Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 46(6); 2012 > Article
-
Original Article
Methylation and Immunoexpression ofp16INK4a Tumor Suppressor Gene in Primary Breast Cancer Tissue and Their Quantitativep16INK4a Hypermethylation in Plasma by Real-Time PCR - Jae Jun Lee, Eunkyung Ko1, Junhun Cho, Ha Young Park, Jeong Eon Lee1, Seok Jin Nam1, Duk-Hwan Kim2, Eun Yoon Cho
-
Korean Journal of Pathology 2012;46(6):554-561.
DOI: https://doi.org/10.4132/KoreanJPathol.2012.46.6.554
Published online: December 26, 2012
Department of Pathology, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea.
1Department of Surgery, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea.
2Department of Genome Research, Samsung Biomedical Research Institute, Sungkyunkwan University School of Medicine, Seoul, Korea.
- Corresponding Author: Eun Yoon Cho, M.D. Department of Pathology, Samsung Medical Center, Sungkyunkwan University School of Medicine, 81 Irwon-ro, Gangnam-gu, Seoul 135-710, Korea. Tel: +82-2-3410-2800, Fax: +82-2-3410-0025, eunyoon.cho@samsung.com
© 2012 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- The p16INK4a gene methylation has been reported to be a major tumorigenic mechanism.
-
Methods
- We evaluated the methylation status of the p16INK4a genes in 231 invasive breast cancer and 90 intraductal carcinoma specimens using a methylation-specific polymerase chain reaction and p16 protein expression using immunohistochemistry. The quantity of cell-free methylated p16INK4a DNA in the plasma samples of 200 patients with invasive breast cancer was also examined using a fluorescence-based real-time polymerase chain reaction assay.
-
Results
- The frequencies of p16INK4a methylation in invasive and intraductal tumors were 52.8% (122/231) and 57.8% (52/90), respectively. The p16 protein was overexpressed in 145 of the 231 invasive carcinomas (62.8%) and 63 of the 90 intraductal carcinomas (70%). High p16 expression in invasive carcinomas correlated significantly with a high histologic grade, a negative estrogen receptor and progesterone receptor status, p53 immunoreactivity and high Ki-67 expression with immunohistochemistry. In addition, the methylation index of p16INK4a was significantly higher in the cancer patients than the normal controls (p<0.001).
-
Conclusions
- High p16 immunoreactivity correlated with a loss of differentiation in breast carcinomas and high frequency of p16INK4a promoter methylation in both invasive and intraductal carcinomas, suggesting it may be involved in the pathogenesis of breast cancer.
- Study material and DNA extraction
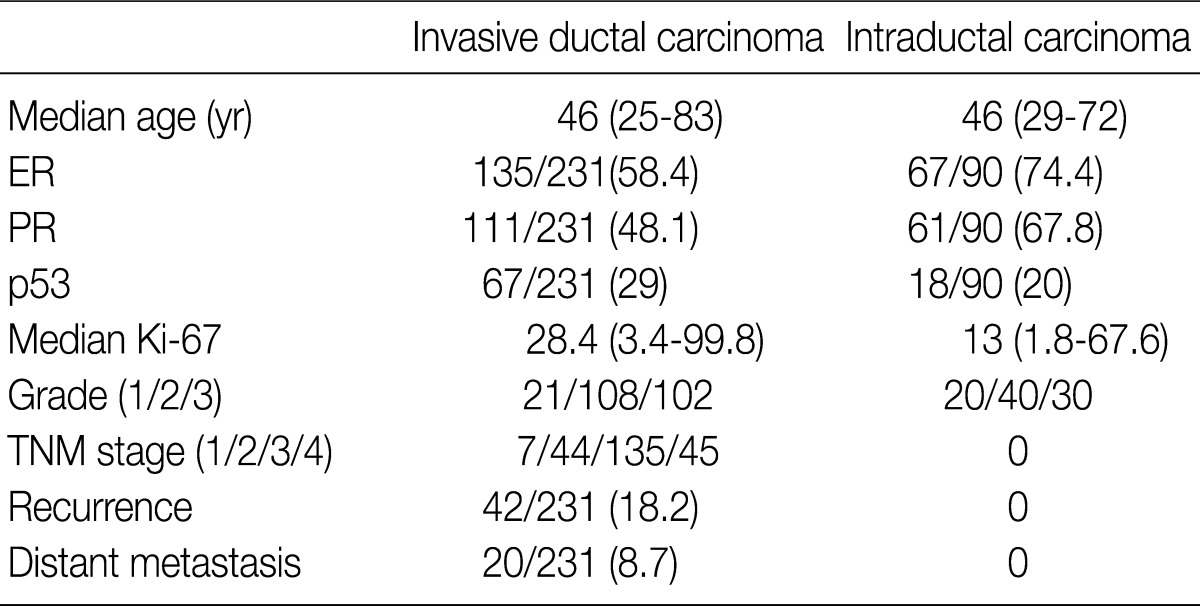
- We obtained tumor specimens from 231 patients with invasive breast carcinoma and from 200 matched preoperative plasma and tumor specimens from 90 patients with intraductal carcinoma who underwent surgical resection at Samsung Medical Center from May 2002 through December 2004. All of the tissues had been fixed in 10% neutral buffered formalin and embedded in paraffin and were evaluated for a wide variety of pathologic features by examination of the hematoxylin and eosin (H&E)-stained tissue sections and the immunohistochemically-stained slides by two investigators in a blinded fashion without knowledge of the molecular data. The medical records were reviewed for clinical information. The tumor grade and stage were determined based on the American Joint Committee on Cancer guidelines. The clinicopathological findings of the patients are presented in Table 1. All of the patients were women, and the median age was 46 years (range, 25 to 83 years). The mean number of metastatic lymph nodes was 4.95 (range, 1 to 27; median, 3.00). The survival data were obtained from a review of the patient's chart in our hospital and from the National Cancer Registry. The mean follow-up time was 45.5 (range, 1 to 63) months. Recurrence-free survival was defined as the number of months from diagnosis to the occurrence of an event (local recurrence/metastasis). Plasma specimens were also obtained from 189 healthy individuals (109 men and 80 women) as controls.
- For DNA extraction, we used fresh-frozen tissues for invasive carcinoma and formalin-fixed, paraffin-embedded tissues for intraductal carcinoma. The H&E-stained sections were reviewed to confirm the tumor areas and tumor volume before DNA extraction. The tumor-rich areas (>75%) were extracted from 0.1% methylene blue-stained serial 10 µm tissue sections, and the areas corresponding to tumor parenchyma were carefully microdissected from the surrounding stromal tissues. DNA was extracted using a DNeasy tissue kit (Qiagen, Valencia, CA, USA) according to the manufacturer's instructions.
- Assembly of tissue microarrays
- The specimens were fixed in 10% buffered formalin, and embedded in paraffin. The representative areas on the H&E-stained sections were carefully selected and marked on the individual paraffin blocks. Two tissue cores (1 mm in diameter) were obtained from the regions of interest in each tumor block. The tissue cores were then inserted in a recipient paraffin block according to the manufacturer's instructions (Isu Abxis Co., Ltd., Cambridge, UK).
- Immunohistochemistry
- After deparaffinization and rehydration, the 4 µm thick sections on the silane-coated slides were heat-pretreated in a citrate buffer (pH 7.3 at 92℃ in a microwave oven) and immunostained using specific antibodies directed against p16 (G175-405, Pharmingen, Hamburg, Germany), Her-2 (CB11, Novocastra Laboratories, Newcastle, UK), estrogen receptor (ER; 6F11, Novocastra Laboratories), progesterone receptor (PR; 1A6, Novocastra Laboratories), p53 (Zymed, South San Francisco, CA, USA), and Ki-67 (MIB I, Dianova, Hamburg, Germany). An avidin-biotin technique was applied using 3,3'-diaminobenzidine (DAB) for visualization and hematoxylin as the nuclear counterstain. Visual estimation of the nuclear and/or cytoplasmic staining of p16 was performed. A cell was considered positive for p16 protein if it contained staining within the nucleus, cytoplasm, or both the nucleus and cytoplasm. The number of stained tumor cells was determined semiquantitatively. The proportion of positive cells was classified into five categories: 0 (1-4% positive tumor cells), 1 (5-10%), 2 (10-50%), 3 (51-80%), or 4 (>80%). The cases with more than 5% positive cells were defined as positive. The staining intensity was scored as follows: 1 (weak), 2 (moderate), or 3 (strong). The scores for both parameters were multiplied to generate the immunoreactivity score (IS) for each case. As a result, the IS range was from 0 to 12. Each lesion was examined and scored separately by two pathologists, and the cases with discrepant scores were discussed until agreement was reached between the pathologists.
- Protein expression was determined in a blinded manner for the methylation analyses and vice versa. The distribution of Her-2 was assessed according to the following scoring system: 0, no immunoreactivity or immunoreactivity in less than 10% of the tumor cells; 1+, faint and incomplete staining of more than 10% of the tumor cells; 2+, weak to moderate complete membrane immunoreactivity in more than 10% of the tumor cells; 3+, moderate to strong complete membrane immunoreactivity in more than 30% of the tumor cells. The semiquantitative histochemical scores were used to record the results of the ER and PR staining according to the Allred Scoring System.18 The p53 staining in the cell nuclei was assessed, and the patients with staining in more than 5% of the tumor cells were regarded as positive.
- DNA modification and methylation analysis
- One microgram of genomic DNA was modified using sodium bisulfite with the EZ DNA methylation kit (Zymo Research, Orange, CA, USA) according to the manufacturer's instructions. Modified DNA was stored in aliquots at -20℃ until required. The methylation status of the p16INK4a gene was determined by methylation-specific PCR (MSP) using two pairs of primers, as described by Herman et al.3 The MSP primers specific for methylated p16INK4a gene were 5'-TTATTAGAGGGTGGGGCGGATCGC-3' (sense) and 5'-GACCCCGAACCGCGACCGTAA-3' (antisense), and the primers used for the unmethylated p16INK4a gene were 5'-TTATTAGAGGGTGGGGTGGATTGT-3' (sense) and 5'-CCACCTAAATCAACCTCCAACCA-3' (antisense). Amplification was carried out over 35 cycles (1 minute at 94℃, 1 minute at the annealing temperature, and 1 minute at 72℃), followed by 4 minutes at 72℃. The PCR product sizes were 150 bp and 234 bp for the methylated and unmethylated segments, respectively. CpGenome Universal Unmethylated DNA (S7822, Millipore, Billerica, MA, USA) was used as a positive control for the unmethylated alleles, and CpGenome Universal Methylated DNA (S7821, Millipore) was used as a positive control for the methylated alleles. The PCR products were visualized on 2% agarose gels stained with ethidium bromide.
- Quantitative real-time MSP of plasma
- Quantitative real-time MSP was performed. The primer sequences used to amplify the actin gene were 5'-TGGTGATGGAGGAGGTTTAGTAAGT-3' (sense) and 5'-AACCAATAAAACCTACTCCTCCCTTAA-3' (antisense) were used in conjunction with a fluorogenic probe 5'-(FAM)-ACCACCACCCAACACACAATAACAAACACA-(TAMRA)-3', and the primer sequences used for the methylated p16INK4a gene were 5'-GGGGAGAGTAGATAGCGGGC-3' (sense) and 5'-AACCAATCAACCGAAAATTCCATA-3' (antisense) in conjunction with a fluorogenic probe 5'-(FAM)-TACTCCCCGCCGCCGACTCCAT-(TAMRA)-3'. Each 10 µL PCR reaction volume contained 5 µL TaqMan Universal PCR Master Mix No AmpErase UNG (Applied Biosystems, Foster City, CA, USA), 1 pM of each primer, a 0.025 pM probe, and 2 µL template DNA. Assays were run on an ABI Prism 7900 Sequence Detection System (Applied Biosystems). The PCR conditions were as follows: 2 minutes at 50℃, 10 minutes at 95℃, followed by 50 cycles of 15 seconds at 95℃, and 1 minute at 60℃. Real-time quantitative PCR reactions were performed for the detection and quantitation of the bisulfite unconverted methylated version of the p16INK4a gene and the bisulfite-converted unmethylated version of the actin gene. Serial dilutions of methylated or unmethylated control genomic DNAs (Millipore) were used to construct standard curves. The methylation index (MI) in each sample was calculated using the following equation:19
- MI values greater than 0.1% were considered positive.
- Statistical analysis
- Statistical analyses of the immunohistochemical abnormalities were performed using the two-tailed Fisher's exact test with a p-value set as <0.05 to indicate significance. Fisher's exact tests were used to compare aberrant p16 gene expression and p16INK4a methylation with the clinicopathologic parameters of breast cancer. Comparison of the amount of methylated p16INK4a DNA in the plasma between the invasive breast cancer patients and normal controls was performed using the Student's t-test (two-tailed). All statistical comparisons were performed using the statistical software, SPSS (SPSS Inc., Chicago, IL, USA).
MATERIALS AND METHODS
- Methylation analysis
- We analyzed the pattern of p16INK4a methylation in the 231 invasive breast carcinomas and 90 intraductal carcinomas. The results of these experiments are summarized in Table 2 and representatively illustrated in Fig. 1. The p16INK4a promoter was frequently hypermethylated, not only in the invasive tumor samples but also in the intraductal carcinoma specimens. In many of the specimens examined (122/231 [52.8%] for invasive ductal carcinoma and 52/90 [57.8%] for intraductal carcinoma) we observed the presence of the amplification band corresponding to the methylated target sequence. We also detected the unmethylated allele in 114 invasive cancer and 53 intraductal carcinoma specimens, suggesting contamination of the dissected tumor samples with normal cells. Ninety-eight cases of invasive cancer and 43 cases of intraductal carcinoma were fully unmethylated.
- Immunohistochemical analysis
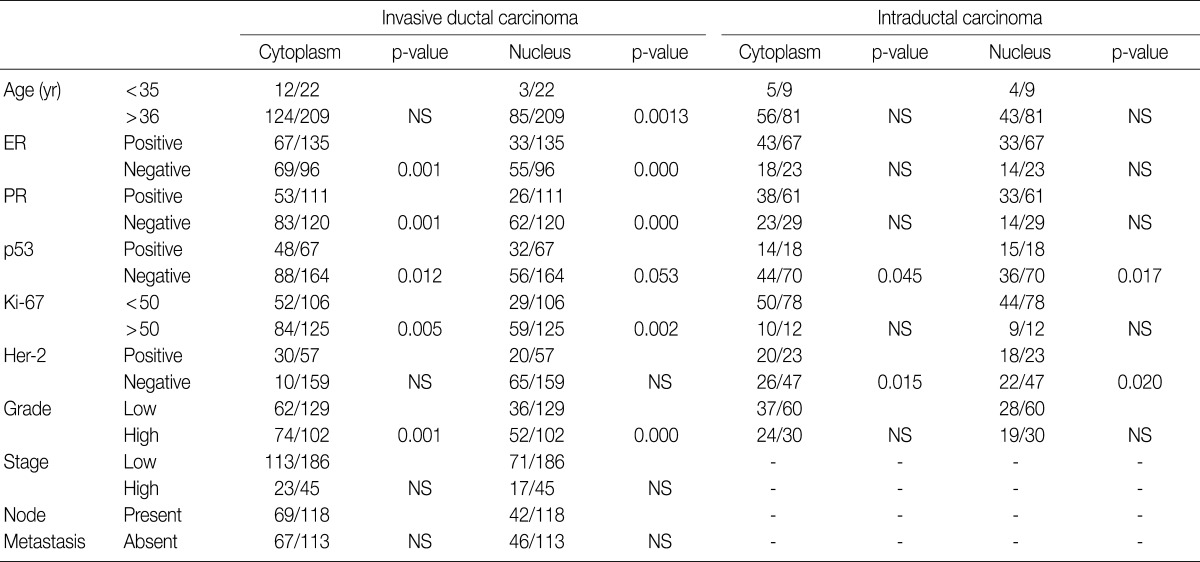
- The majority of breast carcinoma samples moderately or strongly expressed p16 in either the infiltrating or intraductal component, or both (Table 2, Fig. 2). As shown in Table 2, immunohistochemical analysis showed that 145 of 231 invasive carcinomas (62.8%) and 63 of 90 intraductal carcinomas (70%) overexpressed the p16 protein. The reactivity was predominantly nuclear and cytoplasmic or cytoplasmic alone (Fig. 2). The overexpression of p16 in the invasive carcinomas was significantly associated with a high histologic grade, negative ER and PR status, p53 immunoreactivity, and high Ki-67 proliferation index (Table 3). The IS score of the p16 immunoreactivity also correlated with the above variables. No correlation of p16 expression with clinical stage, HER2/neu status, or disease free survival was found.
- Relationship between MSP and immunohistochemical analyses
- In our study, there was no clear association between promoter hypermethylation of p16INK4a and p16 expression. In the invasive carcinomas, 78 of 122 tumors with methylation revealed p16 overexpression (p=0.7) while 44 tumors showed loss of protein. In the intraductal carcinomas, 34 of 52 tumors with methylation had high expression of p16 (p=0.269), while 18 tumors showed loss of protein (Table 2).
- Plasma concentration of methylated p16INK4a promoter DNA in the breast carcinoma patients
- Peripheral blood plasma was obtained from 200 invasive breast cancer patients before surgery and from 189 healthy donors, which were used as normal controls. The amount of methylated p16INK4a gene copies was examined using the quantitative real-time MSP approach and was expressed as the MI (%). Methylated p16INK4a was found in the plasma of 69 of the 200 (34.5%) invasive breast cancer patients and in 2 of the 189 (1%) normal controls. Among the control group, all of the subjects with methylated p16INK4a were men. The mean MI for p16INK4a was 6.37% (range, 0 to 85%) in the cancer patients and 0.06% (range, 0 to 7.3%) in the normal controls (Fig. 3). The mean concentration of methylated p16INK4a in the plasma of the invasive breast cancer patients was higher than that in the normal healthy control group, which was statistically significant (t-test, p=0.016).
RESULTS
- p16INK4a promoter methylation is variably present in breast carcinoma with a prevalence ranging from 4% to 68.4%,3,20-22 and our present study revealed a relatively high frequency of methylation of 52.8% (122/231) and 57.8% (52/90) in the invasive and intraductal tumors, respectively. The high prevalence of p16INK4a methylation in both the invasive and intraductal carcinomas may indicate that the epigenetic alterations of the p16INK4a gene play a significant role in the early stage of mammary carcinogenesis. Recently, Liu et al.23 suggested that p16INK4a methylation in various intraductal proliferative lesions, such as usual ductal hyperplasia, flat epithelial atypia, atypical ductal hyperplasia, low grade intraductal carcinomas, and high grade intraductal carcinomas, was related to the progression of intraductal epithelial proliferative lesions of the breast.
- The p16INK4a gene is epigenetically silenced in many human tumors; however, regardless of the cut-off point of immunoreactivity, our study did not show a firm association between methylation and protein expression. Similar findings have been reported in breast carcinomas and in other malignancies.4,24-26 There are several possible explanations for this discrepancy between immunohistochemistry and promoter hypermethylation. For example, the discrepancy may be caused by tumor heterogeneity. It is possible that both p16-positive cells and -negative cells were present in some cases, and that the predominant p16-positive cells were analyzed for p16INK4a promoter hypermethylation. Another explanation may be the regulation of protein expression by post-transcriptional alterations other than promoter methylation. Moreover, the immunoreactivity of p16 proteins has been reported to be variable according to different anti-p16 antibodies. The G175-405 antibody used in our study has been reported to be the most specific.27
- The present study showed that p16 protein overexpression was correlated with several phenotypic parameters of the tumors indicative of a poor prognosis, although it was not associated with disease-free survival or overall survival. This suggested that the high expression of p16 correlated with an aggressive tumor phenotype. Some reports in the literature are in agreement with the above findings,5-7,28 whereas other reports did not find any correlation.29 The pathophysiological role of p16 in the oncogenesis of breast carcinoma should be focused in further investigation. p16 is a tumor suppressor, and it is accepted that the loss of p16 expression is related to tumorigenesis. Therefore, our data could be interpreted as overexpression of p16 as an attempt to compensate for the oncogenic changes of other proteins, such as retinoblastoma or TP53.30 In the present study, high expression of p16 correlated with both p53 protein positivity and a high Ki-67 proliferation index. The p16 overexpression in the patients with high proliferative activity may indicate that p16 is inactive or not sufficient to limit cell growth in breast carcinomas.
- By using a quantitative PCR approach, we found that methylated p16INK4a DNA was present in the plasma of both the cancer patients and healthy subjects. However, the invasive breast cancer patients had significantly higher frequencies and concentrations of methylated p16INK4a DNA than the normal controls. Our results suggest that a quantitative measurement of methylated p16INK4a DNA concentrations in the plasma of cancer patients might be a useful tool for cancer screening and monitoring of treatment effectiveness. Nevertheless, a large-scale prospective study of plasma from cancer patients is necessary to further investigate the feasibility of methylated p16INK4a DNA as a tumor screening marker in invasive breast cancer.
- In conclusion, the high frequency of p16INK4a methylation in invasive breast carcinoma and intraductal carcinoma suggest that p16INK4a methylation could be a common and early event in breast carcinogenesis. The overexpression of p16 in the invasive carcinomas was correlated with several phenotypic parameters of the tumors indicative of poor prognosis, although it was not associated with disease free survival or overall survival. Therefore, this suggested that the high level of p16 expression may be an indicator of poor prognosis. Moreover, the concentrations of p16INK4a methylated DNA was significantly higher in the plasma of the invasive breast cancer patients compared with the healthy individuals, suggesting that quantitative measurement of plasma methylated p16INK4a gene concentrations might be a useful and non-invasive tool for cancer screening and monitoring treatment response.
DISCUSSION
- 1. Kamb A, Gruis NA, Weaver-Feldhaus J, et al. A cell cycle regulator potentially involved in genesis of many tumor types. Science 1994; 264: 436-440. ArticlePubMed
- 2. Serrano M, Hannon GJ, Beach D. A new regulatory motif in cell-cycle control causing specific inhibition of cyclin D/CDK4. Nature 1993; 366: 704-707. ArticlePubMedPDF
- 3. Herman JG, Merlo A, Mao L, et al. Inactivation of the CDKN2/p16/MTS1 gene is frequently associated with aberrant DNA methylation in all common human cancers. Cancer Res 1995; 55: 4525-4530. PubMed
- 4. Di Vinci A, Perdelli L, Banelli B, et al. p16(INK4a) promoter methylation and protein expression in breast fibroadenoma and carcinoma. Int J Cancer 2005; 114: 414-421. ArticlePubMed
- 5. Emig R, Magener A, Ehemann V, et al. Aberrant cytoplasmic expression of the p16 protein in breast cancer is associated with accelerated tumour proliferation. Br J Cancer 1998; 78: 1661-1668. ArticlePubMedPMCPDF
- 6. Han S, Ahn SH, Park K, et al. p16INK4a protein expression is associated with poor survival of the breast cancer patients after CMF chemotherapy. Breast Cancer Res Treat 2001; 70: 205-212. ArticlePubMed
- 7. Milde-Langosch K, Bamberger AM, Methner C, Rieck G, Löning T. Expression of cell cycle-regulatory proteins rb, p16/MTS1, p27/KIP1, p21/WAF1, cyclin D1 and cyclin E in breast cancer: correlations with expression of activating protein-1 family members. Int J Cancer 2000; 87: 468-472. ArticlePubMed
- 8. Kersting M, Friedl C, Kraus A, Behn M, Pankow W, Schuermann M. Differential frequencies of p16(INK4a) promoter hypermethylation, p53 mutation, and K-ras mutation in exfoliative material mark the development of lung cancer in symptomatic chronic smokers. J Clin Oncol 2000; 18: 3221-3229. ArticlePubMed
- 9. Kresty LA, Mallery SR, Knobloch TJ, et al. Alterations of p16(INK4a) and p14(ARF) in patients with severe oral epithelial dysplasia. Cancer Res 2002; 62: 5295-5300. PubMed
- 10. Lee JH, Park SJ, Abraham SC, et al. Frequent CpG island methylation in precursor lesions and early gastric adenocarcinomas. Oncogene 2004; 23: 4646-4654. ArticlePubMedPDF
- 11. Bearzatto A, Conte D, Frattini M, et al. p16(INK4A) hypermethylation detected by fluorescent methylation-specific PCR in plasmas from non-small cell lung cancer. Clin Cancer Res 2002; 8: 3782-3787. PubMed
- 12. Hsiung DT, Marsit CJ, Houseman EA, et al. Global DNA methylation level in whole blood as a biomarker in head and neck squamous cell carcinoma. Cancer Epidemiol Biomarkers Prev 2007; 16: 108-114. ArticlePubMedPDF
- 13. Kawakami K, Brabender J, Lord RV, et al. Hypermethylated APC DNA in plasma and prognosis of patients with esophageal adenocarcinoma. J Natl Cancer Inst 2000; 92: 1805-1811. ArticlePubMed
- 14. Lee TL, Leung WK, Chan MW, et al. Detection of gene promoter hypermethylation in the tumor and serum of patients with gastric carcinoma. Clin Cancer Res 2002; 8: 1761-1766. PubMed
- 15. Sharma G, Mirza S, Prasad CP, Srivastava A, Gupta SD, Ralhan R. Promoter hypermethylation of p16INK4a, p14ARF, cyclinD2 and Slit2 in serum and tumor DNA from breast cancer patients. Life Sci 2007; 80: 1873-1881. ArticlePubMed
- 16. Zhang YJ, Wu HC, Shen J, et al. Predicting hepatocellular carcinoma by detection of aberrant promoter methylation in serum DNA. Clin Cancer Res 2007; 13: 2378-2384. ArticlePubMedPDF
- 17. Zitt M, Zitt M, Müller HM. DNA methylation in colorectal cancer: impact on screening and therapy monitoring modalities? Dis Markers 2007; 23: 51-71. ArticlePubMedPMCPDF
- 18. Allred DC, Harvey JM, Berardo M, Clark GM. Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod Pathol 1998; 11: 155-168. PubMed
- 19. Fackler MJ, McVeigh M, Mehrotra J, et al. Quantitative multiplex methylation-specific PCR assay for the detection of promoter hypermethylation in multiple genes in breast cancer. Cancer Res 2004; 64: 4442-4452. ArticlePubMedPDF
- 20. Brenner AJ, Paladugu A, Wang H, Olopade OI, Dreyling MH, Aldaz CM. Preferential loss of expression of p16(INK4a) rather than p19(ARF) in breast cancer. Clin Cancer Res 1996; 2: 1993-1998. PubMed
- 21. Hui R, Macmillan RD, Kenny FS, et al. INK4a gene expression and methylation in primary breast cancer: overexpression of p16INK4a messenger RNA is a marker of poor prognosis. Clin Cancer Res 2000; 6: 2777-2787. PubMed
- 22. Nielsen NH, Roos G, Emdin SO, Landberg G. Methylation of the p16(Ink4a) tumor suppressor gene 5'-CpG island in breast cancer. Cancer Lett 2001; 163: 59-69. ArticlePubMed
- 23. Liu T, Niu Y, Feng Y, et al. Methylation of CpG islands of p16(INK4a) and cyclinD1 overexpression associated with progression of intraductal proliferative lesions of the breast. Hum Pathol 2008; 39: 1637-1646. ArticlePubMed
- 24. Bachmann IM, Straume O, Akslen LA. Altered expression of cell cycle regulators cyclin D1, p14, p16, CDK4 and Rb in nodular melanomas. Int J Oncol 2004; 25: 1559-1565. ArticlePubMed
- 25. Khouja MH, Baekelandt M, Nesland JM, Holm R. The clinical importance of Ki-67, p16, p14, and p57 expression in patients with advanced ovarian carcinoma. Int J Gynecol Pathol 2007; 26: 418-425. ArticlePubMed
- 26. Saegusa M, Machida BD, Okayasu I. Possible associations among expression of p14(ARF), p16(INK4a), p21(WAF1/CIP1), p27(KIP1), and p53 accumulation and the balance of apoptosis and cell proliferation in ovarian carcinomas. Cancer 2001; 92: 1177-1189. ArticlePubMed
- 27. Geradts J, Hruban RH, Schutte M, Kern SE, Maynard R. Immunohistochemical p16INK4a analysis of archival tumors with deletion, hypermethylation, or mutation of the CDKN2/MTS1 gene. A comparison of four commercial antibodies. Appl Immunohistochem Mol Morphol 2000; 8: 71-79. PubMed
- 28. Dublin EA, Patel NK, Gillett CE, Smith P, Peters G, Barnes DM. Retinoblastoma and p16 proteins in mammary carcinoma: their relationship to cyclin D1 and histopathological parameters. Int J Cancer 1998; 79: 71-75. ArticlePubMed
- 29. Geradts J, Wilson PA. High frequency of aberrant p16(INK4A) expression in human breast cancer. Am J Pathol 1996; 149: 15-20. PubMedPMC
- 30. Leong WF, Chau JF, Li B. p53 Deficiency leads to compensatory up-regulation of p16INK4a. Mol Cancer Res 2009; 7: 354-360. ArticlePubMedPDF
REFERENCES
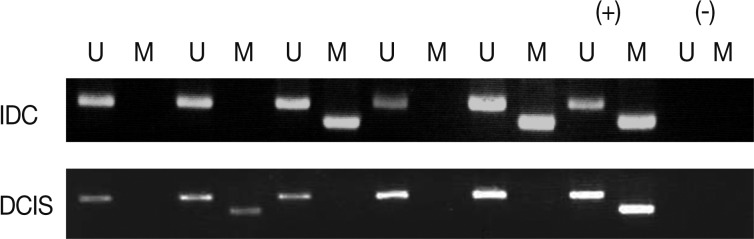

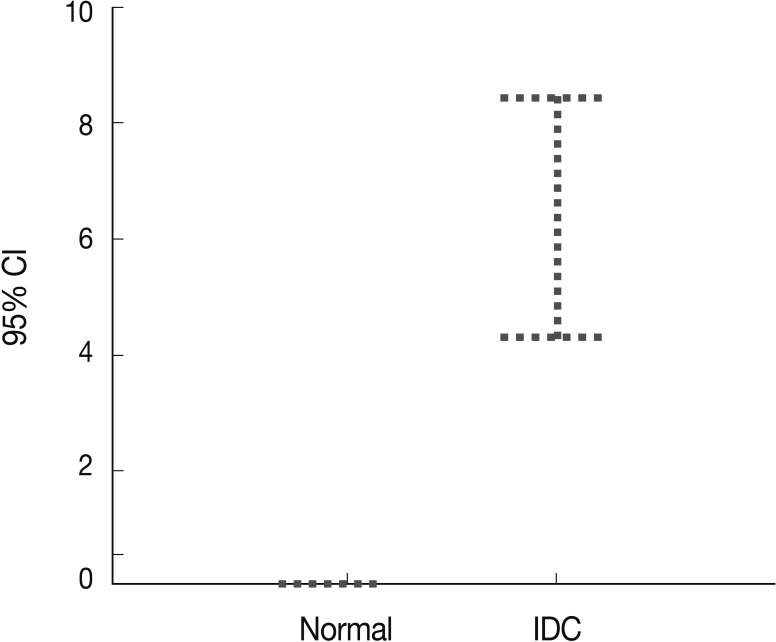
Figure & Data
References
Citations

- Progress in epigenetic research of breast cancer: a bibliometric analysis since the 2000s
Hua Yang, Yu Fang, Haijuan Wang, Ting Lu, Qihua Chen, Hui Liu
Frontiers in Oncology.2025;[Epub] CrossRef - Epigenetic Silencing of p16INK4a gene in Sporadic Breast Cancer
Satya P. Singh, Mallika Tewari, Alok K. Singh, Raghvendra R. Mishra, Hari S. Shukla
Indian Journal of Surgical Oncology.2023; 14(4): 822. CrossRef - Pathogenesis and Potential Therapeutic Targets for Triple-Negative Breast Cancer
Chia-Jung Li, Yen-Dun Tony Tzeng, Yi-Han Chiu, Hung-Yu Lin, Ming-Feng Hou, Pei-Yi Chu
Cancers.2021; 13(12): 2978. CrossRef - Mechanisms of resistance to estrogen receptor modulators in ER+/HER2− advanced breast cancer
Jin Zhang, Qianying Wang, Qing Wang, Jiangran Cao, Jiafu Sun, Zhengmao Zhu
Cellular and Molecular Life Sciences.2020; 77(4): 559. CrossRef - Aberrantly Methylated cfDNA in Body Fluids as a Promising Diagnostic Tool for Early Detection of Breast Cancer
Igor Stastny, Pavol Zubor, Karol Kajo, Peter Kubatka, Olga Golubnitschaja, Zuzana Dankova
Clinical Breast Cancer.2020; 20(6): e711. CrossRef - Epigenetic modulation of BRCA‐1 and MGMT genes, and histones H4 and H3 are associated with breast tumors
Parisa Paydar, Gholamreza Asadikaram, Hamid Zeynali Nejad, Hamed Akbari, Moslem Abolhassani, Vahid Moazed, Mohammad Hadi Nematollahi, Ghasem Ebrahimi, Hossein Fallah
Journal of Cellular Biochemistry.2019; 120(8): 13726. CrossRef - The 9p21 locus: A potential therapeutic target and prognostic marker in breast cancer
Mahdi Rivandi, Mohammad‐Sadegh Khorrami, Hamid Fiuji, Soodabeh Shahidsales, Malihe Hasanzadeh, Mir Hadi Jazayeri, Seyed Mahdi Hassanian, Gordon A. Ferns, Nafiseh Saghafi, Amir Avan
Journal of Cellular Physiology.2018; 233(7): 5170. CrossRef - p16INK4a overexpression as a predictor of survival in ocular surface squamous neoplasia
Sheetal Chauhan, Seema Sen, Anjana Sharma, Seema Kashyap, Radhika Tandon, Mandeep S Bajaj, Neelam Pushker, Murugesan Vanathi, Shyam S Chauhan
British Journal of Ophthalmology.2018; 102(6): 840. CrossRef - Liquid biopsy prediction of axillary lymph node metastasis, cancer recurrence, and patient survival in breast cancer
Ju-Han Lee, Hoiseon Jeong, Jung-Woo Choi, Hwa Eun Oh, Young-Sik Kim
Medicine.2018; 97(42): e12862. CrossRef - EZH2 inhibition sensitizes tamoxifen‑resistant breast cancer cells through cell cycle regulation
Si Chen, Fan Yao, Qinghuan Xiao, Qiannan Liu, Yikun Yang, Xuejuan Li, Guanglie Jiang, Takayoshi Kuno, Yue Fang
Molecular Medicine Reports.2017;[Epub] CrossRef - Aberrant promoter methylation of cancer-related genes in human breast cancer
Liang Wu, Ye Shen, Xianzhen Peng, Simin Zhang, Ming Wang, Guisheng Xu, Xianzhi Zheng, Jianming Wang, Cheng Lu
Oncology Letters.2016; 12(6): 5145. CrossRef - Centrosome aberrations in human mammary epithelial cells driven by cooperative interactions between p16INK4a deficiency and telomere-dependent genotoxic stress
Daniel Domínguez, Purificación Feijoo, Aina Bernal, Amaia Ercilla, Neus Agell, Anna Genescà, Laura Tusell
Oncotarget.2015; 6(29): 28238. CrossRef - Relationships Betweenp16Gene Promoter Methylation and Clinicopathologic Features of Colorectal Cancer: A Meta-Analysis of 27 Cohort Studies
Yan-Zhi Chen, Dan Liu, Yu-Xia Zhao, He-Tong Wang, Ya Gao, Ying Chen
DNA and Cell Biology.2014; 33(10): 729. CrossRef - Endocrine disruption of the epigenome: a breast cancer link
Kevin C Knower, Sarah Q To, Yuet-Kin Leung, Shuk-Mei Ho, Colin D Clyne
Endocrine-Related Cancer.2014; 21(2): T33. CrossRef - Cyclin-dependent kinase inhibitors, p16 and p27, demonstrate different expression patterns in thymoma and thymic carcinoma
Mutsuko Omatsu, Toshiaki Kunimura, Tetsuya Mikogami, Akira Shiokawa, Atsuko Masunaga, Tomoko Nagai, Akihiko Kitami, Takashi Suzuki, Mitsutaka Kadokura
General Thoracic and Cardiovascular Surgery.2014; 62(11): 678. CrossRef - Limoniastrum guyonianum aqueous gall extract induces apoptosis in human cervical cancer cells involving p16INK4A re-expression related to UHRF1 and DNMT1 down-regulation
Mounira Krifa, Mahmoud Alhosin, Christian D Muller, Jean-Pierre Gies, Leila Chekir-Ghedira, Kamel Ghedira, Yves Mély, Christian Bronner, Marc Mousli
Journal of Experimental & Clinical Cancer Research.2013;[Epub] CrossRef - Diagnostic and prognostic value of circulating tumor-related DNA in cancer patients
Diego M Marzese, Hajime Hirose, Dave S B Hoon
Expert Review of Molecular Diagnostics.2013; 13(8): 827. CrossRef - Epigallocatechin-3-gallate and trichostatin A synergistically inhibit human lymphoma cell proliferation through epigenetic modification of p16INK4a
DAN-SEN WU, JIAN-ZHEN SHEN, AI-FANG YU, HAI-YING FU, HUA-RONG ZHOU, SONG-FEI SHEN
Oncology Reports.2013; 30(6): 2969. CrossRef
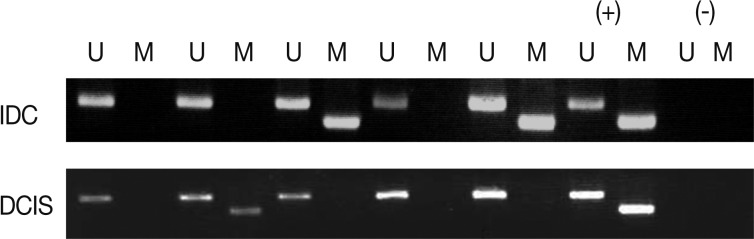
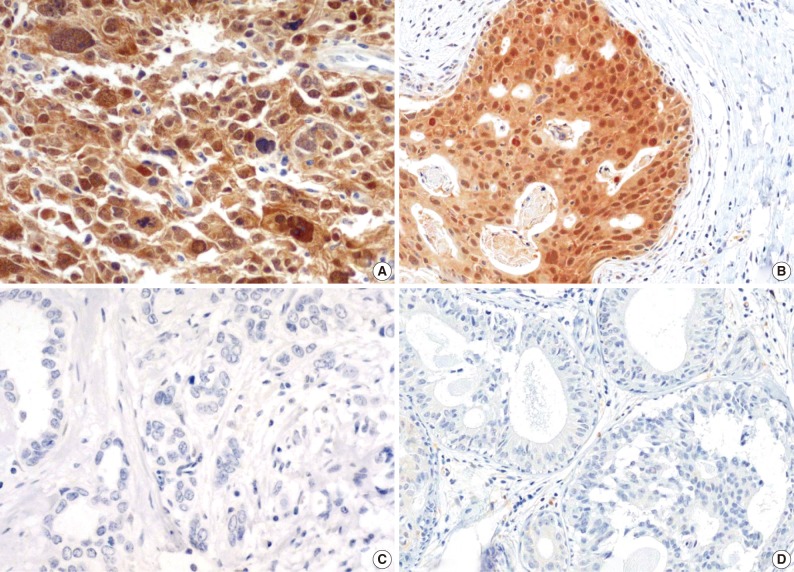
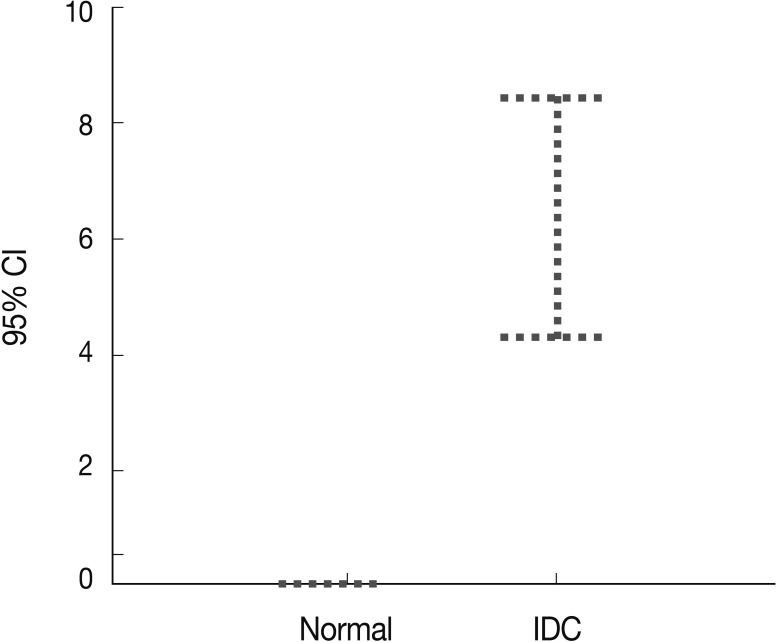
Fig. 1
Fig. 2
Fig. 3



Values are presented as number (% or range). ER, estrogen receptor; PR, progesterone receptor; TNM, tumor-node-metastasis.
Values are presented as number (%). IDC, invasive ductal carcinoma; NS; not significant; DCIS, ductal carcinoma
NS, not significant; ER, estrogen receptor; PR, progesterone receptor.

 E-submission
E-submission
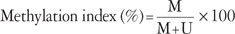

 PubReader
PubReader Cite this Article
Cite this Article




