUJI FILM MultiGauge V3.0 software (Fjji Film), and the background level was subtracted from the POI level. The data were presented as mean±standard deviation (SD) values. In addition, coefficient of variation (CV) was used for comparing the expression stability of genes in various organs. CV is defined as the SD divided by the mean, multiplied by 100%.2,5 The expression stability was evaluated based on the assumption that a lower CV value means lower variations of POI expression levels within each group. CV values equal to or less than 10% were used for selecting stable genes in a previous study,6 the same criterion was adopted in this study.
MATERIALS AND METHODS
- All seven housekeeping genes used in this study were expressed in all ten different tissues (Fig. 1). The signal intensities of POI measured by MultiGauge V3.0 are shown in Table 2. The CV value was calculated using the mean and SD, and the result was rounded off to two decimal places (Table 3). The expression stability of each housekeeping gene was represented using CV values across tissue types (Fig. 2). The unit of the vertical axis for CV values was expressed on a logarithmic scale to represent 10% or lower differences in each gene effectively. Among the ten organs, the cerebrum, cerebellum, ventricle, femoral muscle spleen, and kidney were shown to have stable HPRT1 expression by low CV values (≤10%); the cerebrum, cerebellum, ventricle, kidney, and spleen showed stable PPIA expression; the cerebrum, cerebellum, and kidney showed stable YWHAZ expression; and the cerebrum, ventricle, psoas muscle, and kidney, showed stable GAPDH expression. All organ tissues were found to have CV values over 10% for GYS1, TBP, and ACTB. Of the various organs examined, the femoral muscle showed the lowest CV value with GYS1. Similarly, the cerebrum showed the lowest CV value with TBP, while the kidney showed the lowest variation with ACTB.
- The expression stability of POI in each tissue was also evaluated using CV values, with the horizontal axis representing each gene and the vertical axis for the CV value on a logarithmic scale (Fig. 3). Among the seven genes, HPRT1, PPIA, YWHAZ, and GAPDH were shown to be stably expressed in the cerebrum with low CV values (≤10%); HPRT1, PPIA, and YWHAZ were stably expressed in the cerebellum; HPRT1, PPIA, and GAPDH, were stably expressed in the ventricle; GAPDH, was stably expressed in psoas muscle; HPRT1, was stably expressed in femoral muscle; HPRT1 and PPIA were stably expressed in the spleen; and HPRT1, PPIA, YWHAZ, and GAPDH, were stably expressed in the kidney. In the atrium, liver, and aorta, all genes were found to have CV values over 10%. Of the seven genes, PPIA showed the lowest CV value in the atrium while ACTB had the lowest CV values in the liver and aorta.
- In summary, HPRT1 in the cerebrum, cerebellum, ventricle, femoral muscle, spleen, and kidney, PPIA in the cerebrum, cerebellum, ventricle, spleen, and kidney, YWHAZ in the cerebrum, cerebellum, and kidney, and GAPDH in the cerebrum, ventricle, psoas muscle, and kidney showed 10% or lower CV values. Of the ten organs used in the study, the atrium, liver, and aorta showed high CV values greater than 10% with all housekeeping genes. Therefore, these three organs appear to be less useful tissues for studies based on constant gene expression.
RESULTS
- In protein expression studies, it is important to identify housekeeping genes that show comparatively stable expression patterns across body tissues. Expression of certain housekeeping genes can vary widely depending on different tissue types and disease processes involved. Suitable housekeeping genes as an internal control under certain defined conditions have been reported in previous studies. Rho et al.7 have reported that RPL29 only or the combination of RPL29 and B2M was most appropriate for stomach tissues; B2M only or the combination of B2M and GAPDH for stomach cancer cell lines;7 ACTB and SDHA, for the gene expression analysis of breast cancer among typical reference genes;8 and GAPDH, RPS17, RPL30, RPLP0, and TFRC, in studies on post-mortem brain samples from patients with major psychiatric disorders.9 According to gene expression studies of mice myocardial infarction models, HPRT, RPL13A, and TPT1 are the appropriate reference genes;10 GUSB, PPIA, and TBP, are used for serous ovarian cancer;11 and EEF1A1 only or the combination of EEFA1 and GAPDH, are commonly used for cervical cancer.12 In swine skeletal muscles, PPIA, HPRT, and eEF-1γ are the genes that showed the most stable expression.13 In the rat hippocampus after resuscitation from a cardiac arrest, a combination of CypA and Pgk1 should be considered at four and 21 days after injury, whereas CypA and Gapdh is the best combination at seven days.14 CrypA is most favorable if restriction to a single reference gene is required for all time points.14 These stable housekeeping genes were identified based on mRNA expression analyses. However, in the present study, we identified the stable housekeeping genes based on protein expression.
- The expression stabilities of seven housekeeping genes (HPRT1, PPIA, GYS1, TBP, YWHAZ, GAPDH, and ACTB) in the rat cerebrum, cerebellum, cardiac ventricle and atrium, psoas major muscle, femoral muscle, liver, spleen, kidney, and aorta tissues were evaluated through protein quantitation, Western blotting, and Multi-gauge software (Fuji Film). The results revealed that four genes-HPRT1, PPIA, YWHAZ, and GAPDH-showed 10% or lower CV values, which implied that they are more stably expressed across tissues compared to other housekeeping genes. The CV is one of the most used statistical measures by researchers in the evaluation of experimental precision. In field experiments, if the CV is below 10%, it is considered low. CV values from 10% to 20% are considered moderate, implying good precision, while those from 20% to 30% are considered high, indicating low precision. Finally, CV values above 30% are considered very high, indicating very low precision.15,16 Listed in order of stability, HPRT1 is a stably expressed gene in the spleen, kidney, cerebrum, cerebellum, ventricle, and femoral muscle; PPIA is stably expressed in the cerebellum, kidney, spleen, cerebrum, and ventricle; YWHAZ, is stably expressed in the cerebellum, kidney, and cerebrum; and GAPDH is stably expressed in the cerebrum, kidney, ventricle, and psoas muscle. In particular, the gene that showed the most stable expression was PPIA in the cerebellum. For GYS1, TBP, and ACTB, all organ tissues were found to have CV values over 10%. Furthermore, among the ten selected organs, the atrium, liver, and aorta were considered as less desirable tissues for housekeeping gene expression studies because of unstable expression with CV values over 10%. In studies on muscle samples, different results have been reported. Gu et al.3 reported TOP2B, HSPCB, and YWHAZ genes were appropriate for five types of muscle tissue (diaphragm muscle, heart, longissimus dorsi muscle, psoas major muscle, and tongue). In swine skeletal muscles, PPIA, HPRT, and eEF-1γ were considered to be the most stable genes.13 However, in this study, YWHAZ was not a suitable gene in the cardiac ventricle, atrium, psoas muscle, and femoral muscle, and PPIA was not stable in femoral muscle. These differences are thought to stem from differences in the protocols used.
- The internal organs are protected by the skin and soft tissues from the environment, so they are less affected by external factors. However, the degradation of body organ proteins is also influenced by temperature or pH. In skeletal muscle, some proteins showed a temperature-dependent cleavage over the first 48 hours postmortem. For example, the transformation of calcineurin A was completed within 24 hours. In contrast, protein phosphatase 2A increased within the first 24 hours after which it degraded at 21℃, but remained stable for up to 96 hours at 5℃ and 10℃.17 In forensic medicine, further studies could define appropriate correction factors that would be applied to compensate for the effect of temperature changes due to environmental conditions such as clothing, water suspension, leaf and other coverings, air movement, and surface properties.18 Melody et al.19 suggested that there might be an important linkage between the intercellular environment in the postmortem skeletal muscle and the activation of µ-calpain. Moreover, the rate of postmortem pH decline within the first six hours after death was shown to influence the rate of µ-calpain activity and autolysis that plays a pivotal role in regulating early postmortem proteolysis.19 Tissue accessibility, postmortem stability, and asepsis are all important attributes for postmortem interval changes, which are afforded by some tissues (e.g., muscle), but not by others (e.g., lung).17
- Based on previous studies, we searched more reference studies on postmortem proteolysis associated with postmortem intervals, but even data of protein expression on housekeeping genes were very rare. Once the pattern of postmortem protein degradation in the body is established, it can be used to estimate postmortem intervals, because some housekeeping genes analyzed in this study were stable in certain tissues. Since some animal and human proteins share high sequence homology, it is likely that the results from animal experiments will reflect human postmortem changes.17 In order to compare protein degradation during the postmortem interval, housekeeping genes that do not show differences in expression among individuals before death are needed as references, regardless of the damage or disease that caused the death. The current study is believed to offer a basic foundation for identification of proper housekeeping genes that would be selected for internal controls across various tissue types during postmortem intervals.
- As mentioned earlier, almost all other studies have been performed under diseased states. In contrast, the present study may help select stable housekeeping genes from normal body tissues other than the tissues under an altered condition or disease. Baseline data for further study on protein degradation patterns of internal control genes were presented in the current study. Accordingly, the results of this study may be used for determination of the elapsed time after death. However, in order to quantitate the patterns of postmortem protein degradation, other factors including temperature or pH should be considered in future studies.
DISCUSSION
Acknowledgments
Acknowledgments
- 1. Greer S, Honeywell R, Geletu M, Arulanandam R, Raptis L. Housekeeping genes; expression levels may change with density of cultured cells. J Immunol Methods 2010; 355: 76-79. ArticlePubMed
- 2. Liu NK, Xu XM. beta-tubulin is a more suitable internal control than beta-actin in western blot analysis of spinal cord tissues after traumatic injury. J Neurotrauma 2006; 23: 1794-1801. PubMed
- 3. Gu YR, Li MZ, Zhang K, et al. Evaluation of endogenous control genes for gene expression studies across multiple tissues and in the specific sets of fat- and muscle-type samples of the pig. J Anim Breed Genet 2011; 128: 319-325. ArticlePubMed
- 4. Zuo B, Xiong YZ, Deng CY, et al. Polymorphism, linkage mapping and expression pattern of the porcine skeletal muscle glycogen synthase (GYS1) gene. Anim Genet 2005; 36: 254-257. ArticlePubMed
- 5. Dawson B, Trapp RG. Basic and clinical biostatistics. New York: Lange Medical Books/McGraw-Hill, 2004.
- 6. Chia CY, Lim CW, Leong WT, Ling MH. High expression stability of microtubule affinity regulating kinase 3 (MARK3) makes it a reliable reference gene. IUBMB Life 2010; 62: 200-203. ArticlePubMed
- 7. Rho HW, Lee BC, Choi ES, Choi IJ, Lee YS, Goh SH. Identification of valid reference genes for gene expression studies of human stomach cancer by reverse transcription-qPCR. BMC Cancer 2010; 10: 240.ArticlePubMedPMCPDF
- 8. Gur-Dedeoglu B, Konu O, Bozkurt B, Ergul G, Seckin S, Yulug IG. Identification of endogenous reference genes for qRT-PCR analysis in normal matched breast tumor tissues. Oncol Res 2009; 17: 353-365. ArticlePubMed
- 9. Abasolo N, Torrell H, Roig B, Moyano S, Vilella E, Martorell L. RT-qPCR study on post-mortem brain samples from patients with major psychiatric disorders: reference genes and specimen characteristics. J Psychiatr Res 2011; 45: 1411-1418. ArticlePubMed
- 10. Everaert BR, Boulet GA, Timmermans JP, Vrints CJ. Importance of suitable reference gene selection for quantitative real-time PCR: special reference to mouse myocardial infarction studies. PLoS One 2011; 6: e23793.ArticlePubMedPMC
- 11. Li YL, Ye F, Hu Y, Lu WG, Xie X. Identification of suitable reference genes for gene expression studies of human serous ovarian cancer by real-time polymerase chain reaction. Anal Biochem 2009; 394: 110-116. ArticlePubMed
- 12. Shen Y, Li Y, Ye F, Wang F, Lu W, Xie X. Identification of suitable reference genes for measurement of gene expression in human cervical tissues. Anal Biochem 2010; 405: 224-229. ArticlePubMed
- 13. Feng X, Xiong Y, Qian H, Lei M, Xu D, Ren Z. Selection of reference genes for gene expression studies in porcine skeletal muscle using SYBR green qPCR. J Biotechnol 2010; 150: 288-293. ArticlePubMed
- 14. Langnaese K, John R, Schweizer H, Ebmeyer U, Keilhoff G. Selection of reference genes for quantitative real-time PCR in a rat asphyxial cardiac arrest model. BMC Mol Biol 2008; 9: 53.ArticlePubMedPMC
- 15. Couto MF, Peternelli LA, Barbosa MH. Classification of the coefficients of variation for sugarcane crops. Cienc Rural 2013; 43: 957-961. Article
- 16. Fritsche-Neto R, Vieira RA, Scapim CA, Miranda GV, Rezende LM. Updating the ranking of the coefficients of variation from maize experiments. Acta Sci Agron 2012; 34: 99-101. Article
- 17. Poloz YO, O'Day DH. Determining time of death: temperature-dependent postmortem changes in calcineurin A, MARCKS, CaMKII, and protein phosphatase 2A in mouse. Int J Legal Med 2009; 123: 305-314. ArticlePubMedPDF
- 18. Althaus L, Stuckradt S, Henssge C, Bajanowski T. Cooling experiments using dummies covered by leaves. Int J Legal Med 2007; 121: 112-114. ArticlePubMedPDF
- 19. Melody JL, Lonergan SM, Rowe LJ, Huiatt TW, Mayes MS, Huff-Lonergan E. Early postmortem biochemical factors influence tenderness and water-holding capacity of three porcine muscles. J Anim Sci 2004; 82: 1195-1205. PubMed
REFERENCES
Fig. 1Representative photos of Western blot analysis. (A) HPRT1 exhibits reproducibility in the cerebrum, cerebellum, spleen, but shows notable variation in the aorta. (B) GYS1 exhibits reproducibility in the cerebrum, cerebellum and femoral muscle, but it shows only faint bands in the liver, spleen, and kidney. (C) ACTB exhibits relatively reproducible expression level in the cerebrum, cerebellum, atrium, liver and kidney. Unlike other housekeeping genes, ACTB exhibits reproducibility in the aorta (individual rat numbers are shown as case no. 1 through no. 8).
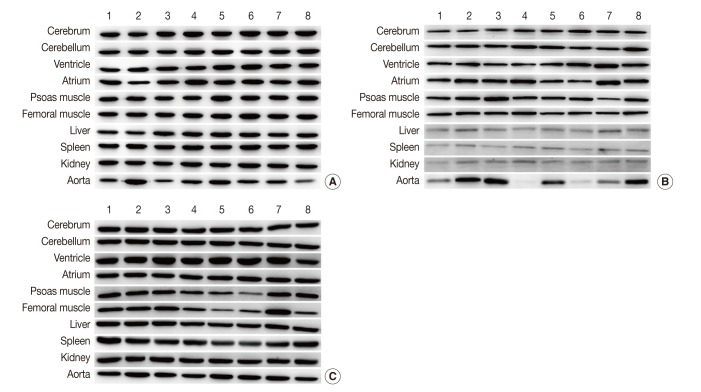

Fig. 2Expression stability of each gene represents as coefficient of variation (CV) value across tissue types. (A) HPRT1 shows stability in order of the spleen, kidney, cerebrum, cerebellum, ventricle, and femoral muscle, but a remarkably high CV value in the aorta. (B) PPIA shows the highest stability in the cerebellum. (C) With GYS1, all organ tissues are found to have CV values over 10%. The femoral muscle is relatively stable compared to other genes. (D) With TBP, all organ tissues are found to have CV values over 10%. (E) YWHAZ exhibits the highest stability in cerebellum, whereas it shows prominently high CV values in the femoral muscle and aorta. (F) GAPDH demonstrates the highest stability in the cerebrum, but significant variance in the aorta. (G) With ACTB, all organ tissues are found to have CV values over 10%. However, the kidney is relatively stable compared to other genes. Femoral m., femoral muscle; psoas m., Psoas muscle.
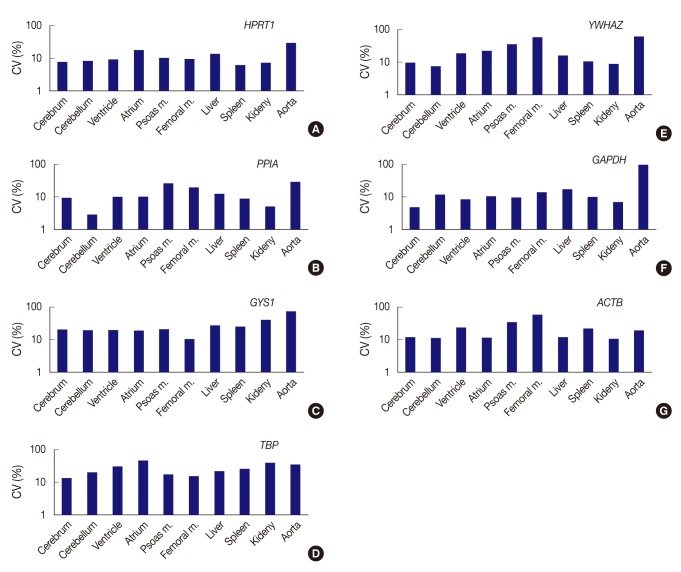

Fig. 3Comparison of expression stability between genes in each tissue type. (A) In the cerebrum, GAPDH demonstrates the highest stability, while GYS1 shows the least stable expression. (B) In the cerebellum, PPIA shows the highest stability, while TBP was the least stable. (C) In the ventricle, GAPDH exhibits the highest stability, while TBP was the least stable. (D) In the atrium, all seven genes are found to have coefficient of variation (CV) values over 10%. However, PPIA is relatively stable compared to other genes. (E) In psoas muscle, GAPDH shows the highest stability. (F) In femoral muscle, HPRT1 exhibits the highest stability. YWHAZ and ACTB demonstrate the highest variability in the femoral muscle. (G) In the liver, all genes are found to have CV values over 10%. However, ACTB is relatively stable compared to other genes. (H) In the spleen, HPRT1 demonstrates the highest stability. (I) In the kidney, PPIA shows the highest stability. (J) In aorta, all seven genes demonstrate high CV values over 10%.
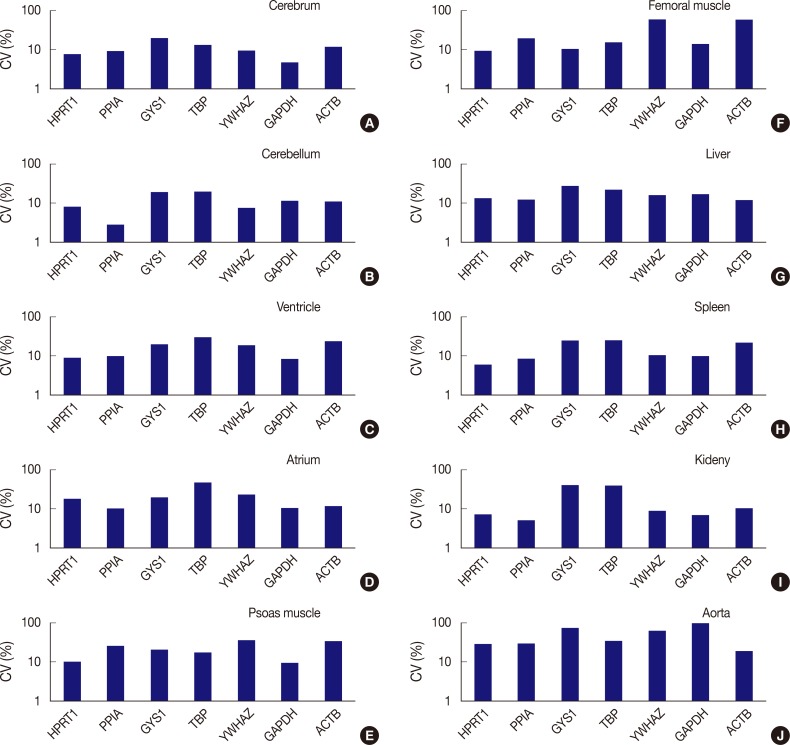

Table 1.List of antibodies used
Table 2.Average signal intensities in eight rats for comparison of protein expression in each gene across tissue types
Table 3.Coefficient of variation values (%) of each antibody according to tissue types
Figure & Data
References
Citations
Citations to this article as recorded by 

- scCompass: An Integrated Multi‐Species scRNA‐seq Database for AI‐Ready
Pengfei Wang, Wenhao Liu, Jiajia Wang, Yana Liu, Pengjiang Li, Ping Xu, Wentao Cui, Ran Zhang, Qingqing Long, Zhilong Hu, Chen Fang, Jingxi Dong, Chunyang Zhang, Yan Chen, Chengrui Wang, Guole Liu, Hanyu Xie, Yiyang Zhang, Meng Xiao, Shubai Chen, Haiping
Advanced Science.2025;[Epub] CrossRef - Evolutionary Insights into the Length Variation of DNA Damage Response Proteins Across Eukaryotes
Dominic Wiredu-Boakye, Laurence Higgins, Ondřej Gahura, Anzhelika Butenko, Guy Leonard, Mark A Freeman, Árni Kristmundsson, Karen Moore, Jamie W Harrison, Shani Mac Donald, Vyacheslav Yurchenko, Bryony A P Williams, Richard Chahwan, Courtney Stairs
Genome Biology and Evolution.2025;[Epub] CrossRef - Tissue-Specific Reference Genes in Rats: Insights into Cardiac, Muscular, and Skeletal Systems
Emre Can Günaydın, Beyza Göncü
International Journal of Life Sciences and Biotechnology.2025; 8(3): 219. CrossRef -
Reference Genes for Gene Expression Profiling in Mouse Models of
Listeria Monocytogenes
Infection
Lethicia Souza Tavares, Roberta Lane Oliveira-Silva, Marcelo Tigre Moura, Jéssica Barboza da Silva, Ana Maria Benko-Iseppon, José Vitor Lima-Filho
BioTechniques.2024; 76(3): 104. CrossRef - Sarcolipin relates to fattening, but not sarco/endoplasmic reticulum Ca2+-ATPase uncoupling, in captive migratory gray catbirds
Cory R. Elowe, Maria Stager, Alexander R. Gerson
Journal of Experimental Biology.2024;[Epub] CrossRef - Assessment of the alpha 7 nicotinic acetylcholine receptor as an imaging marker of cardiac repair-associated processes using NS14490
Victoria J. M. Reid, Wesley K. X. McLoughlin, Kalyani Pandya, Holly Stott, Monika Iškauskienė, Algirdas Šačkus, Judit A. Marti, Dominic Kurian, Thomas M. Wishart, Christophe Lucatelli, Dan Peters, Gillian A. Gray, Andrew H. Baker, David E. Newby, Patrick
EJNMMI Research.2024;[Epub] CrossRef - The interaction effect of high intensity interval swimming and gallic acid consumption on pain tolerance mechanisms in an animal model of Parkinson's disease
Raheleh Rastegarian, Mehrzad Moghadasi, Zahra Mosalanezhad, Reza Zeinalebadi
The Neuroscience Journal of Shefaye Khatam.2024; 12(4): 10. CrossRef - Identification of differentially expressed genes at different post-natal development stages of longissimus dorsi muscle in Tianzhu white yak
Bingang Shi, Xuehong Shi, Zhi Zuo, Shijie Zhao, Zhidong Zhao, Jiqing Wang, Huitong Zhou, Yuzhu Luo, Jiang Hu, Jon G.H. Hickford
Gene.2022; 823: 146356. CrossRef - Specialized androgen synthesis in skeletal muscles that actuate elaborate social displays
Eric R. Schuppe, Daniel Tobiansky, Franz Goller, Matthew J. Fuxjager
Journal of Experimental Biology.2022;[Epub] CrossRef - Deep-ultraviolet laser ablation sampling for proteomic analysis of tissue
Remilekun O. Lawal, Luke T. Richardson, Chao Dong, Fabrizio Donnarumma, Touradj Solouki, Kermit K. Murray
Analytica Chimica Acta.2021; 1184: 339021. CrossRef - Protective effects of grape seed proanthocyanidins against iron overload-induced renal oxidative damage in rats
Shaojun Yun, Dongyang Chu, Xingshuai He, Wenfang Zhang, Cuiping Feng
Journal of Trace Elements in Medicine and Biology.2020; 57: 126407. CrossRef - Gene Expression Variation of Candidate Endogenous Control Genes Across Latitudinal Populations of the Bank Vole (Clethrionomys glareolus)
Lucie Němcová, Silvia Marková, Petr Kotlík
Frontiers in Ecology and Evolution.2020;[Epub] CrossRef - High-Fat Diet Alters the Expression of Reference Genes in Male Mice
Xiuqin Fan, Hongyang Yao, Xuanyi Liu, Qiaoyu Shi, Liang Lv, Ping Li, Rui Wang, Tiantian Tang, Kemin Qi
Frontiers in Nutrition.2020;[Epub] CrossRef - The applicability of forensic time since death estimation methods for buried bodies in advanced decomposition stages
Stefan Pittner, Valentina Bugelli, M. Eric Benbow, Bianca Ehrenfellner, Angela Zissler, Carlo P. Campobasso, Roelof-Jan Oostra, Maurice C. G. Aalders, Richard Zehner, Lena Lutz, Fabio C. Monticelli, Christian Staufer, Katharina Helm, Vilma Pinchi, Joseph
PLOS ONE.2020; 15(12): e0243395. CrossRef - Doublecortin-like Kinase 1 Regulates α-Synuclein Levels and Toxicity
Gabriel E. Vázquez-Vélez, Kristyn A. Gonzales, Jean-Pierre Revelli, Carolyn J. Adamski, Fatemeh Alavi Naini, Aleksandar Bajić, Evelyn Craigen, Ronald Richman, Sabrina M. Heman-Ackah, Matthew J.A. Wood, Maxime W.C. Rousseaux, Huda Y. Zoghbi
The Journal of Neuroscience.2020; 40(2): 459. CrossRef - The role of extracellular matrix stiffness in regulating cytoskeletal remodeling via vinculin in synthetic smooth muscle cells
Kai Shen, Harshavardhan Kenche, Hua Zhao, Jinping Li, Jasimine Stone
Biochemical and Biophysical Research Communications.2019; 508(1): 302. CrossRef - Combined proteomic and miRNome analyses of mouse testis exposed to an endocrine disruptors chemicals mixture reveals altered toxicological pathways involved in male infertility
Julio Buñay, Eduardo Larriba, Daniel Patiño-Garcia, Paulina Urriola-Muñoz, Ricardo D Moreno, Jesús del Mazo
MHR: Basic science of reproductive medicine.2019; 25(3): 156. CrossRef - Postmortem proteomics to discover biomarkers for forensic PMI estimation
Kyoung-Min Choi, Angela Zissler, Eunjung Kim, Bianca Ehrenfellner, Eunji Cho, Se-in Lee, Peter Steinbacher, Ki Na Yun, Jong Hwan Shin, Jin Young Kim, Walter Stoiber, Heesun Chung, Fabio Carlo Monticelli, Jae-Young Kim, Stefan Pittner
International Journal of Legal Medicine.2019; 133(3): 899. CrossRef - Aging‐associated changes in hippocampal glycogen metabolism in mice. Evidence for and against astrocyte‐to‐neuron lactate shuttle
Dominika Drulis‐Fajdasz, Agnieszka Gizak, Tomasz Wójtowicz, Jacek R. Wiśniewski, Dariusz Rakus
Glia.2018; 66(7): 1481. CrossRef - New insights into the distribution, protein abundance and subcellular localisation of the endogenous peroxisomal biogenesis proteins PEX3 and PEX19 in different organs and cell types of the adult mouse
Claudia Colasante, Jiangping Chen, Barbara Ahlemeyer, Rocio Bonilla-Martinez, Srikanth Karnati, Eveline Baumgart-Vogt, Stephan N. Witt
PLOS ONE.2017; 12(8): e0183150. CrossRef - Identification of valid endogenous control genes for determining gene expression in C6 glioma cell line treated with conditioned medium from adipose-derived stem cell
I.C. Iser, R.P. de Campos, A.P.S. Bertoni, M.R. Wink
Biomedicine & Pharmacotherapy.2015; 75: 75. CrossRef - Sclerosing Mucoepidermoid Carcinoma in the Parotid Gland: Literature Review
Chang-Ki Woo, Bae-Hyun Kim, Byung-Joo Lee, Jin-Choon Lee
Korean Journal of Otorhinolaryngology-Head and Neck Surgery.2012; 55(8): 508. CrossRef
Evaluation of Protein Expression in Housekeeping Genes across Multiple Tissues in Rats
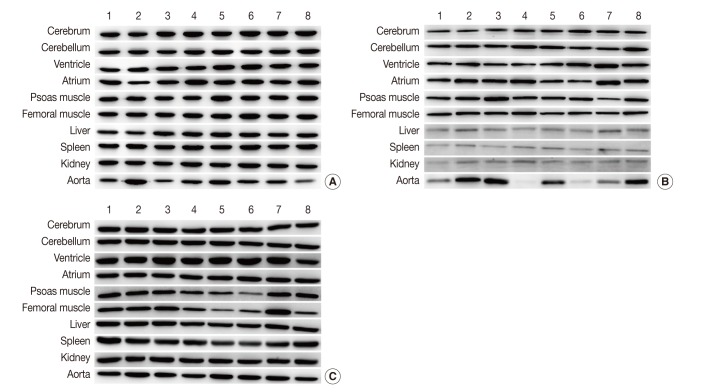
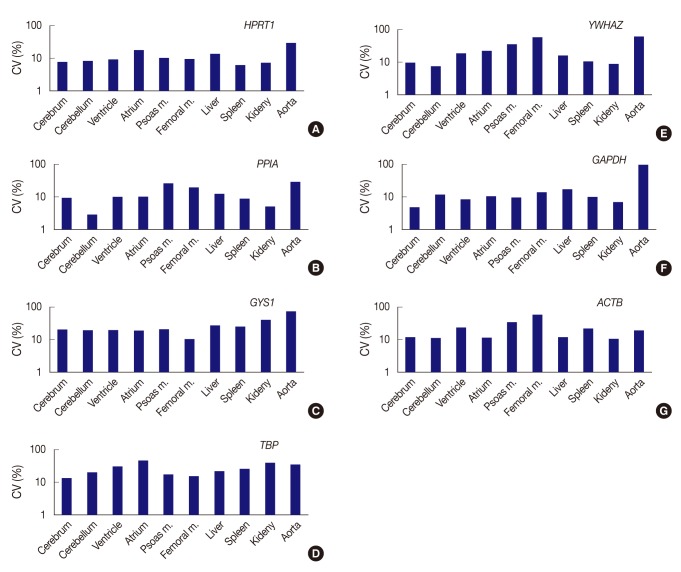
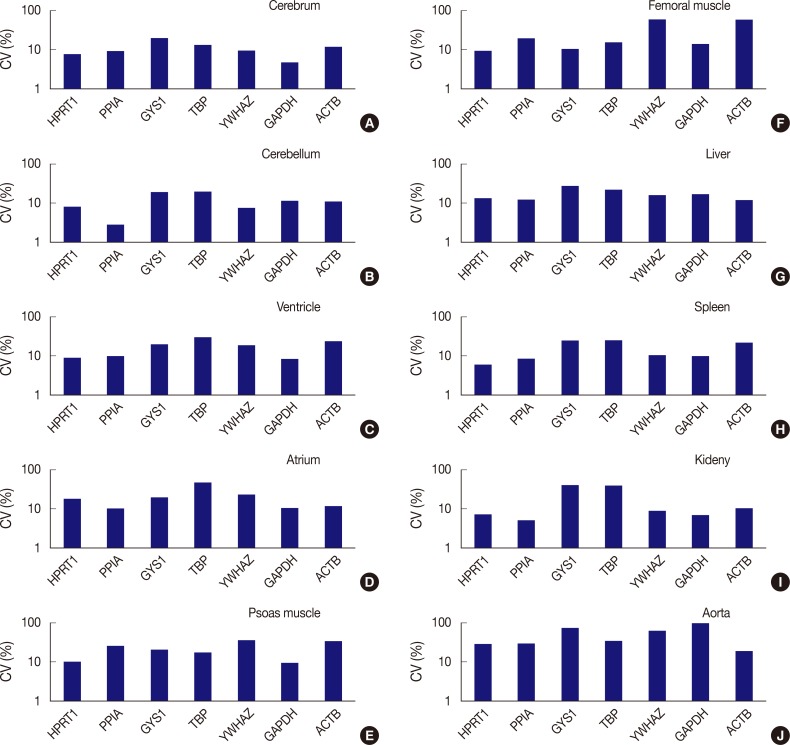
Fig. 1 Representative photos of Western blot analysis. (A) HPRT1 exhibits reproducibility in the cerebrum, cerebellum, spleen, but shows notable variation in the aorta. (B) GYS1 exhibits reproducibility in the cerebrum, cerebellum and femoral muscle, but it shows only faint bands in the liver, spleen, and kidney. (C) ACTB exhibits relatively reproducible expression level in the cerebrum, cerebellum, atrium, liver and kidney. Unlike other housekeeping genes, ACTB exhibits reproducibility in the aorta (individual rat numbers are shown as case no. 1 through no. 8).
Fig. 2 Expression stability of each gene represents as coefficient of variation (CV) value across tissue types. (A) HPRT1 shows stability in order of the spleen, kidney, cerebrum, cerebellum, ventricle, and femoral muscle, but a remarkably high CV value in the aorta. (B) PPIA shows the highest stability in the cerebellum. (C) With GYS1, all organ tissues are found to have CV values over 10%. The femoral muscle is relatively stable compared to other genes. (D) With TBP, all organ tissues are found to have CV values over 10%. (E) YWHAZ exhibits the highest stability in cerebellum, whereas it shows prominently high CV values in the femoral muscle and aorta. (F) GAPDH demonstrates the highest stability in the cerebrum, but significant variance in the aorta. (G) With ACTB, all organ tissues are found to have CV values over 10%. However, the kidney is relatively stable compared to other genes. Femoral m., femoral muscle; psoas m., Psoas muscle.
Fig. 3 Comparison of expression stability between genes in each tissue type. (A) In the cerebrum, GAPDH demonstrates the highest stability, while GYS1 shows the least stable expression. (B) In the cerebellum, PPIA shows the highest stability, while TBP was the least stable. (C) In the ventricle, GAPDH exhibits the highest stability, while TBP was the least stable. (D) In the atrium, all seven genes are found to have coefficient of variation (CV) values over 10%. However, PPIA is relatively stable compared to other genes. (E) In psoas muscle, GAPDH shows the highest stability. (F) In femoral muscle, HPRT1 exhibits the highest stability. YWHAZ and ACTB demonstrate the highest variability in the femoral muscle. (G) In the liver, all genes are found to have CV values over 10%. However, ACTB is relatively stable compared to other genes. (H) In the spleen, HPRT1 demonstrates the highest stability. (I) In the kidney, PPIA shows the highest stability. (J) In aorta, all seven genes demonstrate high CV values over 10%.
Fig. 1
Fig. 2
Fig. 3
Evaluation of Protein Expression in Housekeeping Genes across Multiple Tissues in Rats
| Antibody | Molecular weight (kDa) | Company | Dilution | Organ(s) |
|---|---|---|---|---|
| HPRT1 | 24 | Abcam | 1:16,000 | Psoas muscle, femoral muscle |
| 1:32,000 | Cerebrum, cerebellum, ventricle, atrium, aorta, liver, kidney, spleen | |||
| PPIA | 18 | Cell Signaling | 1:1,000 | Ventricle, atrium, liver, kidney, spleen |
| 1:8,000 | Cerebrum, cerebellum, aorta, psoas muscle, femoral muscle | |||
| GYS1 | 85 | Abcam | 1:1,000 | Cerebrum, cerebellum, aorta, liver, kidney, spleen |
| 1:8,000 | Ventricle, atrium, psoas muscle, femoral muscle | |||
| TBP | 38 | Abcam | 1:1,000 | Ventricle, atrium, liver, kidney, spleen, cerebrum, cerebellum, aorta, psoas muscle, and femoral muscle |
| YWHAZ | 30 | Millipore | 1:4,000 | Ventricle, atrium, psoas muscle, femoral muscle |
| 1:8,000 | Aorta, liver, kidney, spleen | |||
| 1:16,000 | Cerebrum, cerebellum | |||
| GAPDH | 37 | Cell Signaling | 1:64,000 | Ventricle, atrium, liver, kidney, spleen, cerebrum, cerebellum, aorta, psoas muscle, femoral muscle |
| ACTB | 42 | Sigma | 1:50,000 | Ventricle, atrium, liver, kidney, spleen, cerebrum, cerebellum, aorta, psoas muscle, femoral muscle |
| HPRT1 | PPIA | GYS1 | TBP | YWHAZ | GAPDH | ACTB | |
|---|---|---|---|---|---|---|---|
| Cerebrum | 9.82 ± 0.75 | 10.23 ± 0.95 | 9.07 ± 1.84 | 10.49 ± 1.41 | 11.27 ± 1.10 | 11.58 ± 0.56 | 9.36 ± 1.12 |
| Cerebellum | 9.85 ± 0.81 | 10.92 ± 0.31 | 9.57 ± 1.84 | 9.79 ± 1.96 | 11.18 ± 0.84 | 9.17 ± 1.06 | 9.35 ± 1.05 |
| Ventricle | 9.77 ± 0.89 | 9.92 ± 0.99 | 10.05 ± 1.97 | 5.63 ± 1.72 | 10.17 ± 1.92 | 11.40 ± 0.96 | 6.77 ± 1.60 |
| Atrium | 9.72 ± 1.72 | 10.91 ± 1.10 | 9.83 ± 1.87 | 4.65 ± 2.10 | 10.14 ± 2.28 | 9.35 ± 0.98 | 8.10 ± 0.93 |
| Psoas muscle | 9.81 ± 0.99 | 9.33 ± 2.38 | 10.10 ± 2.08 | 9.71 ± 1.69 | 7.15 ± 2.55 | 11.57 ± 1.11 | 5.79 ± 1.95 |
| Femoral muscle | 10.04 ± 0.94 | 9.85 ± 1.90 | 10.01 ± 1.05 | 9.39 ± 1.44 | 6.12 ± 3.54 | 11.55 ± 1.62 | 7.50 ± 4.33 |
| Liver | 9.03 ± 1.22 | 10.43 ± 1.28 | 1.99 ± 0.54 | 4.91 ± 1.08 | 9.51 ± 1.53 | 11.2 ± 1.92 | 9.51 ± 1.14 |
| Spleen | 9.90 ± 0.61 | 10.41 ± 0.92 | 2.87 ± 0.72 | 9.50 ± 2.42 | 9.08 ± 0.98 | 10.60 ± 1.07 | 10.33 ± 2.29 |
| Kidney | 9.57 ± 0.69 | 10.37 ± 0.53 | 1.88 ± 0.76 | 3.11 ± 1.21 | 9.77 ± 0.87 | 10.41 ± 0.73 | 7.73 ± 0.81 |
| Aorta | 10.99 ± 3.17 | 9.00 ± 2.57 | 7.33 ± 5.38 | 6.41 ± 2.19 | 5.61 ± 3.44 | 9.60 ± 9.08 | 10.63 ± 2.01 |
| HPRT1 | PPIA | GYS1 | TBP | YWHAZ | GAPDH | ACTB | |
|---|---|---|---|---|---|---|---|
| Cerebrum | 7.64 | 9.29 | 20.29 | 13.44 | 9.76 | 4.84 | 11.97 |
| Cerebellum | 8.22 | 2.84 | 19.23 | 20.02 | 7.51 | 11.56 | 11.23 |
| Ventricle | 9.11 | 9.98 | 19.60 | 30.55 | 18.88 | 8.42 | 23.63 |
| Atrium | 17.70 | 10.08 | 19.02 | 45.16 | 22.49 | 10.48 | 11.48 |
| Psoas muscle | 10.09 | 25.51 | 20.59 | 17.40 | 35.66 | 9.59 | 33.68 |
| Femoral muscle | 9.36 | 19.29 | 10.49 | 15.34 | 57.84 | 14.03 | 57.73 |
| Liver | 13.51 | 12.27 | 27.14 | 22.00 | 16.09 | 17.14 | 11.99 |
| Spleen | 6.16 | 8.84 | 25.09 | 25.47 | 10.79 | 10.09 | 22.17 |
| Kidney | 7.21 | 5.11 | 40.43 | 38.91 | 8.90 | 7.01 | 10.48 |
| Aorta | 28.84 | 28.56 | 73.40 | 34.17 | 61.32 | 94.58 | 18.91 |
Table 1. List of antibodies used
Table 2. Average signal intensities in eight rats for comparison of protein expression in each gene across tissue types
Values are presented as mean±standard deviation.
Table 3. Coefficient of variation values (%) of each antibody according to tissue types

 E-submission
E-submission
 PubReader
PubReader Cite this Article
Cite this Article




