Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 54(6); 2020 > Article
-
Original Article
Evaluation of human papillomavirus (HPV) prediction using the International Endocervical Adenocarcinoma Criteria and Classification system, compared to p16 immunohistochemistry and HPV RNA in-situ hybridization -
Hezhen Ren1
 , Jennifer Pors1
, Jennifer Pors1 , Christine Chow2
, Christine Chow2 , Monica Ta2
, Monica Ta2 , Simona Stolnicu3
, Simona Stolnicu3 , Robert Soslow4
, Robert Soslow4 , David Huntsman1,5
, David Huntsman1,5 , Lynn Hoang1,2,6
, Lynn Hoang1,2,6
-
Journal of Pathology and Translational Medicine 2020;54(6):480-488.
DOI: https://doi.org/10.4132/jptm.2020.07.18
Published online: August 31, 2020
1Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, BC, Canada
2Genetic Pathology Evaluation Center (GPEC), Vancouver, BC, Canada
3Department of Pathology, University of Medicine, Pharmacy, Sciences and Technology of Targu Mures, Targu Mures, Romania
4Department of Pathology, Memorial Sloan Kettering Cancer Center, New York, NY, USA
5Molecular Oncology, British Columbia Cancer Research Centre, Vancouver, BC, Canada
6Department of Anatomical Pathology, Vancouver General Hospital, Vancouver, BC, Canada
- Corresponding Author: Lynn Hoang, MD, Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver General Hospital, 1215-910 West 10th Avenue, Vancouver, BC V5Z 1M9, Canada Tel: +1-604-875-4731, Fax: +1-604-875-4497, E-mail: Lien.Hoang@vch.ca
© 2020 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- The International Endocervical Adenocarcinoma Criteria and Classification (IECC) separated endocervical adenocarcinomas into human papillomavirus (HPV) associated (HPVA) and non–HPV-associated (NHPVA) categories by morphology alone. Our primary objective was to assess the accuracy of HPV prediction by the IECC system compared to p16 immunohistochemistry and HPV RNA in-situ hybridization (RISH). Our secondary goal was to directly compare p16 and HPV RISH concordance.
-
Methods
- Cases were classified by IECC and stained for p16 and HPV RISH on tissue microarray, with discordant p16/HPV RISH cases re-stained on whole tissue sections. Remaining discordant cases (p16/HPV, IECC/p16, IECC/HPV discordances) were re-reviewed by the original pathologists (n = 3) and external expert pathologists (n = 2) blinded to the p16 and HPV RISH results. Final IECC diagnosis was assigned upon independent agreement between all reviewers.
-
Results
- One hundred and eleven endocervical adenocarcinomas were classified originally into 94 HPVA and 17 NHPVA cases. p16 and HPV RISH was concordant in 108/111 cases (97%) independent of the IECC. HPV RISH and p16 was concordant with IECC in 103/111 (93%) and 106/111 (95%), respectively. After expert review, concordance improved to 107/111 (96%) for HPV RISH. After review of the eight discordant cases, one remained as HPVA, four were reclassified to NHPVA from HPVA, two were unclassifiable, and one possibly represented a mixed usual and gastric-type adenocarcinoma.
-
Conclusions
- p16 and HPV RISH have excellent concordance in endocervical adenocarcinomas, and IECC can predict HPV status in most cases. Focal apical mitoses and apoptotic debris on original review led to the misclassification of several NHPVA as HPVA.
- Selection criteria and tumor classification
- All ECAs diagnosed on resection (hysterectomy or trachelectomy) were identified from the archives of Vancouver General Hospital. Biopsies were excluded. Cases with prior neoadjuvant chemoradiation rendering histologic assessment difficult were excluded. Additional specimens from the same patient were excluded. Slides of all ECAs were originally reviewed and reclassified under the IECC criteria (L.H., J.P., and H.R.). The observers had variable expertise (junior attending, senior resident, junior resident), and the IECC assignment was made with all observers in agreement. As per the IECC, ECAs were assigned as HPVA based on the presence of apical mitotic features and apoptotic bodies present at scanning magnification. The HPVA ECAs were further substratified into usual-type, villoglandular-type, mucinous intestinal-type, mucinous signet ring-type, invasive stratified mucin-producing intraepithelial lesion (iSMILE), and mucinous not otherwise specified (NOS), as defined previously [11,31]. When these features were absent, the slides were reexamined at ×200. Cases were classified as NHPVA if HPVA features were absent. The NHPVA ECAs were further substratified into gastric-type, clear cell, mesonephric, endometrioid, and serous carcinoma based on their morphological features.
- Tissue microarray construction
- A tissue microarray (TMA) was constructed using formalin fixed paraffin embedded blocks. At least two 0.6 mm cores were taken from each case using the Pathology Devices TMArrayer (Pathology Devices, San Diego, CA, USA).
- IHC and RNA in-situ hybridization
- p16 (clone E6H4, mouse monoclonal, 1:5 dilution, Roche Diagnostics, Laval, Canada) IHC was performed on 4-µm sections using the Dako Omnis (Agilent, Santa Clara, CA, USA) and Dako EnVision FLEX+ detection system as per manufacturer recommendations. Sections were mounted onto Dako FLEX microscope slides, air-dried for 20 minutes and baked at 60°C for 20 minutes. The heat-induced antigen retrieval method was performed using Envision FLEX Target Retrieval Solution (Dako) in Dako PT Link.
- HPV RISH was performed using the Advanced Cell Diagnostics (ACD) RNAScope 2.5 assay (Cat# 322300 ACD, Advanced Cell Diagnostics, Newark, NJ, USA) with RNAscope Probe HPV-HR18 (Cat#312591 ACD) which targets E6 and E7 mRNA for 18 high-risk HPV subtypes (HPV 16, 18, 26, 31, 33, 35, 39, 45, 51, 52, 53, 56, 58, 59, 66, 68, 73, and 82), as per manufacturer instructions [11]. In brief, 4-µm sections were baked at 60°C and air-dried. The slides were briefly treated with hydrogen peroxide for 10 minutes and then underwent target retrieval for 15 minutes and protease treatment for 30 minutes. Hybridization with probe was done at 40°C for 2 hours followed by series of signal amplification. Signal detection was achieved through DAB staining before counterstaining with hematoxylin. Both RNAscope Positive Control Probe (Hs-PPIB, Cat# 313901 ACD) and RNAscope Negative Control Probe (DapB, Cat#310043 ACD) were used on separate sections for quality control purposes.
- p16 IHC and HPV RISH were repeated on whole 4-µm sections when there was equivocal staining or when there was a discrepancy between p16 and HPV RISH scores on TMA. Cases not within the TMA were stained using whole sections directly.
- p16 IHC and HPV RISH scoring
- p16 IHC was scored positive only when there was strong and continuous diffuse staining in neoplastic glands (nuclear or nuclear and cytoplasmic) as previously established [32]. Absent, weak, or patchy staining was scored as negative. HPV RISH was scored positive only when there was dot-like nuclear staining as detailed previously [20].
- Review of discordant cases
- All cases which had discordance (either p16 IHC-HPV RISH, IECC-HPV RISH, IECC-p16 IHC) underwent re-review by the original institutional pathologists as well as by two external expert pathologists (R.S. and S.S.) who were blinded to the p16 IHC and HPV RISH findings. The final IECC diagnostic designation was determined when all observers (original pathologists and two expert pathologists) completely agreed on the diagnosis. Any cases without complete agreement were considered unclassifiable.
MATERIALS AND METHODS
- Case selection
- A total of 129 ECAs were initially acquired. After exclusion of cases with biopsy specimen only, neoadjuvant therapy, insufficient tissue, or repeats, 111 cases were included in the study. The 111 cases grouped by the IECC included: 94 HPVA cases (63 usual-type, 4 intestinal, 4 villoglandular, 1 iSMILE, and 22 mucinous NOS) and 17 NHPVA cases (12 gastric, 4 mesonephric, and 1 NOS).
- Eighteen cases in the tissue microarray had both p16 IHC and HPV RISH repeated on whole tissue sections. Twelve of 18 cases (66%) had a change in their p16 IHC and/or HPV RISH status after repeat staining on whole tissue sections. Of these 12 cases, six (50%) were positive for p16 IHC and negative for HPV RISH on TMA but showed positive staining for both p16 IHC and HPV RISH on whole sections. Four cases (33%) had absent or necrotic tumor cells on both p16 IHC and HPV RISH TMAs and showed positive staining for both p16 IHC and HPV RISH on whole sections. Two cases (17%) had positive p16 IHC on TMA but no tumor cells on HPV RISH TMA, and whole sections confirmed HPV RISH positivity. Three cases were not included in the construction of the TMA initially, and were stained using only whole sections.
- Assessment of p16 IHC, HPV RISH, and IECC accuracy on original review
- On initial assessment, p16 IHC and HPV RISH were concordant in 108/111 cases (97%) independent of the IECC assignment Eighty-eight cases were positive for both p16 IHC and HPV RISH, and 20 cases were negative for both p16 IHC and HPV RISH. The three cases that were discordant were all p16 positive and HPV RISH negative. These three cases are discussed in detail below.
- On original review, the IECC assignment was concordant with HPV RISH in 103/111 cases (93%) and with p16 IHC in 106/111 cases (95%) (Table 1). Altogether, there were a total of eight cases with p16 IHC/HPV RISH and/or IECC/HPV RISH and/or IECC/p16 discordance. These eight discordant cases included seven HPVA cases (one usual-type, one mucinous intestinal-type, and five mucinous NOS [3 are mentioned above in the preceding paragraph]) that were all found to be HPV RISH negative. There was one NHPVA case (gastric-type) that was found to be HPV RISH positive, which is described in detail below.
- Histology re-examination with by original reviewers and by external expert reviewers
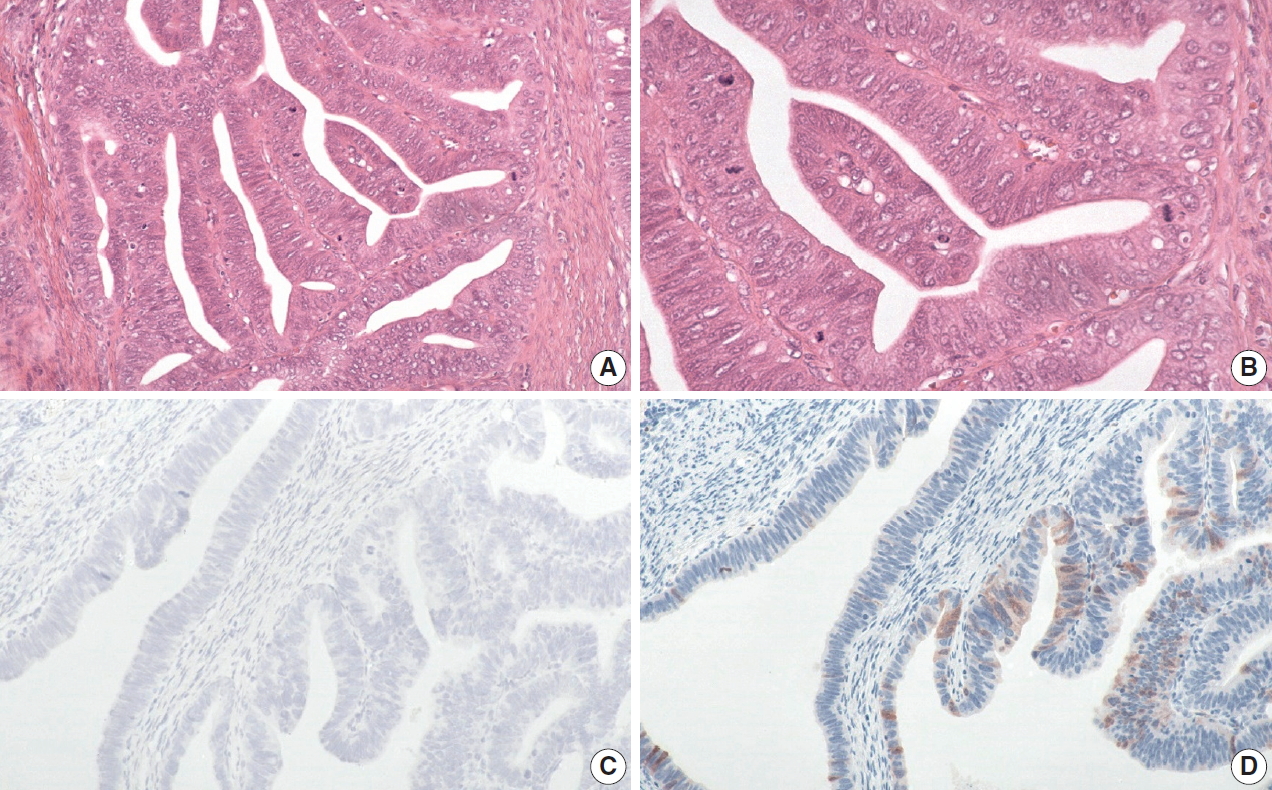
- The histology of each of the eight discordant cases was reviewed again by the original internal reviewers as well as independently by two expert external reviewers (Table 2). Histology review of the one discordant usual-type ECA demonstrated classic HPVA features with apoptotic bodies and floating mitosis. The two external reviewers also independently thought it was HPVA. Its classification remained unchanged as HPVA usual-type ECA, despite the negativity for both p16 IHC and HPV RISH (Fig. 1).
- One case initially called HPVA-mucinous intestinal-type, showed an ECA with ample eosinophilic cytoplasm with well-defined cell borders, with histologic features in keeping with NHPVA-gastric type ECA. Focally, the presence of goblet cells was identified, which was the initial rationale for classifying it as a HPVA-mucinous intestinal-type. Upon review, the intestinal-type ECA was reclassified as gastric-type ECA. The two external reviewers also called this NHPVA.
- Of the five HPVA-mucinous NOS cases, two were negative for both p16 IHC and HPV RISH. On reassessment of both cases, there was only very focal apoptotic bodies and apical mitoses present. There were also other areas in the same tumor which showed ample intracytoplasmic mucin. Both cases were reclassified as a NHPVA-gastric type. The two external reviewers also agreed independently on the diagnosis.
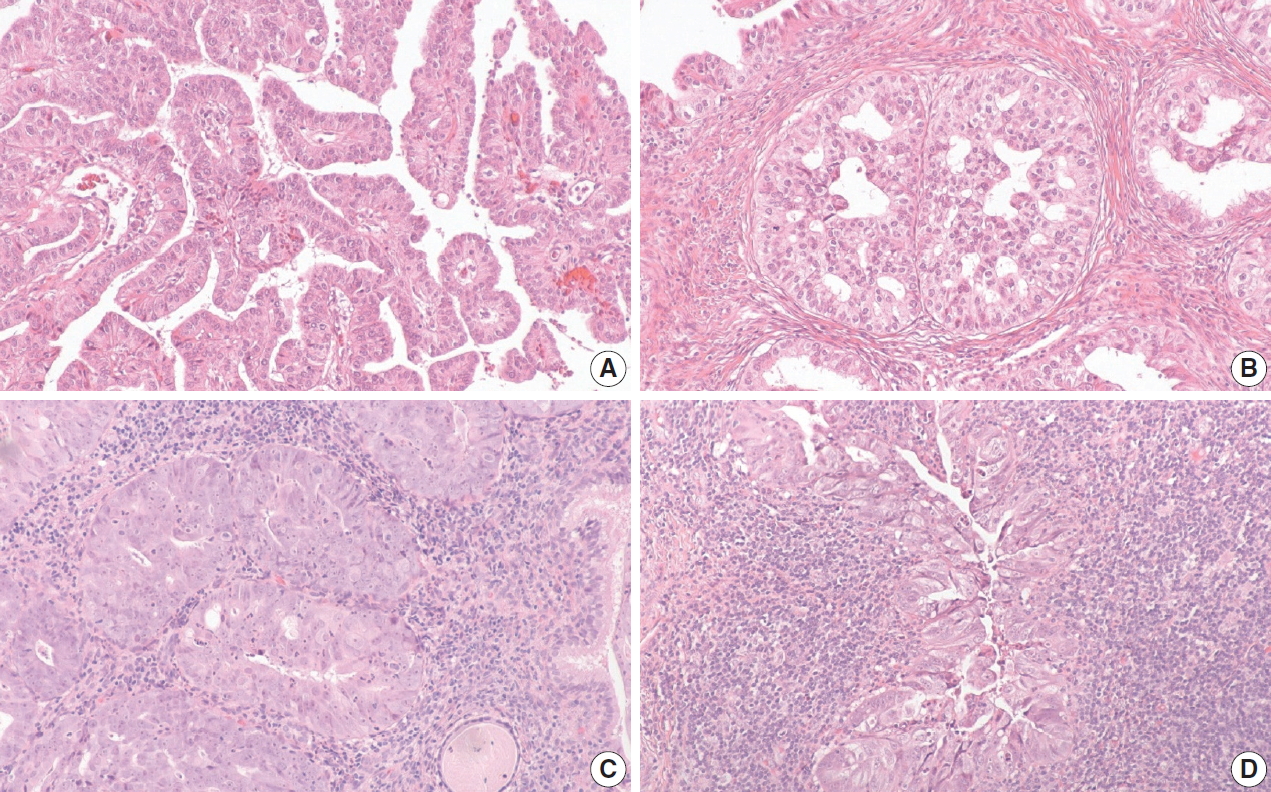
- Of the five HPVA-mucinous NOS cases, three were positive for p16 IHC and negative for HPV RISH. These cases were difficult to reclassify even after external reviews. One case exhibited villoglandular and glandular architecture. There were admixed columnar cells with vague mucinous cytoplasm and cuboidal cells forming small papillae (Fig. 2A, B). While all reviewers agreed that this case belonged in the NHPVA category, the precise histotype could not be agreed upon by morphology alone, yielding a final designation of NHPVA-NOS. The second case showed mostly solid nested growth and the cells exhibited severe cytologic atypia and admixed neutrophils. The nuclei were enlarged with conspicuous nucleoli. There was no agreement in histotype and this case was deemed unclassifiable (Fig. 2C). IHC performed in the clinical pathology workup showed positivity for estrogen receptor and negativity for monoclonal carcinoembryonic antigen and vimentin. The third case was similar. It comprised of nests of mucinous cells with severe cytologic atypia. There was a background of dense lymphoplasmacytic inflammation. This case was also deemed unclassifiable (Fig. 2D). No additional IHC was performed in the clinical pathology workup; however, this patient did have a history of cervical intraepithelial neoplasia 2 and adenocarcinoma-in-situ on a prior loop electrosurgical excision procedure specimen.
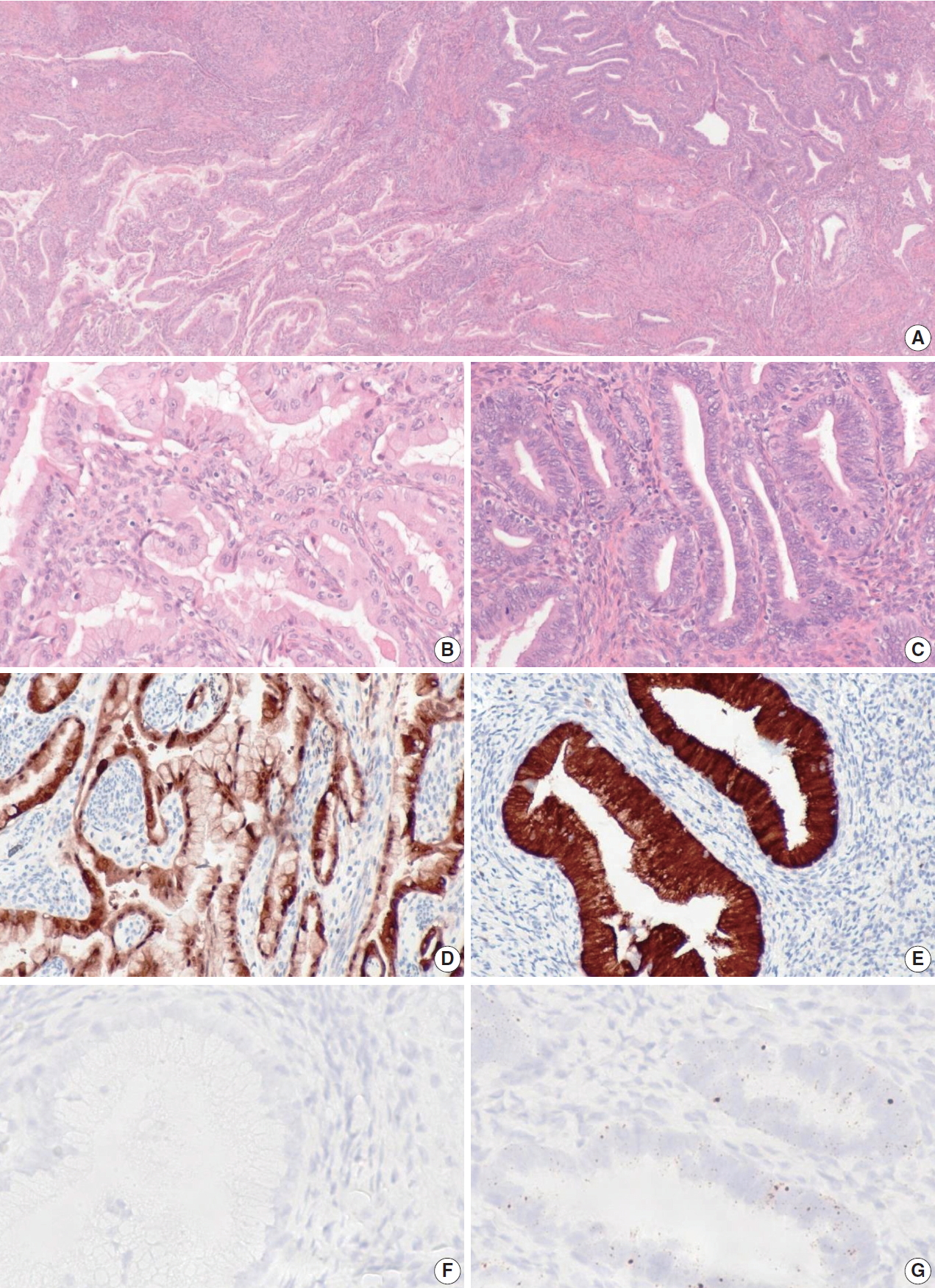
- When the one discordant NHPVA-gastric type ECA was rereviewed, the histology demonstrated intermixed gastric-type and usual-type morphology, which were spatially distinct (Fig. 3). The two histotypes showed positive p16 IHC but a distinct HPV RISH profile. The usual-type component was positive for HPV RISH and more superficial, while the gastric-type component was negative for HPV RISH and more deep and infiltrative. Therefore, this case was reclassified as an ECA of mixed type. This patient presented with International Federation of Gynecology and Obstetrics stage IIB disease and was treated with adjuvant chemotherapy and radiation. Follow up after 5 years revealed no evidence of disease recurrence.
RESULTS
- In our large series of cervical adenocarcinomas, we found that p16 IHC is concordant with HPV RISH in 97% of cases and is an excellent surrogate biomarker for high-risk HPV. On original review, p16 IHC and HPV RISH was concordant with IECC in 106/111 (95%) and 103/111 (93%) of cases, which remained unchanged for p16 IHC and improved to 107/111 (96%) for HPV RISH after expert review. Our study reiterates that the morphological guidelines denoted by the IECC are robust and can be relied on as the first step in reaching the correct ECA classification in the vast majority of cases.
- Our results are in keeping with recent findings by Chen et al. [27], who demonstrated 100% sensitivity of p16 IHC for HPV RISH in HPVA ECAs, but demonstrated a lower specificity (88.89%). We also found 100% sensitivity of p16 IHC for HPV RISH, and a lower specificity (86.9%). Our p16 IHC and IECC concordance is also similar to that presented previously by Hodgson et al. [10] (92% in HPVA, 100% in NHPVA). However, their study demonstrated a higher false negative rate for p16 IHC in HPVA (5 of 63, 8% of cases). We only saw false-negative p16 IHC in one case (1%). The difference between our concordance rate is likely because any discordance in our study prompted staining of p16 on whole tissue sections.
- It has been shown that p16 IHC can be positive in up to one-third of NHPVA-gastric type, which is a pitfall for pathologists [31]. In our series, our one hybrid ECA showed diffuse p16 IHC staining in the gastric-type component, two cases of gastric-type ECA showed patchy staining and the remainder of the gastrictype ECA showed negligible staining. We did not actually encounter a high rate of true p16 IHC positivity in our NHPVA-gastric cases. However, we did have three p16 IHC+/HPV RISH–cases which were reclassified after expert review as one NHPVA-NOS and two unclassifiable adenocarcinomas.
- The HPVA-mucinous group posed the greatest challenge on our initial histologic assessment. Five HPVA-mucinous were reclassified after review (2 into NHPVA-gastric, 1 into NHPVA-NOS, and 2 were unclassifiable). In some of the cases, the low-power architecture (villoglandular), mucin-deplete columnar cells and goblet cells, prompted an initial diagnosis of HPVA. More often, it was the finding of very focal areas of luminal mitotic figures or apoptotic bodies that prompted a diagnosis of HPVA. The original reviews were performed by pathologists with varying levels of training and we believe this was a reflection of experience. Novice pathologists appear to take a more prescriptive approach, following the IECC rules very literally, such that any evidence of luminal mitoses or apoptotic debris renders a diagnosis of HPVA. The more seasoned pathologists appear to take a broader approach, incorporating the entire constellation of morphologic features and only considering luminal mitoses and apoptotic bodies when they are more conspicuous. This is an important finding to emphasize as many pathologists new to the IECC system may encounter this same problem. In practice, more time would be spent on each case assessing for these features as well as any adjacent in-situ lesions (adenocarcinoma in situ, lobular endocervical glandular hyperplasia) that would be informative for real-life practice. Even after careful review, some mucinous NOS cases were not classifiable, albeit a very small number (2/111, 2% of cases). In practice, additional immunohistochemical markers available in the clinical pathology laboratory (such as estrogen receptor, Napsin, p53) would likely allow for a refined diagnosis in these cases. For example, one of the unclassified cases had limited IHC workup which was positive for estrogen receptor and negative for vimentin and monoclonal carcinoembryonic antigen. It was subsequently signed out as invasive poorly differentiated adenocarcinoma, consistent with cervical primary, favor endometrioid.
- Using morphology alone, we have identified one hybrid case with both NHPVA and HPVA features. Like the 13 cases reported by Wada et al. [33], the hybrid case contained features of both gastric and usual-type ECA. However, the cases presented in their study demonstrated either positive HPV or negative HPV in both gastric and usual-type components using HPV DNA ISH. Using HPV RISH, we showed a different staining pattern in the two components. Although a collision tumor is a possibility, we hypothesize that this tumor started as an HPV driven neoplasm which then gained additional mutations allowing for oncogenesis independent of HPV, with a subclone of the tumor presenting as loss of HPVA and gain of NHPVA morphologic features. The favorable prognosis after 5 years also supports an HPVA origin.
- There are a couple of minor limitations of this study to highlight. First, the HPV RISH kit detects 18 high-risk HPV subtypes, but more than 200 types of HPV have been recognized [34]. This could explain why the one usual-type ECA was negative for HPV RISH. Second, although p16 IHC is a robust surrogate marker for HPV infection, p16 IHC can be falsely negative in HPVA tumors in the setting of methylation induced-inactivation and allelic loss [35,36]. We did not assess for these advanced molecular features.
- In summary, p16 IHC and HPV RISH have excellent concordance in endocervical adenocarcinomas. IECC classification demonstrated strong concordance with HPV testing using p16 IHC and HPV RISH, with HPV RISH being slightly superior to p16. Classification by IECC was excellent in novice pathologists, with only a slight improvement with the addition of expert pathologists. Of the IECC/HPV RISH or IECC/p16 IHC discordant cases, the majority were mucinous (i.e., mucinous intestinal-type, mucinous NOS and gastric-types). Therefore, in any cases with mucinous cytoplasm, pathologists should be extra careful to examine for conspicuous HPV-related histologic features, along with any associated in-situ lesions. The presence of goblet cells or very focal apoptotic bodies or luminal mitoses are not enough to designate a case as HPVA. Ambiguous cases may warrant referral for HPV RISH or PCR.
DISCUSSION
Ethics Statement
All procedures performed in the current study were approved by the Institutional Review Board (IRB) (H18-01652, October 1, 2018) in accordance with the 1964 Helsinki declaration and its later amendments. Formal written informed consent was not required with a waiver by the appropriate IRB.
Author contributions
Conceptualization: LH, DH. Data curation: HR, JP. Formal analysis: HR, LH. Investigation: CC, MT. Methodology: CC, MT. Supervision: LH, DH. Validation: LH. Visualization: HR, LH. Writing—original draft: HR, LH. Writing—review & editing: SS, RS, LH. Approval of final manuscript: all authors.
Conflicts of Interest
The authors declare that they have no potential conflicts of interest.
Funding Statement
This study has supported by the Carraressi Foundation, Sumiko Kobayashi Marks Memorial Fund, OVCare and the UBC & VGH Hospital Foundations.
|
Concordant |
Discordanta |
Unclassifiable | Overall concordance, n (%) | |||
|---|---|---|---|---|---|---|
| HPVA/HPV+ | NHPVA/HPV– | HPVA/HPV– | NHPVA/HPV+ | |||
| Original review | 87 | 16 | 7 | 1 | 0 | 103/111 (92.8) |
| Expert review | 87 | 20 | 1 | 1 | 2 | 107/111 (96.4) |
|
|
||||||
| HPVA/p16+ | NHPVA/p16– | HPVA/p16– | NHPVA/p16+ | |||
|
|
||||||
| Original review | 90 | 16 | 4 | 1 | 0 | 106/111 (95.4) |
| Expert review | 87 | 19 | 1 | 2 | 2 | 106/111 (95.4) |
- 1. Adegoke O, Kulasingam S, Virnig B. Cervical cancer trends in the United States: a 35-year population-based analysis. J Womens Health (Larchmt) 2012; 21: 1031-7. ArticlePubMedPMC
- 2. Smith HO, Tiffany MF, Qualls CR, Key CR. The rising incidence of adenocarcinoma relative to squamous cell carcinoma of the uterine cervix in the United States: a 24-year population-based study. Gynecol Oncol 2000; 78: 97-105. ArticlePubMed
- 3. Stolnicu S, Hoang L, Soslow RA. Recent advances in invasive adenocarcinoma of the cervix. Virchows Arch 2019; 475: 537-49. ArticlePubMedPMCPDF
- 4. McAlpine JN, Leung SC, Cheng A, et al. Human papillomavirus (HPV)-independent vulvar squamous cell carcinoma has a worse prognosis than HPV-associated disease: a retrospective cohort study. Histopathology 2017; 71: 238-46. ArticlePubMedPDF
- 5. Nooij LS, Ter Haar NT, Ruano D, et al. Genomic characterization of vulvar (pre)cancers identifies distinct molecular subtypes with prognostic significance. Clin Cancer Res 2017; 23: 6781-9. ArticlePubMedPDF
- 6. Ang KK, Harris J, Wheeler R, et al. Human papillomavirus and survival of patients with oropharyngeal cancer. N Engl J Med 2010; 363: 24-35. ArticlePubMedPMC
- 7. Fakhry C, Westra WH, Li S, et al. Improved survival of patients with human papillomavirus-positive head and neck squamous cell carcinoma in a prospective clinical trial. J Natl Cancer Inst 2008; 100: 261-9. ArticlePubMedPDF
- 8. Rischin D, Young RJ, Fisher R, et al. Prognostic significance of p16INK4A and human papillomavirus in patients with oropharyngeal cancer treated on TROG 02.02 phase III trial. J Clin Oncol 2010; 28: 4142-8. ArticlePubMedPMC
- 9. Posner MR, Lorch JH, Goloubeva O, et al. Survival and human papillomavirus in oropharynx cancer in TAX 324: a subset analysis from an international phase III trial. Ann Oncol 2011; 22: 1071-7. ArticlePubMedPMCPDF
- 10. Hodgson A, Park KJ, Djordjevic B, et al. International endocervical adenocarcinoma criteria and classification: validation and interobserver reproducibility. Am J Surg Pathol 2019; 43: 75-83. PubMedPMC
- 11. Stolnicu S, Barsan I, Hoang L, et al. International Endocervical Adenocarcinoma Criteria and Classification (IECC): a new pathogenetic classification for invasive adenocarcinomas of the endocervix. Am J Surg Pathol 2018; 42: 214-26. PubMedPMC
- 12. Prigge ES, Arbyn M, von Knebel Doeberitz M, Reuschenbach M. Diagnostic accuracy of p16(INK4a) immunohistochemistry in oropharyngeal squamous cell carcinomas: a systematic review and meta-analysis. Int J Cancer 2017; 140: 1186-98. ArticlePubMedPDF
- 13. Jordan RC, Lingen MW, Perez-Ordonez B, et al. Validation of methods for oropharyngeal cancer HPV status determination in US cooperative group trials. Am J Surg Pathol 2012; 36: 945-54. ArticlePubMedPMC
- 14. Wang F, Flanagan J, Su N, et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J Mol Diagn 2012; 14: 22-9. PubMedPMC
- 15. Schache AG, Liloglou T, Risk JM, et al. Validation of a novel diagnostic standard in HPV-positive oropharyngeal squamous cell carcinoma. Br J Cancer 2013; 108: 1332-9. ArticlePubMedPMCPDF
- 16. Keung ES, Souers RJ, Bridge JA, et al. Comparative performance of high-risk human papillomavirus RNA and DNA in situ hybridization on college of American pathologists proficiency tests. Arch Pathol Lab Med 2020; 144: 344-9. ArticlePubMedPDF
- 17. Satgunaseelan L, Chia N, Suh H, et al. p16 expression in cutaneous squamous cell carcinoma of the head and neck is not associated with integration of high risk HPV DNA or prognosis. Pathology 2017; 49: 494-8. ArticlePubMed
- 18. Augustin J, Outh-Gauer S, Mandavit M, et al. Evaluation of the efficacy of the 4 tests (p16 immunochemistry, polymerase chain reaction, DNA, and RNA in situ hybridization) to evaluate a human papillomavirus infection in head and neck cancers: a cohort of 348 French squamous cell carcinomas. Hum Pathol 2018; 78: 63-71. ArticlePubMed
- 19. Bishop JA, Ma XJ, Wang H, et al. Detection of transcriptionally active high-risk HPV in patients with head and neck squamous cell carcinoma as visualized by a novel E6/E7 mRNA in situ hybridization method. Am J Surg Pathol 2012; 36: 1874-82. ArticlePubMedPMC
- 20. Evans MF, Peng Z, Clark KM, et al. HPV E6/E7 RNA in situ hybridization signal patterns as biomarkers of three-tier cervical intraepithelial neoplasia grade. PLoS One 2014; 9: e91142. ArticlePubMedPMC
- 21. Roldan Urgoiti GB, Gustafson K, Klimowicz AC, Petrillo SK, Magliocco AM, Doll CM. The prognostic value of HPV status and p16 expression in patients with carcinoma of the anal canal. PLoS One 2014; 9: e108790. ArticlePubMedPMC
- 22. Zappacosta R, Colasante A, Viola P, et al. Chromogenic in situ hybridization and p16/Ki67 dual staining on formalin-fixed paraffinembedded cervical specimens: correlation with HPV-DNA test, E6/E7 mRNA test, and potential clinical applications. Biomed Res Int 2013; 2013: 453606.ArticlePubMedPMCPDF
- 23. Aumayr K, Susani M, Horvat R, et al. P16INK4A immunohistochemistry for detection of human papilloma virus-associated penile squamous cell carcinoma is superior to in-situ hybridization. Int J Immunopathol Pharmacol 2013; 26: 611-20. ArticlePubMedPDF
- 24. Winters R, Trotman W, Adamson CS, et al. Screening for human papillomavirus in basaloid squamous carcinoma: utility of p16(INK4a), CISH, and PCR. Int J Surg Pathol 2011; 19: 309-14. ArticlePubMedPDF
- 25. Evans MF, Matthews A, Kandil D, Adamson CS, Trotman WE, Cooper K. Discrimination of ‘driver’ and ‘passenger’ HPV in tonsillar carcinomas by the polymerase chain reaction, chromogenic in situ hybridization, and p16(INK4a) immunohistochemistry. Head Neck Pathol 2011; 5: 344-8. ArticlePubMedPMCPDF
- 26. Sheng Z, Minato H, Sasagawa T, et al. Detection of high-risk human papillomavirus subtypes in cervical glandular neoplasia by in situ hybridization. Int J Clin Exp Pathol 2013; 6: 2168-77. PubMedPMC
- 27. Chen T, Li J, Wang S, Ning Y, Zhou X, Wang Y. High-risk HPV E6/E7 mRNA in situ hybridization in endocervical glandular neoplasia: performance compared with p16(INK4a) and Ki67 immunochemistry. Am J Transl Res 2019; 11: 6498-506. PubMedPMC
- 28. Peng WX, Kure S, Ishino K, et al. P16-positive continuous minimal deviation adenocarcinoma and gastric type adenocarcinoma in a patient with Peutz-Jeghers syndrome. Int J Clin Exp Pathol 2015; 8: 5877-82. PubMedPMC
- 29. Houghton O, Jamison J, Wilson R, Carson J, McCluggage WG. p16 Immunoreactivity in unusual types of cervical adenocarcinoma does not reflect human papillomavirus infection. Histopathology 2010; 57: 342-50. ArticlePubMed
- 30. Lu S, Shen D, Zhao Y, Kang N, Wang X. Primary endocervical gastric-type adenocarcinoma: a clinicopathologic and immunohistochemical analysis of 23 cases. Diagn Pathol 2019; 14: 72.ArticlePubMedPMCPDF
- 31. Stolnicu S, Barsan I, Hoang L, et al. Diagnostic algorithmic proposal based on comprehensive immunohistochemical evaluation of 297 invasive endocervical adenocarcinomas. Am J Surg Pathol 2018; 42: 989-1000. ArticlePubMedPMC
- 32. Singh N, Gilks CB, Wong RW, McCluggage WG, Herrington CS. Interpretation of p16 immunohistochemistry in lower anogenital tract neoplasia [Internet]. Derby: British Association of Gynaecological Pathologists, 2018 [cited 2019 Dec 11]. Available from: https://www.bgcs.org.uk/wp-content/uploads/2019/05/BAGP-UKNEQAScIQC-project-p16-interpretation-guide-2018.pdf .
- 33. Wada T, Ohishi Y, Kaku T, et al. Endocervical adenocarcinoma with morphologic features of both usual and gastric types: clinicopathologic and immunohistochemical analyses and high-risk HPV detection by in situ hybridization. Am J Surg Pathol 2017; 41: 696-705. PubMed
- 34. Burd EM. Human papillomavirus and cervical cancer. Clin Microbiol Rev 2003; 16: 1-17. ArticlePubMedPMCPDF
- 35. Poetsch M, Hemmerich M, Kakies C, et al. Alterations in the tumor suppressor gene p16(INK4A) are associated with aggressive behavior of penile carcinomas. Virchows Arch 2011; 458: 221-9. ArticlePubMedPDF
- 36. Nuovo GJ, Plaia TW, Belinsky SA, Baylin SB, Herman JG. In situ detection of the hypermethylation-induced inactivation of the p16 gene as an early event in oncogenesis. Proc Natl Acad Sci U S A 1999; 96: 12754-9. ArticlePubMedPMC
REFERENCES
Figure & Data
References
Citations

- Role of human papillomavirus status in the classification, diagnosis, and prognosis of malignant cervical epithelial tumors and precursor lesions
Simona Stolnicu
Die Pathologie.2026; 47(S1): 97. CrossRef - EdgeNeXt-SEDP for cervical adenocarcinoma HPV-associated and non-HPV-associated diagnosis and decision support
Qi Chen, Hao Wang, Hao Zhang, Zhenkun Zhu, Xi Wei
Life Sciences.2025; 380: 123931. CrossRef - Cytology and histology of endocervical glandular lesions: a review with emphasis on recent developments
Natalie Banet, Karen L. Talia
Pathology.2025; 57(7): 817. CrossRef - Joint detection of multiple HPV-testing technologies and evaluation of clinicopathological characteristics discriminate between HPV-independent and low-copy HPV-associated cervical squamous cell carcinoma (CSCC) -an analysis of 3869 cases
Linghui Lu, Tianqi Liu, Shunni Wang, Jing Li, Feiran Zhang, Yan Ning, Yiqin Wang
Gynecologic Oncology.2023; 170: 59. CrossRef - Incidence and Clinicopathologic Characteristics of Human Papillomavirus–independent Invasive Squamous Cell Carcinomas of the Cervix
Simona Stolnicu, Douglas Allison, Aaron M. Praiss, Basile Tessier-Cloutier, Amir Momeni Boroujeni, Jessica Flynn, Alexia Iasonos, Rene Serrette, Lien Hoang, Andrei Patrichi, Cristina Terinte, Anna Pesci, Claudia Mateoiu, Ricardo R. Lastra, Takako Kiyokawa
American Journal of Surgical Pathology.2023; 47(12): 1376. CrossRef - Testing Algorithms for the Diagnosis of Malignant Glandular Tumors of the Uterine Cervix Histotyped per the International Endocervical Adenocarcinoma Criteria and Classification (IECC) System
Máire A. Duggan, Qiuli Duan, Ruth M. Pfeiffer, Mary Anne Brett, Sandra Lee, Mustapha Abubakar, Martin Köbel, Monica Rodriguez, Aylin Sar
Applied Immunohistochemistry & Molecular Morphology.2022; 30(2): 91. CrossRef - Local and Metastatic Relapses in a Young Woman with Papillary Squamous Cell Carcinoma of the Uterine Cervix
Ha Young Woo, Hyun-Soo Kim
Diagnostics.2022; 12(3): 599. CrossRef - Clinical correlation of lymphovascular invasion and Silva pattern of invasion in early-stage endocervical adenocarcinoma: proposed binary Silva classification system
Simona Stolnicu, Lien Hoang, Noorah Almadani, Louise De Brot, Glauco Baiocchi, Graziele Bovolim, Maria Jose Brito, Georgia Karpathiou, Antonio Ieni, Esther Guerra, Takako Kiyokawa, Pavel Dundr, Carlos Parra-Herran, Sofia Lérias, Ana Felix, Andres Roma, An
Pathology.2022; 54(5): 548. CrossRef - Reproducibility of Morphologic Parameters of the International Endocervical Adenocarcinoma Criteria and Classification System and Correlation With Clinicopathologic Parameters: A Multi-Institutional Study
Pinar Bulutay, Nihan Haberal, Özlem Özen, Özlem Erdem, Emine H. Zeren, İbrahim Kulac, Çagatay Taskiran, Dogan Vatansever, Ali Ayhan, Nilgün Kapucuoğlu
International Journal of Gynecological Pathology.2022; 41(5): 447. CrossRef - HPV-Negative Cervical Cancer: A Narrative Review
Francesca Arezzo, Gennaro Cormio, Vera Loizzi, Gerardo Cazzato, Viviana Cataldo, Claudio Lombardi, Giuseppe Ingravallo, Leonardo Resta, Ettore Cicinelli
Diagnostics.2021; 11(6): 952. CrossRef - International Endocervical Adenocarcinoma Criteria and Classification (IECC): An Independent Cohort With Clinical and Molecular Findings
Hezhen Ren, Noorah Almadani, Jennifer Pors, Samuel Leung, Julie Ho, Christine Chow, Monica Ta, Kay J. Park, Simona Stolnicu, Robert Soslow, David Huntsman, Blake C. Gilks, Lynn Hoang
International Journal of Gynecological Pathology.2021; 40(6): 533. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
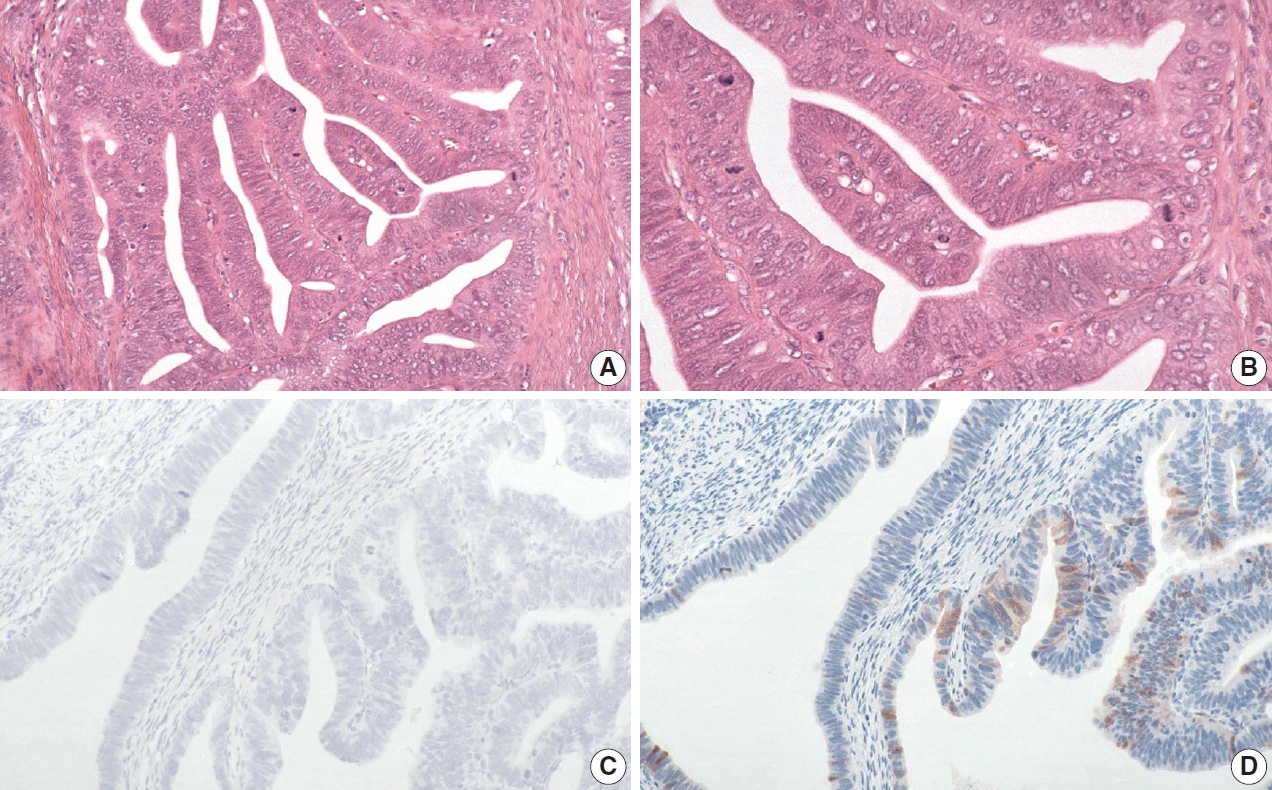
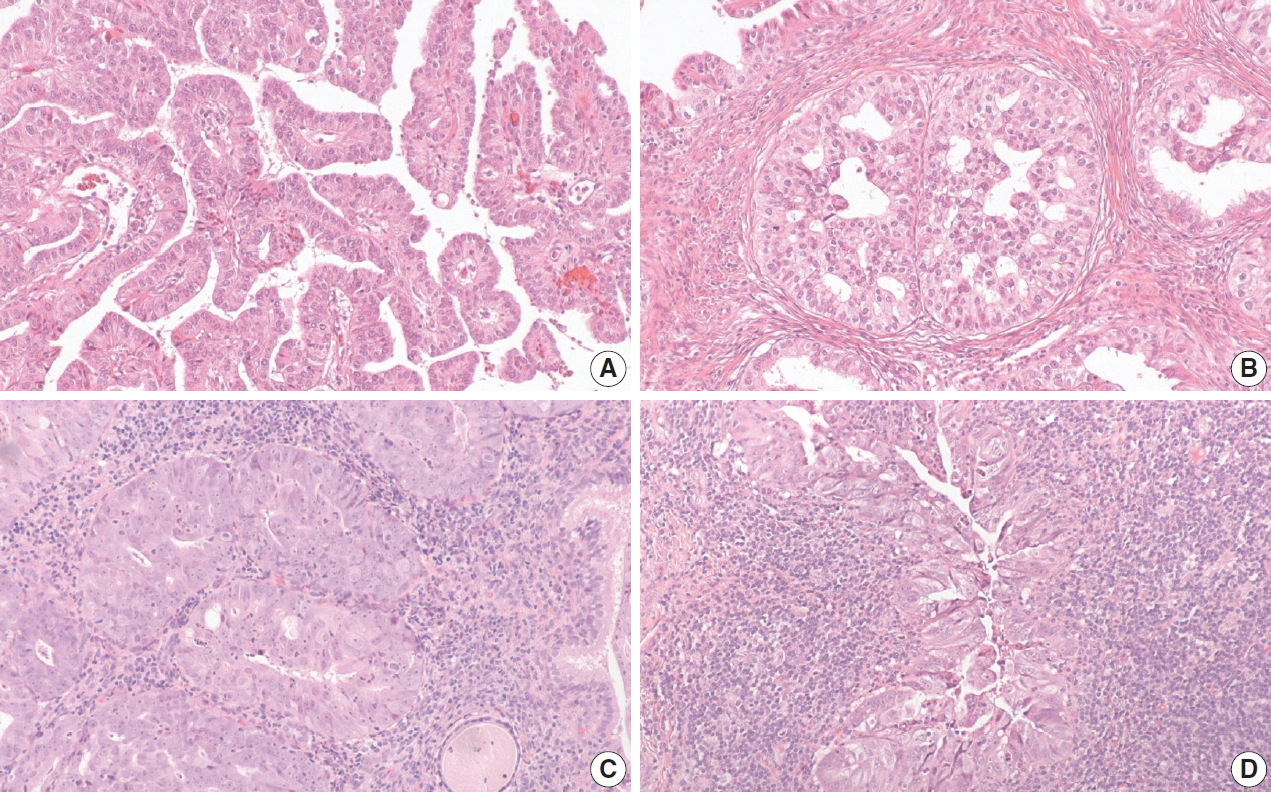
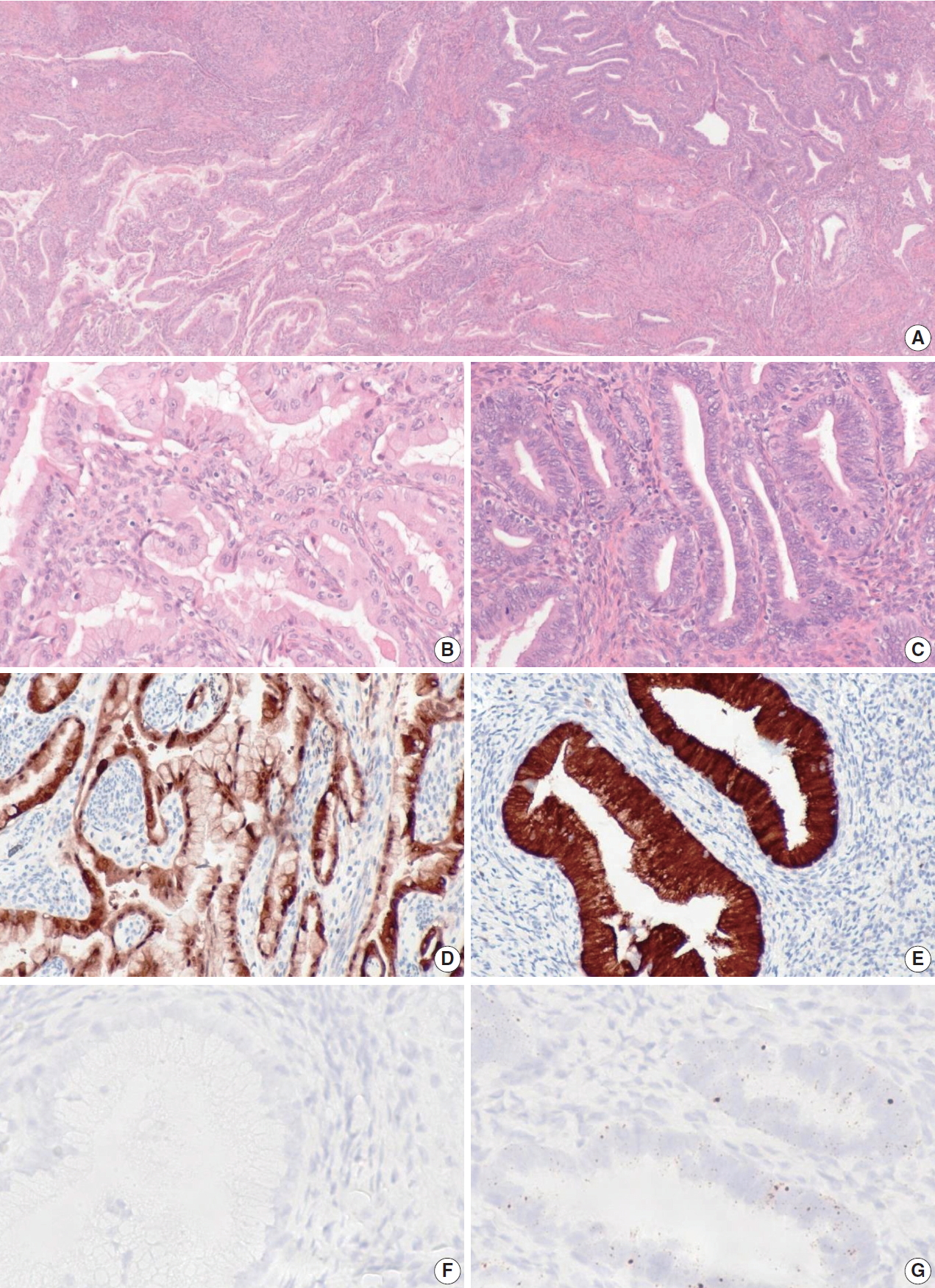
Fig. 1.
Fig. 2.
Fig. 3.
| Concordant |
Discordant |
Unclassifiable | Overall concordance, n (%) | |||
|---|---|---|---|---|---|---|
| HPVA/HPV+ | NHPVA/HPV– | HPVA/HPV– | NHPVA/HPV+ | |||
| Original review | 87 | 16 | 7 | 1 | 0 | 103/111 (92.8) |
| Expert review | 87 | 20 | 1 | 1 | 2 | 107/111 (96.4) |
| HPVA/p16+ | NHPVA/p16– | HPVA/p16– | NHPVA/p16+ | |||
| Original review | 90 | 16 | 4 | 1 | 0 | 106/111 (95.4) |
| Expert review | 87 | 19 | 1 | 2 | 2 | 106/111 (95.4) |
| Original histotype | p16 | HPV RISH | Histology review | Reclassified |
|---|---|---|---|---|
| HPVA-usual | Neg | Neg | Internal review: Demonstrated HPVA features, with morphology in keeping with usual-type | HPVA-usual |
| External reviewer 2: HPVA-usual | ||||
| External reviewer 3: HPVA-usual | ||||
| HPVA-mucinous (intestinal) | Neg | Neg | Internal review: Goblet cells present; Lack of significant HPVA features, with morphology in keeping with gastric-type | NHPVA-gastric |
| External reviewer 2: NHPVA-gastric | ||||
| External reviewer 3: NHPVA-gastric | ||||
| HPVA-mucinous NOS | Neg | Neg | Internal review: Lack of significant HPVA features, with morphology in keeping with gastric-type | NHPVA-gastric |
| External reviewer 2: NHPVA-gastric or endometrioid | ||||
| External reviewer 3: NHPVA-gastric or endometrioid | ||||
| HPVA-mucinous NOS | Neg | Neg | Internal review: Lack of significant HPVA features, with morphology in keeping with gastric-type | NHPVA-gastric |
| External reviewer 2: NHPVA-gastric | ||||
| External reviewer 3: NHPVA-gastric | ||||
| HPVA-mucinous NOS | Pos | Neg | Internal review: Lack of significant HPVA features; Mixture of cuboidal and columnar cells with vague intracytoplasmic mucin; Likely NHPVA-gastric | NHPVA-NOS |
| External reviewer 2: NHPVA-gastric or endometrioid | ||||
| External reviewer 3: NHPVA-clear cell | ||||
| HPVA-mucinous NOS | Pos | Neg | Internal review: Usual or adenocarcinoma NOS unclassified, need HPV status | Unclassifiable |
| External reviewer 2: HPVA-usual or NOS | ||||
| External reviewer 3: Unclassifiable | ||||
| HPVA-mucinous NOS | Pos | Neg | Internal review: Nested cells with amphophilic cytoplasm and dense lymphoplasmacytic infiltrate–Unclassifiable | Unclassifiable |
| External reviewer 2: HPVA-usual | ||||
| External reviewer 3: Unclassifiable | ||||
| NHPVA-gastric | Pos | Mixed | Internal review: HPVA features focally present, hybrid morphology in keeping with mixed usual and gastric-type | Mixed (usual and gastric) |
| Not available for external review |
HPV, human papillomavirus; RISH, RNA in-situ hybridization; IHC, immunohistochemistry; IECC, International Endocervical Adenocarcinoma Criteria and Classification; HPVA, HPV associated. Discordant cases are detailed in Table 2.
IHC, immunohistochemistry; HPV, human papillomavirus; RISH, RNA in-situ hybridization; HPVA, HPV associated; Neg, negative; NHPVA, non–HPV-associated; Pos, positive; NOS, not otherwise indicated.

 E-submission
E-submission





