Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 53(6); 2019 > Article
-
Original Article
MicroRNA-374a Expression as a Prognostic Biomarker in Lung Adenocarcinoma -
Yeseul Kim
 , Jongmin Sim1
, Jongmin Sim1 , Hyunsung Kim
, Hyunsung Kim , Seong Sik Bang
, Seong Sik Bang , Seungyun Jee
, Seungyun Jee , Sungeon Park
, Sungeon Park , Kiseok Jang
, Kiseok Jang
-
Journal of Pathology and Translational Medicine 2019;53(6):354-360.
DOI: https://doi.org/10.4132/jptm.2019.10.01
Published online: October 24, 2019
Department of Pathology, Hanyang University College of Medicine, Seoul, Korea
1Department of Pathology, Samsung Medical Center, Seoul, Korea
- Corresponding Author: Kiseok Jang, MD, PhD, Department of Pathology, Hanyang University College of Medicine, 222-1 Wangsimni-ro, Seongdong-gu, Seoul 04763, Korea Tel: +82-2-2290-8248, Fax: +82-2-2296-7502, E-mail: medartisan@hanyang.ac.kr
© 2019 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- Lung cancer is the most common cause of cancer-related death, and adenocarcinoma is the most common histologic subtype. MicroRNA is a small non-coding RNA that inhibits multiple target gene expression at the post-transcriptional level and is commonly dysregulated in malignant tumors. The purpose of this study was to analyze the expression of microRNA-374a (miR-374a) in lung adenocarcinoma and correlate its expression with various clinicopathological characteristics.
-
Methods
- The expression level of miR-374a was measured in 111 formalin-fixed paraffin-embedded lung adenocarcinoma tissues using reverse transcription-quantitative polymerase chain reaction assays. The correlation between miR-374a expression and clinicopathological parameters, including clinical outcome, was further analyzed.
-
Results
- High miR-374 expression was correlated with advanced pT category (chi-square test, p=.004) and pleural invasion (chi-square test, p=.034). Survival analysis revealed that patients with high miR-374a expression had significantly shorter disease-free survival relative to those with low miR-374a expression (log-rank test, p=.032).
-
Conclusions
- miR-374a expression may serve as a potential prognostic biomarker for predicting recurrence in early stage lung adenocarcinoma after curative surgery.
- Patients and tumor samples
- A total of 184 consecutive cases of curative surgery for primary lung adenocarcinoma conducted at Hanyang University Hospital, Seoul, Korea, from 2003 to 2014 were retrospectively selected. Of the total 184 cases, 37 (20.1%) were excluded due to the lack of a suitable FFPE sample and 36 of the remaining patients were excluded because of the poor quality of FFPE RNAs. Finally, qRT-PCR for miR-374a and the U6 control was successful in 111 cases and further statistical analyses were performed. None of the patients received preoperative therapy. Clinicopathological data were collected and reviewed from the medical records and histopathological reports, and additional review of archived pathologic slides was conducted. The clinicopathological parameters included patient age, sex, tumor size, T category, pleural invasion, lymph node metastasis, American Joint Committee on Cancer (AJCC) tumor stage, histological grade, lymphovascular invasion, and perineural invasion. Histologic grading was determined by conventional histological criteria, including architectural abnormalities and cytologic atypia. Most cases with histologic grade 1 were categorized as lepidic predominant subtype, cases with grade 2 were categorized as acinar or papillary predominant subtype, and cases with grade 3 were categorized as solid or micropapillary predominant subtype based on the new 2015 World Health Organization classification. Disease-free survival (DFS) was measured from the date of operation until recurrence or death. Overall survival (OS) was measured from the date of operation until the time of death or the last follow-up.
- RNA extraction and qRT-PCR
- To identify areas most representative of tumors and non-necrotic tumor sections, we reviewed and marked histologic slides. Three or four 10-µm-thick tissue sections from each block were collected. Total RNA was isolated from the FFPE tumor tissues using miRNeasy FFPE kits (Qiagen, Hilden, Germany) according to the manufacturer’s instruction. The concentration and purity of extracted RNA was measured using a NanoDrop 2000 spectrophotometer (NanoDrop Technologies, Waltham, MA, USA).
- Universal cDNA synthesis kits (Exiqon, Vedbaek, Denmark) were used to convert RNA into cDNA. qRT-PCR experiments were performed with a mixture of diluted cDNA samples, an miR-374a specific primer set (Exiqon), and an ExiLENT SYBR Green Master mix (Exiqon). PCR was performed using a CFX96 thermocycler (Bio-Rad, Hercules, CA, USA) under the following conditions: 95°C for 15 minutes followed by 45 cycles of 95°C for 10 seconds and 60°C for 1 minute. After PCR amplification, melting curve analysis was performed to confirm the specificity of PCR products. The expression level of miR-374a was calculated using the 2-ΔCt method relative to the U6 small nuclear RNA (RNU6B).
- Statistical analysis
- Statistical analysis was performed using SPSS ver. 21.0 software (IBM Corp., Armonk, NY, USA). Chi-square tests were used to evaluate associations between miR-374a expression and various clinicopathological parameters of pulmonary adenocarcinoma patients. Kaplan-Meier survival curves for both DFS and OS were plotted, and the log-rank test was applied to establish the level of significance. The Cox proportional hazard regression model was employed in both univariable and multivariable survival analyses. A p-value of <.05 was defined as statistically significant.
- Ethics statement
- This study was approved by the Institutional Review Board (IRB) of Hanyang University Hospital (IRB file no. 2016-07-038) with a waiver of informed consent.
MATERIALS AND METHODS
- Clinicopathological characteristics of patients enrolled in the study
- Two-thirds of patients were over 60 years of age (n=68, 61.3%) and more females than men (n=65, 58.6%) were included in the study. Most of the cases were categorized as histologic grade 2 (n=78, 69.6%) and AJCC stage I (n=76, 68.5%). Lymph node metastasis was found in 25 cases (22.5%), pleural invasion in 46 cases (41.4%), lymphovascular invasion in 40 cases (36.0%), and perineural invasion in 21 cases (18.9%). The clinicopathological characteristics are summarized in Table 1.
- Correlations between miR-374a expression and clinicopathological characteristics in pulmonary adenocarcinoma
- The quantitative measures of relative miR-374a expression in all 111 lung adenocarcinoma samples were as follows: mean, 0.048; median, 0.066; standard deviation, 1.585; and range, 0.000–11.794. The patients were divided into two groups (low vs high) according to the median value of miR-374a expression. We analyzed the associations between miR-374a expression and clinicopathological characteristics, including age, sex, AJCC stage group, primary tumor (T category), lymph node metastasis, pleural invasion, histologic grade, lymphovascular invasion, and perineural invasion. High miR-374a expression was correlated with advanced T category (chi-square test, p=.004) and the presence of pleural invasion (chi-square test, p=.034) (Table 2).
- Prognostic value of miR-374a expression in pulmonary adenocarcinoma
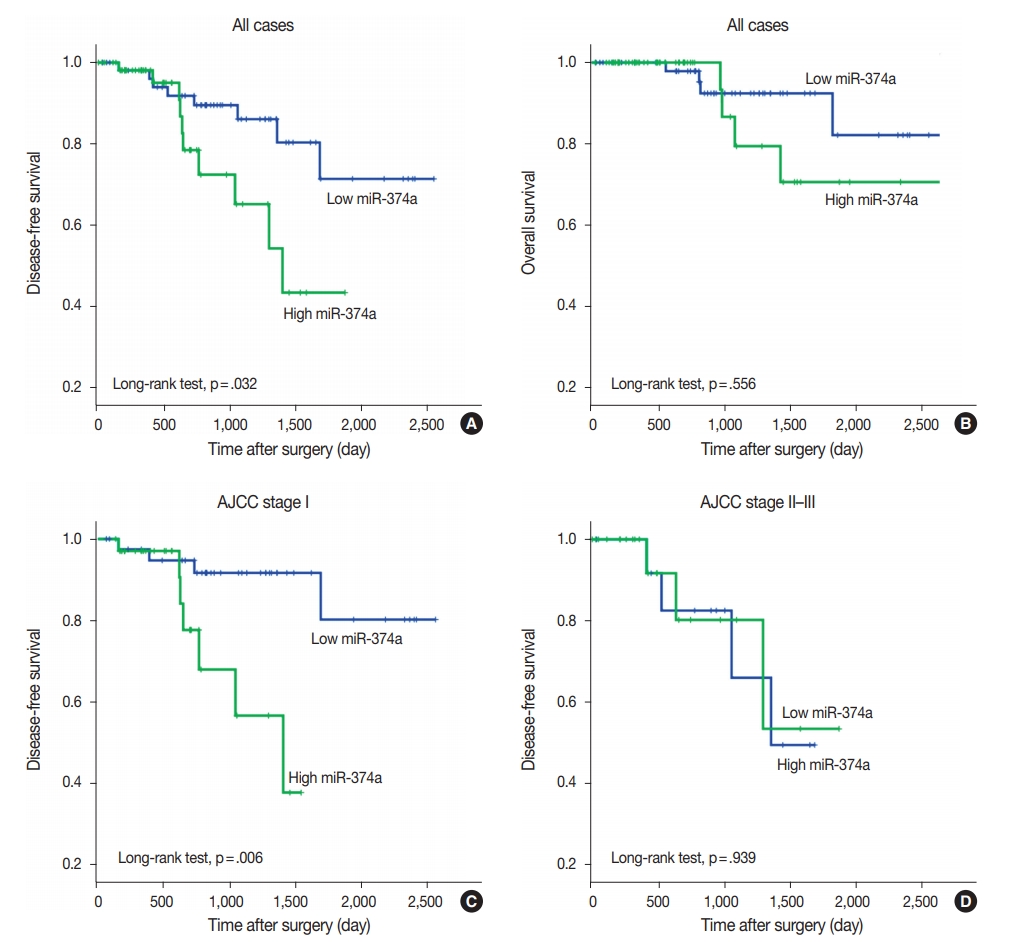
- The median follow-up interval of patients for OS and DFS was 27.3 months (range, 0.9 to 118.6 months) and 25.7 months (range, 0.9 to 85.0 months), respectively. Local recurrence or metastasis was found in 18 patients, and 9 patients died during the follow-up period. On univariate survival analysis, AJCC stage, T category, lymph node metastasis, histologic grade, lymphovascular invasion, and perineural invasion were revealed as prognostic factors for DFS and/or OS (Table 3). Patients with high miR-374a expression showed shorter DFS compared to those with low miR-374a expression, and the difference was statistically significant (log-rank test, p=.032) (Fig. 1). However, miR-374a expression level was not an independent prognostic factor in multivariable survival analysis. The OS of the patients with high miR-374a expression was also inferior to those with low expression; however, the difference failed to demonstrate a statistical significance. In addition, stage-stratified survival analyses revealed that high miR-374a expression was significantly associated with shorter DFS only in patients with AJCC stage I (log-rank test, p=.006).
RESULTS
- Fresh frozen tissue has been mostly used for high-throughput miRNA expression profiling for human cancers. However, sampling and long-term storage of fresh tumor tissue is laborious and expensive in daily clinical practice. As tumor specimens are routinely stored as FFPE blocks after diagnostic process in pathology archives and can be linked with medical records, emerging interest has been directed toward whether FFPE tissue can be used for studies of miRNA biomarkers. Interestingly, miRNAs appear to be stable in FFPE samples, even exhibiting near total mRNA degradation [17]. Preservation of miRNA in FFPE tumor samples has been confirmed in previous studies, and the results obtained from qRT-PCR analyses, miRNA expression profiling, and even next-generation sequencing studies are consistent with those of fresh frozen tissue [18-21]. These results suggest that quantifying miRNA expression level using FFPE tissue can enable the identification of clinically-available miRNA biomarkers in human malignancies.
- One of the clinical significances of measuring miRNA expression level in resected tumor tissues is to identify patients at high risk of recurrence or cancer-related death after curative surgery. Zheng et al. [22] reported that NSCLC patients with low miR-195 expression showed poor overall survival. In the study of Chen et al. [23], low miRNA-148a expression was significantly correlated with high histologic grade, frequent lymph node metastasis, and a poor OS in NSCLC. Further, miR-21, as an oncogenic miRNA, has been reported as a biomarker for predicting recurrence, metastasis, and resistance of radiation and chemotherapy in NSCLC [24-26]. In our study, high miR-374a expression correlated with advanced pT category, pleural invasion, and poor DFS in lung adenocarcinoma. Zhao et al. [16] investigated miR-374a expression level in 158 NSCLC tissues by in situ hybridization. They found that miR-374a was highly expressed in NSCLC relative to non-neoplastic tissues and correlated with lymph node metastasis and poor clinical outcome [16]. However, in the study of 38 Estonian NSCLC cases (18 squamous cell carcinomas and 20 adenocarcinomas), low miR-374a expression, as by miRNA microarray, was associated with poor survival. This discrepancy may result from the difference in miRNA detection methods and study population, such as ethnicity and histologic type. Lack of reproducibility between studies suggests that further validation with a larger case series and optimization of detection methods is needed for clarification.
- Most previous studies have reported miR-374a as an oncogenic miRNA in various cancer types. In studies with osteosarcoma cell lines, tumor cells transiently transfected with miR-374a mimic showed an increase in cell proliferation and colony formation by downr-egulating FOXO1 and/or Axin2 [7,27]. Downregulation of miR-374a inhibited cell proliferation, migration, and invasion in breast cancer cell lines (MDA-MB-23a and MCF-7) by targeting LACTB [11]. In triple-negative breast cancer, ARRB1 was a direct target of miR-374a, and knockdown of miR-374a attenuated cell proliferation and migration in vitro [13]. Downregulation of miR-374a inhibited tumor growth and decreased Ki-67 proliferation index in xenograft mice [13]. However, some studies have reported miR-374a as a tumor suppressive miRNA. Induced overexpression of miR-374a in colon cancer cell lines inhibited cell proliferation and invasion based on in vitro assays and a xenograft model by targeting cyclin D1 [14]. Nasopharyngeal cancer cells transfected with miR-374a mimic inhibited cell proliferation and invasion in vitro, and showed a decreased number of metastatic lesions in a mouse model [28]. These results suggest that miR-374a may have dual roles in tumor progression, and the role depends on the major target pathway in a given cancer subtype.
- In conclusion, our data show that high miR-374a expression correlates with advanced pT category and pleural invasion. High expression of miR-374a may serve as a potential prognostic biomarker for predicting recurrence in early stage lung adenocarcinoma after curative surgery. Further validation studies for a prognostic biomarker and related mechanism should be explored.
DISCUSSION
Author contributions
Conceptualization: KJ.
Data curation: YK, JS, HK, SSB, SJ, SP.
Formal analysis: JS, YK, KJ.
Funding acquisition: KJ.
Investigation: JS, YK, KJ.
Methodology: KJ.
Project administration: KJ.
Resources: JS, KJ.
Supervision: KJ.
Validation: JS, YK, KJ.
Visualization: YK, KJ.
Writing—original draft preparation: YK, KJ.
Writing—review & editing: YK, KJ.
Conflicts of Interest
The authors declare that they have no potential conflicts of interest.
Funding
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Korean government (Ministry of Science, ICT & Future Planning) (NRF-2015R1C1 A1A01056091) and Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2018R1D1A1B07048798).
Acknowledgments
- 1. Cronin KA, Lake AJ, Scott S, et al. Annual report to the nation on the status of cancer, part I: national cancer statistics. Cancer 2018; 124: 2785-800. ArticlePubMedPMCPDF
- 2. Kim HC, Jung CY, Cho DG, et al. Clinical characteristics and prognostic factors of lung cancer in Korea: a pilot study of data from the Korean Nationwide Lung Cancer Registry. Tuberc Respir Dis 2019; 82: 118-25. ArticlePDF
- 3. Chen K, Rajewsky N. The evolution of gene regulation by transcription factors and microRNAs. Nat Rev Genet 2007; 8: 93-103. ArticlePubMedPDF
- 4. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281-97. ArticlePubMed
- 5. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857-66. ArticlePubMedPDF
- 6. Trang P, Weidhaas JB, Slack FJ. MicroRNAs as potential cancer therapeutics. Oncogene 2008; 27 Suppl 2: S52-7. ArticlePubMedPDF
- 7. Lu T, Zhang C, Chai MX, An YB, Jia JL. MiR-374a promotes the proliferation of osteosarcoma cell proliferation by targeting Axin2. Int J Clin Exp Pathol 2015; 8: 10776-83. PubMedPMC
- 8. Wang Y, Xin H, Han Z, Sun H, Gao N, Yu H. MicroRNA-374a promotes esophageal cancer cell proliferation via Axin2 suppression. Oncol Rep 2015; 34: 1988-94. ArticlePubMed
- 9. Cai J, Guan H, Fang L, et al. MicroRNA-374a activates Wnt/ β-catenin signaling to promote breast cancer metastasis. J Clin Invest 2013; 123: 566-79. ArticlePubMedPMC
- 10. Li H, Chen H, Wang H, et al. MicroRNA-374a promotes hepatocellular carcinoma cell proliferation by targeting mitogen-inducible gene 6 (MIG-6). Oncol Res 2018; 26: 557-63. ArticlePubMedPMC
- 11. Zhang J, He Y, Yu Y, et al. Upregulation of miR-374a promotes tumor metastasis and progression by downregulating LACTB and predicts unfavorable prognosis in breast cancer. Cancer Med 2018 May 23 https://doi.org/10.1002/cam4.1576. Article
- 12. Ma L, Shao Z, Zhao Y. MicroRNA-374a promotes pancreatic cancer cell proliferation and epithelial to mesenchymal transition by targeting SRCIN1. Pathol Res Pract 2019; 215: 152382.ArticlePubMed
- 13. Son D, Kim Y, Lim S, et al. MiR-374a-5p promotes tumor progression by targeting ARRB1 in triple negative breast cancer. Cancer Lett 2019; 454: 224-33. ArticlePubMed
- 14. Chen Y, Jiang J, Zhao M, et al. MicroRNA-374a suppresses colon cancer progression by directly reducing CCND1 to inactivate the PI3K/AKT pathway. Oncotarget 2016; 7: 41306-19. ArticlePubMedPMC
- 15. Qian D, Chen K, Deng H, et al. MicroRNA-374b suppresses proliferation and promotes apoptosis in T-cell lymphoblastic lymphoma by repressing AKT1 and Wnt-16. Clin Cancer Res 2015; 21: 4881-91. ArticlePubMedPDF
- 16. Zhao M, Xu P, Liu Z, et al. Dual roles of miR-374a by modulated cJun respectively targets CCND1-inducing PI3K/AKT signal and PTEN-suppressing Wnt/β-catenin signaling in non-small-cell lung cancer. Cell Death Dis 2018; 9: 78.PubMedPMC
- 17. Hall JS, Taylor J, Valentine HR, et al. Enhanced stability of microRNA expression facilitates classification of FFPE tumour samples exhibiting near total mRNA degradation. Br J Cancer 2012; 107: 684-94. ArticlePubMedPMCPDF
- 18. Zhang X, Chen J, Radcliffe T, Lebrun DP, Tron VA, Feilotter H. An array-based analysis of microRNA expression comparing matched frozen and formalin-fixed paraffin-embedded human tissue samples. J Mol Diagn 2008; 10: 513-9. ArticlePubMedPMC
- 19. Weng L, Wu X, Gao H, et al. MicroRNA profiling of clear cell renal cell carcinoma by whole-genome small RNA deep sequencing of paired frozen and formalin-fixed, paraffin-embedded tissue specimens. J Pathol 2010; 222: 41-51. ArticlePubMed
- 20. Glud M, Klausen M, Gniadecki R, et al. MicroRNA expression in melanocytic nevi: the usefulness of formalin-fixed, paraffin-embedded material for miRNA microarray profiling. J Invest Dermatol 2009; 129: 1219-24. ArticlePubMed
- 21. Hui AB, Shi W, Boutros PC, et al. Robust global micro-RNA profiling with formalin-fixed paraffin-embedded breast cancer tissues. Lab Invest 2009; 89: 597-606. ArticlePubMedPDF
- 22. Zheng J, Xu T, Chen F, Zhang Y. MiRNA-195-5p functions as a tumor suppressor and a predictive of poor prognosis in non-small cell lung cancer by directly targeting CIAPIN1. Pathol Oncol Res 2019; 25: 1181-90. ArticlePubMedPMCPDF
- 23. Chen Y, Min L, Ren C, et al. MiRNA-148a serves as a prognostic factor and suppresses migration and invasion through Wnt1 in non-small cell lung cancer. PLoS One 2017; 12: e0171751. ArticlePubMedPMC
- 24. Zheng W, Zhao J, Tao Y, et al. MicroRNA-21: a promising biomarker for the prognosis and diagnosis of non-small cell lung cancer. Oncol Lett 2018; 16: 2777-82. PubMedPMC
- 25. Wang XC, Wang W, Zhang ZB, Zhao J, Tan XG, Luo JC. Overexpression of miRNA-21 promotes radiation-resistance of non-small cell lung cancer. Radiat Oncol 2013; 8: 146.PubMedPMC
- 26. Gao W, Lu X, Liu L, Xu J, Feng D, Shu Y. MiRNA-21: a biomarker predictive for platinum-based adjuvant chemotherapy response in patients with non-small cell lung cancer. Cancer Biol Ther 2012; 13: 330-40. PubMed
- 27. He W, Feng L, Xia D, Han N. MiR-374a promotes the proliferation of human osteosarcoma by downregulating FOXO1 expression. Int J Clin Exp Med 2015; 8: 3482-9. PubMedPMC
- 28. Zhen Y, Fang W, Zhao M, et al. MiR-374a-CCND1-pPI3K/AKT-c-JUN feedback loop modulated by PDCD4 suppresses cell growth, metastasis, and sensitizes nasopharyngeal carcinoma to cisplatin. Oncogene 2017; 36: 275-85. ArticlePubMedPDF
REFERENCES
Figure & Data
References
Citations

- Upregulated miR-374a-5p drives psoriasis pathogenesis through WIF1 downregulation and Wnt5a/NF-κB activation
Jing Ma, Lu Gan, Hongying Chen, Lihao Chen, Yu Hu, Chao Luan, Kun Chen, Jiaan Zhang
Cellular Signalling.2024; 119: 111171. CrossRef - Association between the expression level of miRNA‑374a and TGF‑β1 in patients with colorectal cancer
Noha El Din, Reem El‑Shenawy, Rehab Moustafa, Ahmed Khairy, Sally Farouk
World Academy of Sciences Journal.2024;[Epub] CrossRef - Cell-free plasma miRNAs analysis for low invasive lung cancer diagnostics
M. Yu. Konoshenko, P. P. Laktionov, Yu. A. Lancuhaj, S. V. Pak, S. E. Krasilnikov, O. E. Bryzgunova
Advances in Molecular Oncology.2023; 10(2): 78. CrossRef - MicroRNA‑mediated regulation in lung adenocarcinoma: Signaling pathways and potential therapeutic implications (Review)
Jiye Liu, Fei Zhang, Jiahe Wang, Yibing Wang
Oncology Reports.2023;[Epub] CrossRef - Dysregulation of miR-374a is involved in the progression of diabetic retinopathy and regulates the proliferation and migration of retinal microvascular endothelial cells
Zhanhong Wang, Xiao Zhang, Yanjun Wang, Dailing Xiao
Clinical and Experimental Optometry.2022; 105(3): 287. CrossRef - MicroRNA Profile for Diagnostic and Prognostic Biomarkers in Thyroid Cancer
Jong-Lyul Park, Seon-Kyu Kim, Sora Jeon, Chan-Kwon Jung, Yong-Sung Kim
Cancers.2021; 13(4): 632. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
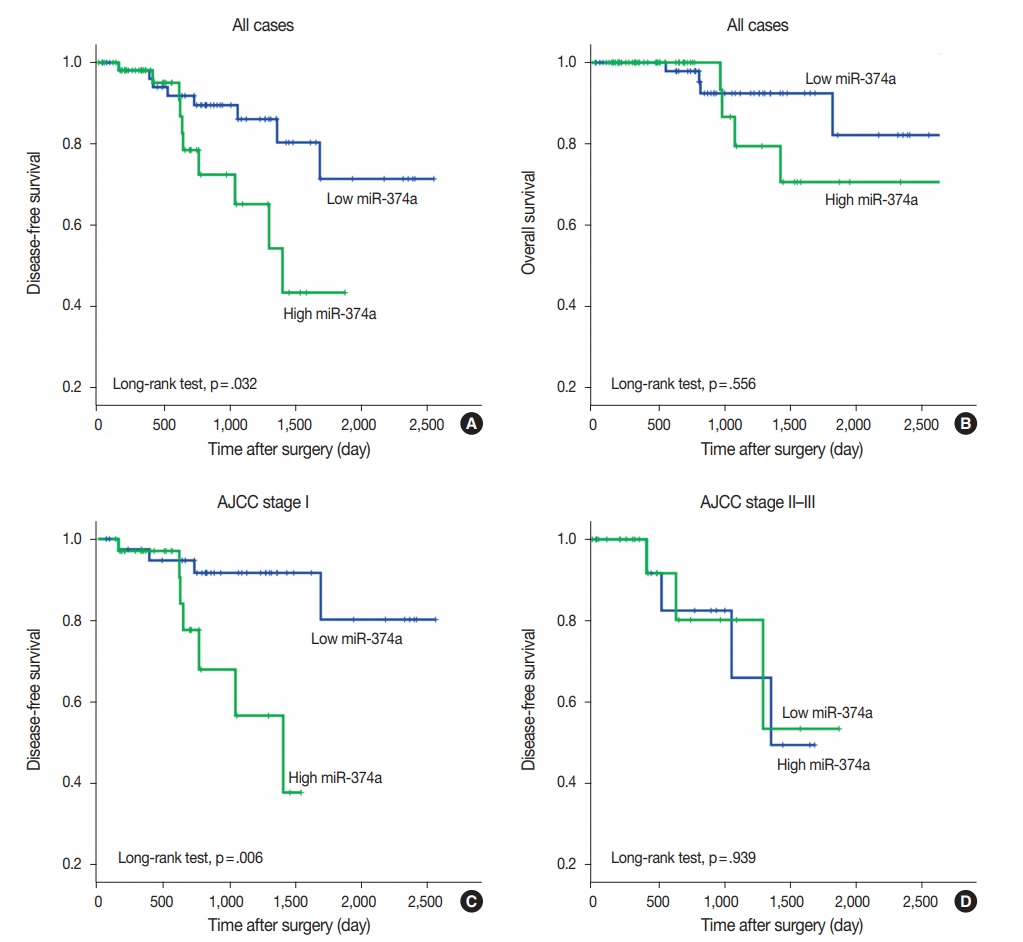
Fig. 1.
| Characteristic | No. (%) (n = 111) |
|---|---|
| Age (yr) | |
| < 60 | 43 (38.7) |
| ≥ 60 | 68 (61.3) |
| Sex | |
| Male | 46 (41.4) |
| Female | 65 (58.6) |
| Histological grade | |
| Grade 1 | 23 (20.7) |
| Grade 2 | 78 (70.3) |
| Grade 3 | 10 (9.0) |
| T category | |
| pT1 | 51 (45.9) |
| pT2 | 53 (47.7) |
| pT3 | 5 (4.5) |
| pT4 | 2 (1.8) |
| AJCC stage group | |
| I | 76 (68.5) |
| II | 17 (15.3) |
| III | 18 (16.2) |
| Lymph node metastasis | |
| pN0 | 86 (77.5) |
| pN1 | 10 (9.0) |
| pN2 | 13 (11.7) |
| pN3 | 2 (1.8) |
| Pleural invasion | |
| PL0 | 65 (58.6) |
| PL1 | 35 (31.5) |
| PL2 | 10 (9.0) |
| PL3 | 1 (0.9) |
| Lymphvascular invasion | |
| Absent | 71 (64.0) |
| Present | 40 (36.0) |
| Perineural invasion | |
| Absent | 90 (81.1) |
| Present | 21 (18.9) |
| Clinicopathological characteristic | No. | miR-374a expression |
Chi-square test |
|
|---|---|---|---|---|
| Low | High | p-value | ||
| Age (yr) | .437 | |||
| < 60 | 43 | 19 (44.2) | 24 (55.8) | |
| ≥ 60 | 68 | 36 (52.9) | 32 (47.1) | |
| Sex | .337 | |||
| Male | 46 | 20 (43.5) | 26 (56.5) | |
| Female | 65 | 35 (53.8) | 30 (46.2) | |
| AJCC stage group | .221 | |||
| I | 76 | 41 (53.9) | 35 (46.1) | |
| II, III | 35 | 14 (40.0) | 21 (60.0) | |
| Primary tumor | .004 | |||
| pT1 | 51 | 33 (64.7) | 18 (35.3) | |
| pT2, pT3, pT4 | 60 | 22 (36.7) | 38 (63.3) | |
| LN metastasis | > .99 | |||
| Negative | 86 | 43 (50.0) | 43 (50.0) | |
| Positive | 25 | 12 (48.0) | 13 (52.0) | |
| Pleural invasion | .034 | |||
| Negative | 65 | 38 (58.5) | 27 (41.5) | |
| Positive | 46 | 17 (37.0) | 29 (63.0) | |
| Histological grade | .490 | |||
| Grade 1 | 23 | 13 (56.5) | 10 (43.5) | |
| Grade 2 and 3 | 88 | 42 (47.7) | 46 (52.3) | |
| Lymphovascular invasion | .695 | |||
| Absent | 71 | 34 (47.9) | 37 (52.1) | |
| Present | 40 | 21 (52.5) | 19 (47.5) | |
| Perineural invasion | .234 | |||
| Absent | 90 | 42 (46.7) | 48 (53.3) | |
| Present | 21 | 13 (61.9) | 8 (38.1) | |
| Disease-free survival |
Overall survival |
|||||
|---|---|---|---|---|---|---|
| HR | 95% CI | p-value | HR | 95% CI | p-value | |
| Age (<60 yr vs ≥60 yr) | 1.129 | 0.607–1.129 | .702 | 2.231 | 0.856–5.817 | .101 |
| Sex (male vs female) | 0.694 | 0.376–1.282 | .244 | 0.469 | 0.200–1.099 | .081 |
| AJCC stage (I vs II, III and IV) | 4.975 | 2.535–9.764 | < .001 | 4.606 | 1.800–11.789 | .001 |
| T category (pT1 vs pT2, pT3, and pT4) | 2.661 | 1.391–5.089 | .003 | 4.663 | 1.713–12.694 | .003 |
| LN metastasis (absent vs present) | 4.314 | 2.313–8.045 | < .001 | 3.087 | 1.331–7.157 | .009 |
| Pleural invasion (absent vs present) | 1.397 | 0.702–2.782 | .341 | 3.488 | 1.406–8.656 | .007 |
| Histologic grade (grade 1 vs grade 2 and 3) | 3.667 | 1.130–11.898 | .030 | 2.367 | 0.552–10.149 | .246 |
| LVI (absent vs present) | 3.969 | 2.053–7.673 | < .001 | 3.677 | 1.488–9.085 | .005 |
| Perineural invasion (absent vs present) | 2.363 | 1.172–4.762 | .016 | 3.855 | 1.524–9.751 | .004 |
| miR-374a expression (low vs high) | 2.740 | 1.054–7.125 | .039 | 1.485 | 0.395–5.590 | .559 |
| miR-374a (low vs high), stage I only | 5.676 | 1.434–22.462 | .013 | 2.844 | 0.464–17.433 | .250 |
AJCC, American Joint Committee on Cancer.
Values are presented as number (%). AJCC, American Joint Committee on Cancer; LN, lymph node.
HR, hazard ratio; CI, confidence interval; AJCC, American Joint Committee on Cancer; LN, lymph node; LVI, lymphovascular invasion.

 E-submission
E-submission



