Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 55(6); 2021 > Article
-
Newsletter
What’s new in molecular genetic pathology 2021: solid tumors and NGS panel selection -
Guoli Chen
 , Patricia C. Tsang
, Patricia C. Tsang
-
Journal of Pathology and Translational Medicine 2021;55(6):421-422.
DOI: https://doi.org/10.4132/jptm.2021.09.01
Published online: November 12, 2021
Geisinger Medical Laboratories, Geisinger Health, Danville and Wilkes-Barre, PA, USA
- Corresponding Author: Patricia C. Tsang, MD Geisinger Medical Laboratories, Geisinger Health, Wilkes-Barre, PA, USA E-mail: patctsang@hotmail.com
- This article has been published jointly, with consent, in both Journal of Pathology and Translational Medicine and PathologyOutlines.com.
© 2021 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 5,559 Views
- 212 Download
Abstract
- The linchpin of precision medicine is molecular genetic and genomic testing. Molecular biomarkers are important for establishing precise diagnoses and for predicting therapeutic responses that enable cancer patients to receive personalized and targeted treatment. Below are highlights of the current considerations in next generation sequencing (NGS) panel selection, and in molecular testing of solid tumors of the lung, digestive system, thyroid and soft tissue.
- • The use of targeted NGS panels (comprising up to several hundred genes) is more common than whole exome sequencing (WES) in clinical practice. Pros and cons of WES or large targeted panels compared to small hotspot panels include the following:
-
- Comprehensive tumor profiling maximizes the chances of uncovering clinically relevant alterations.
- - They require more tumor tissue, cost more for data analysis and storage, and entail more complex interpretation.
- • FDA-approved liquid biopsy using circulating cell-free DNA is an excellent tool when tumor tissue is inaccessible or limited [1]. While liquid biopsy has a faster turnaround time, it has a higher false-negative rate than standard biopsy NGS.
NEXT GENERATION SEQUENCING PANEL SELECTION
- • Molecular profiling is used to predict therapeutic efficacies in NSCLCs and typically includes EGFR, KRAS, ROS1, ALK, MET, BRAF, RET, ERBB2 (HER2), and NTRK [2]. Also useful is PD-L1 (programmed cell death ligand-1) by immunohistochemistry (IHC) to predict response to immune checkpoint inhibitors.
- • EGFR mutations involve 10%-35% of lung adenocarcinoma in Western populations (and as high as 50% in Asian populations) and 3% of lung squamous cell carcinoma (SqCC). Mutations can predict therapeutic response to EGFR-specific tyrosine kinase inhibitors (TKIs).
-
- Acquired resistance inevitably develops, mostly commonly due to EGFR T790M missense mutation, accounting for ~50% of acquired resistance mutations. FDA-approved EGFR-inhibiting drugs that specifically target T790M or exon 20 insertion mutation are available.
- - MET amplification (by FISH or NGS) is responsible for 5%-20% of anti-EGFR resistance.
- • KRAS mutations are detectable in 25%-35% of lung adenocarcinoma, and 5% of SqCC. Activating mutations in codons 12 and 13, and, less commonly, codon 61, predict unfavorable prognosis and EGFR-TKI resistance. A KRAS inhibitor drug that specifically targets the G12C missense mutation (seen in ~13% of NSCLCs) is available.
- • ALK activating rearrangements occur in ~5% of lung adenocarcinoma, particularly in young non-smokers. The most common fusion gene product is EML4-ALK, which predicts response to ALK inhibitors. FISH is the gold standard for testing, but IHC, RT-PCR and NGS can also be used.
- • ROS1 gene fusion (1%-2% of NSCLCs) signifies response to certain ALK inhibitors due to homology between the rearranged ROS1 and ALK genes.
- • MET exon 14 skipping mutations, BRAF V600E mutation, RET gene fusion and NTRK gene fusion can predict therapeutic response to their respective inhibitors.
NON-SMALL CELL LUNG CARCINOMA (NSCLC)
- • EGFR targeted treatment is effective in the absence of KRAS and NRAS mutations. Mutational analyses of KRAS and NRAS genes should include codons 12, 13, 59, 61, 117 and 146. Other drug resistance biomarkers include BRAF mutations and PIK3CA/PTEN deregulation.
- • Despite its low prevalence in CRC (2%–3%), ERBB2 (HER2) amplification is emerging as a potential therapeutic target [3]. Furthermore, activation of ERBB2 (HER2) signaling causes resistance to anti-EGFR therapy in a subset of patients with metastatic CRC.
- • NTRK fusions can be evaluated in CRC due to the availability of targeted therapies.
- • FGFR gene fusion is detected in ~10% of intrahepatic cholangiocarcinoma for which two FGFR-targeted tyrosine kinase inhibitors have been approved - pemigatinib and infigratinib.
COLORECTAL CARCINOMA (CRC) AND CHOLANGIOCARCINOMA
- • Papillary thyroid carcinoma may harbor BRAF mutations (especially V600E), TERT promoter mutations, and, less commonly, RET/PTC1/2/3 rearrangements and RAS mutations.
- • PAX8-PPARγ fusion, RAS mutations, and TERT mutations are enriched in follicular thyroid carcinoma, while BRAF V600E is unusual. RAS mutations are also seen in a minority of benign follicular adenomas.
- • More than 60% of medullary thyroid carcinomas show somatic RET mutations.
- • Anaplastic thyroid carcinomas can harbor TERT, TP53 and/or BRAF mutations.
- • Preoperative molecular testing is valuable in thyroid nodules with indeterminate FNA cytology. The gene panel typically includes TERT, BRAF, PAX8/PPARγ, RAS, RET/PTC and TP53. Below are three commonly used commercial platforms:
-
- ThyroseqTM involves next-generation DNA and RNA sequencing of 112 genes to stratify thyroid nodules as likely benign or likely malignant.
- - Afirma® Gene Sequencing Classifier is an RNA-based test. Similar to ThyroSeqTM, it is mostly a rule-out assay with acceptable rule-in capability.
- - ThyGeNEXT®/ThyraMIR® uses a combination of two tests. If no mutation is found in the first panel by DNA and RNA sequencing, another test is performed using micro-RNA expression.
THYROID CARCINOMA
- • Molecular testing has led to the discovery of new mesenchymal tumor entities. For example, CIC-DUX4 sarcoma is a recently described small round blue cell tumor associated with more aggressive disease than Ewing sarcoma with characteristic EWSR1-FLI1 fusion. Immunohistochemical detection of ETV4, a transcriptional target of CIC-DUX4, is a useful diagnostic tool.
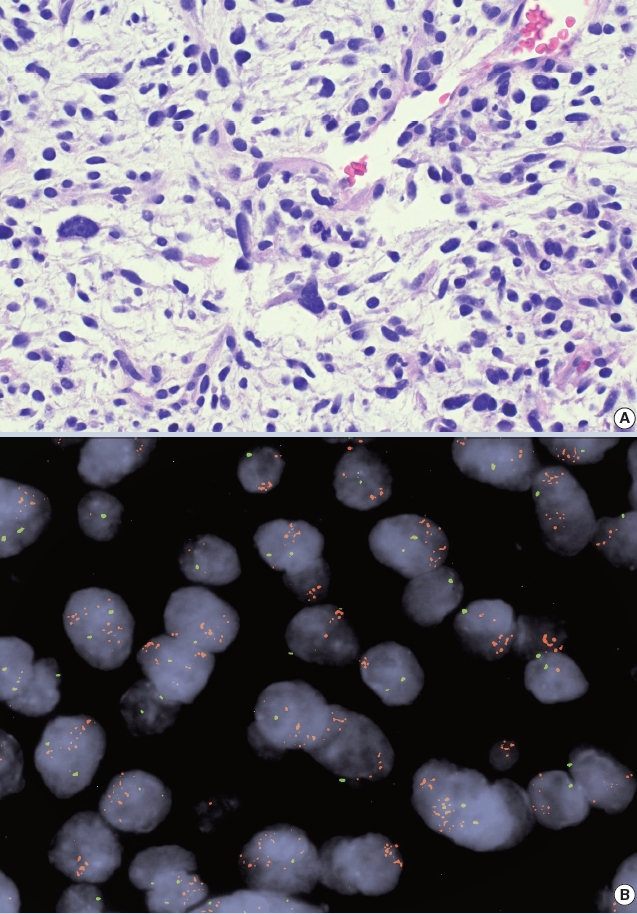
- • Amplification of the MDM2 gene detectable by IHC/FISH/NGS is helpful for confirming a diagnosis of atypical lipomatous tumor/ well-differentiated liposarcoma (~93%) or dedifferentiated liposarcoma (~97%). The oncogenicity of MDM2 is related to TP53 inactivation (Fig. 1).
- • Gene fusion testing by FISH or NGS is helpful for classifying rhabdomyosarcomas [4].
-
- Recurrent gene fusions are detected in ~80% of alveolar rhabdomyosarcoma (ARMS), and involve mostly the FOXO1 gene fusing with either PAX3 or PAX7. Fusion positive ARMS is associated with a worse prognosis than fusion negative ARMS.
- - Embryonal rhabdomyosarcoma (ERMS) characteristically lacks gene fusions, but shows chromosomal losses/gains and gene mutations. ERMS is similar to fusion negative ARMS in clinical behavior.
- - Most congenital/infantile spindle cell and sclerosing rhabdomyosarcomas show NCOA2 in addition to VGLL2 gene fusions. This entity is associated with a favorable prognosis. However, patients who harbor MYOD1 mutations tend to have aggressive disease.
BONE AND SOFT TISSUE TUMORS
- Dr. Chen is an Associate and Attending pathologist at Geisinger Medical Center, where he participates in clinical, academic and teaching services in surgical pathology and molecular diagnostics. He has actively published and lectured on topics related to molecular pathology of solid tumors.
- Dr. Tsang joined the PathologyOutlines editorial board in 2019 and has been its Deputy Editor-in-Chief for Clinical Pathology since 2020. She is a Clinical Associate Professor and Laboratory Medical Director in Geisinger's Northeast Region and has been a practicing molecular pathologist for more than a decade.
Meet the Authors
- 1. Ignatiadis M, Sledge GW, Jeffrey SS. Liquid biopsy enters the clinic: implementation issues and future challenges. Nat Rev Clin Oncol 2021; 18: 297-312. ArticlePubMedPDF
- 2. Imyanitov EN, Iyevleva AG, Levchenko EV. Molecular testing and targeted therapy for non-small cell lung cancer: current status and perspectives. Crit Rev Oncol Hematol 2021; 157: 103194.ArticlePubMed
- 3. Siena S, Di Bartolomeo M, Raghav K, et al. Trastuzumab deruxtecan (DS-8201) in patients with HER2-expressing metastatic colorectal cancer (DESTINY-CRC01): a multicentre, open-label, phase 2 trial. Lancet Oncol 2021; 22: 779-89. ArticlePubMed
- 4. Giannikopoulos P, Parham DM. Rhabdomyosarcoma: how advanced molecular methods are shaping the diagnostic and therapeutic paradigm. Pediatr Dev Pathol 2021; 24: 395-404. ArticlePubMedPDF
REFERENCES
Figure & Data
References
Citations

 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
- Related articles
-
- What's new in molecular genetic pathology 2026: emerging biomarkers for personalized cancer therapies
- What’s new in hematopathology 2025: myeloid neoplasms in the WHO 5th edition and ICC
- What’s new in neuropathology 2024: CNS WHO 5th edition updates
- What’s new in thyroid pathology 2024: updates from the new WHO classification and Bethesda system
- What’s new in genitourinary pathology 2023: WHO 5th edition updates for urinary tract, prostate, testis, and penis

 E-submission
E-submission



