Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 48(1); 2014 > Article
-
Original Article
Distribution of Human Papillomavirus 52 and 58 Genotypes, and Their Expression of p16 and p53 in Cervical Neoplasia - Tae Eun Kim, Hwal Woong Kim, Kyung Eun Lee1
-
Korean Journal of Pathology 2014;48(1):24-29.
DOI: https://doi.org/10.4132/KoreanJPathol.2014.48.1.24
Published online: February 25, 2014
Department of Pathology, Good Moonhwa Hospital, Busan, Korea.
1Department of Clinical Laboratory Science, College of Health Sciences, Catholic University of Pusan, Busan, Korea.
- Corresponding Author: Kyung Eun Lee, Ph.D. Department of Clinical Laboratory Science, College of Health Sciences, Catholic University of Pusan, 57 Oryundae-ro, Geumjeong-gu, Busan 609-757, Korea. Tel: +82-51-510-0562, Fax: +82-51-510-0568, kelee@cup.ac.kr
© 2014 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- This study investigates the prevalence of human papillomavirus (HPV) 52 and 58 genotypes among women residing in Busan, and the expression of p16 and p53 proteins in cervical neoplasia with HPV 52 and 58 infections.
-
Methods
- A total of three hundred fifteen cases were analyzed using the HPV DNA chip test for HPV genotypes, and of these, we retrospectively examined p16 and p53 expression in 62 cases of cervical tissues infected with HPV 52 and 58 using immunohistochemistry.
-
Results
- HPV 52 and 58 genotypes were identified in 62 (54.9%) out of 113 high-risk, HPV-infected cases. Of the cases examined, there were 19 single HPV 52 infections (16.8%), 23 single HPV 58 infections (20.4%), 4 multiple HPV 52 infections (3.5%), and 16 multiple HPV-58 infections (14.2%). Immunoreactivity of p16 and p53 was observed in 41 (66.1%) and 23 (37.1%) of the 62 cases of cervical neoplasia infected with HPV 52 and 58 genotypes, respectively.
-
Conclusions
- This study demonstrates a high prevalence of HPV 52 and 58 genotypes, in addition to HPV 16, among high-risk strains of cervical neoplasia in Korea. These findings suggest that development of more vaccines would be beneficial for the prevention of the various HPV genotypes.
- Patients and tissue samples
- We examined three hundred fifteen cases of liquid-based cytology, and all samples were taken with a cervical brush during a gynecological examination for HPV DNA testing between January 2011 and December 2011. All HPV testing samples were obtained from gynecology outpatient clinic, and the punch biopsies were performed on the same day or within 6 months of HPV testing. A total of 315 cases were analyzed using the HPV DNA chip test (Biomed Lab Co., Seoul, Korea) for HPV genotypes, and of these, we retrospectively examined p16 and p53 expression in 62 cases of cervical tissues infected with HPV 52 and 58 using immunohistochemistry. The histologic grades of diagnoses were classified as either within normal limits (WNL), cervical intraepithelial neoplasia (CIN) I, CIN II, CIN III, squamous cell carcinoma in situ (CIS), or invasive carcinoma. The hematoxylin and eosin-stained slides were reviewed in each case to confirm the surgical pathologist's original diagnosis.
- HPV genotyping
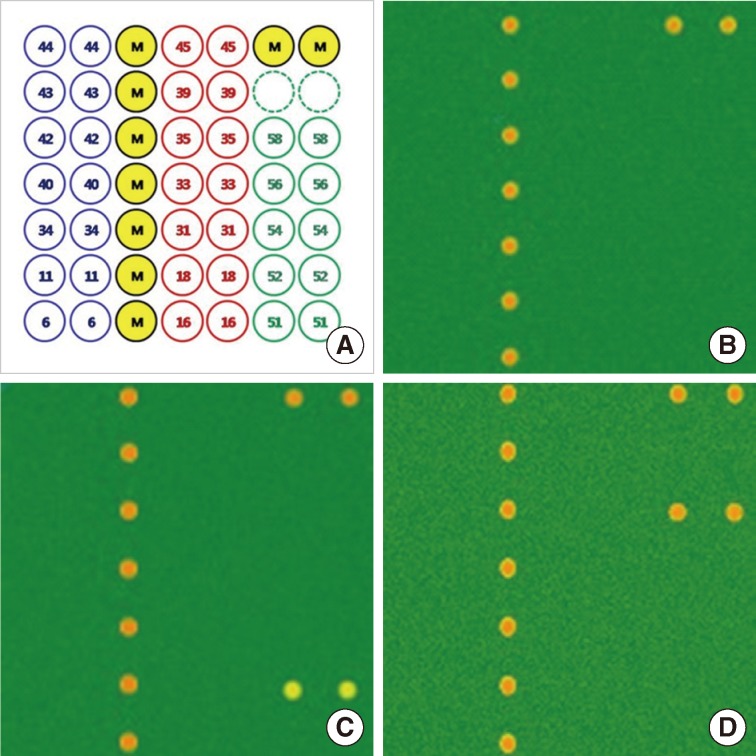
- For HPV genotyping, we used a commercially available HPV DNA chip kit (Biomed Lab), which is a polymerase chain reaction (PCR)-based DNA microarray system. The HPV DNA chip contained 12 types of high-risk HPV (HPV 16, 18, 31, 33, 35, 39, 45, 51, 52, 54, 56, 58) and 7 types of low-risk HPV (HPV 6, 11, 34, 40, 42, 43, 44) strains. The PCR products from all samples were subjected to electrophoresis on 2.5% agarose gels, and size of HPV DNA products was 150 base pairs. After 10 µL of the amplified HPV product was denatured at 95℃ for 5 minutes, it was mixed with a hybridization solution, and then applied to the DNA chip. Hybridized HPV DNA was visualized using a DNA chip scanner (GSI Lumonics, Scanarray Lite, Ottawa, Canada). HPV amplicons were hybridized with the corresponding type of specific oligonucleotide probe and visualized on HPV DNA chip slides as double positive spots.
- Immunohistochemistry
- The immunohistochemical portion of this study, focused on expression of proteins p16 and p53, was performed on formalin-fixed, paraffin-embedded, 4-µm thick tissue sections using the avidin-biotin-peroxidase complex method. Deparaffinization of all the sections was performed through a series of xylene baths, and rehydration was performed with a series of graded alcohol solutions. To enhance immunoreactivity, microwave antigen retrieval was performed at 750 W for 30 minutes in Tris-EDTA buffer (pH 9.0). After blocking endogenous peroxidase activity with 5% hydrogen peroxidase for 10 minutes, primary antibody incubation was performed for 1 hour at room temperature. The primary antibodies were a mouse monoclonal antibodies directed against p16 (JC 8, Santa Cruz Biotechnology, Santa Cruz, CA, USA) and p53 (DO-7, DakoCytomation, Glostrup, Denmark) used in 1:100 dilutions. An Envision Chem Kit (DakoCytomation, Carpinteria, CA, USA) was used as the secondary antibody at room temperature for 30 minutes. After washing the tissue samples in Tris-buffered saline for 10 minutes, 3,3'-diaminobenzidine was used as a chromogen, and Gill's hematoxylin counterstain was applied.
- Evaluation of immunohistochemisty
- All the slides were evaluated without knowledge of any clinicopathologic data. Immunoreactivity for p16 expression was defined by the presence of nuclear and cytoplasmic staining, and p53 expression was defined by the presence of nuclear staining. The percentage scores of immunoreactive tumor cells were categorized into the following four groups: 0 (0%), 1 (1-10%), 2 (11-50%), and 3 (>50%). The staining intensity was also categorized into four groups: 0 (negative), 1 (weak), 2 (moderate), and 3 (strong). A final score was obtained for each case by multiplying the percentage by the intensity score. Finally, tumors with multiplied scores that exceeded 2 (i.e., tumors with moderate and/or strong intensity of >10% of cells) were recorded as positively immunoreactive to p16 and p53; all the other scores were considered negative.
- Statistical analysis
- Statistical analysis was performed using SPSS ver. 17.0 (SPSS Inc., Chicago, IL, USA). The fisher exact test was performed to assess the relationship between the expression patterns of p16 and p53, and results were compared with the histological grades. A p-value less than .05 was considered to be statistically significant.
MATERIALS AND METHODS
- HPV genotypes analysis
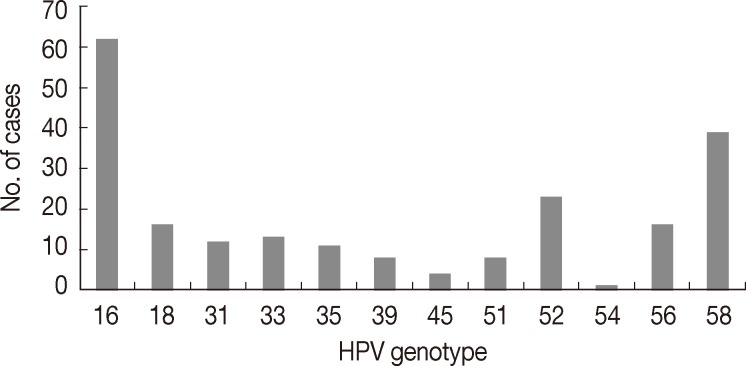
- The mean age of patients was 42.5 years (range, 22 to 83 years). Of the 315 cases, 221 cases (221/315, 70.2%) were found to be HPV DNA positive, of which 51.1% (113/221) had high-risk HPV infections, 4.5% (10/221) had low-risk HPV infections, 27.6% (61/221) had multiple infections and 16.7% (37/221) cases were confirmed to have other types of infections. HPV 16 was the most common genotype (62/221, 28.1%), followed by HPV 58 (39/221, 17.6%), HPV 52 (23/221, 10.4%), HPV 18 (16/221, 7.2%) and HPV 56 (16/221, 7.2%) (Fig. 1).
- HPV 52 and 58 genotypes were identified in 62 (54.9%) of 113 high-risk HPV infected cases. Of those 62, there were; single HPV 52 infections (19/113, 16.8%), single HPV 58 infections (23/113, 20.4%), multiple HPV 52 infections (4/113, 3.5%), and multiple HPV 58 infections (16/113, 14.2%) (Table 1, Fig. 2).
- The histologic diagnoses of the 62 cases of cervical tissues infected with HPV 52 and 58 genotypes were classified as follows: 17 (27.4%) cases of WNL, 20 (32.3%) cases of CIN I, 17 (27.4%) cases of CIN II/III, and 8 (12.9%) cases of CIS (Table 1).
- Expression of p16 and p53 protein
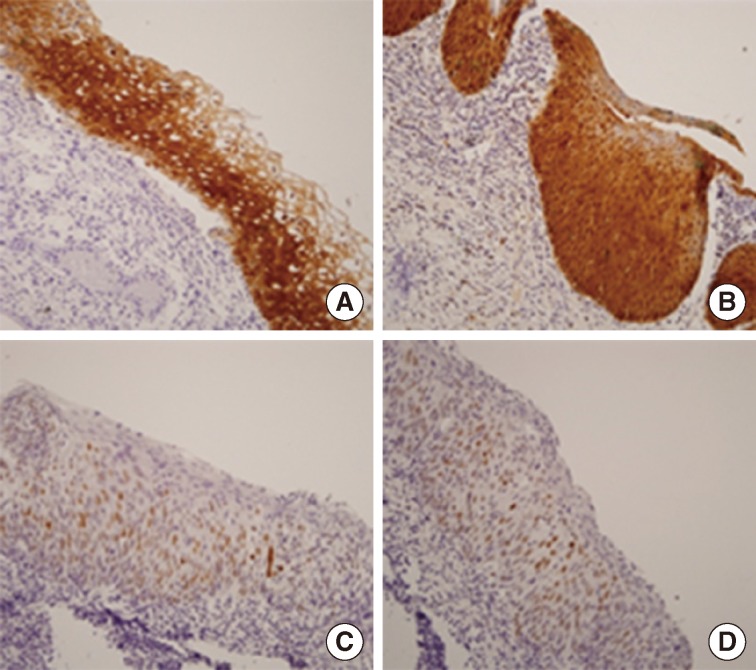
- Among the 62 cases of cervical tissues infected with HPV 52 and 58 genotypes, 41 (66.1%) were positive for p16 expression, and 21 (33.9%) were identified as p16 negative. Immunoreactivity of p16 was found in 58.8% (10/17) of WNL, 60.0% (12/20) of CIN I, 70.6% (12/17) of CIN II/III, and 87.5% (7/8) of CIS. Although it was not statistically significant, p16 overexpression tended to be associated with more advanced histologic grades of CIN (Table 2).
- Of 62 cervical tissues infected with HPV 52 and 58 genotypes, 23 (37.1%) cases were identified as p53 positive, while 39 cases (62.9%) did not express p53. The loss of p16 protein was found in 64.7% (11/17) of WNL, 55.0% (11/20) of CIN I, 70.6% (12/17) of CIN II/III, and 62.5% (5/8) of CIS (Table 2).
- Relationship between the expression of p16 and p53 in HPV 52 and 58 infected cervical neoplasia tissues
- Twenty-one (75.0%) of the 28 cases that positively expressed p16 immunoreactivity also showed negative p53 expression, although p16 overexpression was not significantly associated with the loss of p53 protein (p>.05) (Table 3, Fig. 3).
RESULTS
- While differences in the prevalence of HPV genotypes associated with cervical neoplasia exist between regions, the HPV 16 genotype is the most common strain worldwide.3 In this study, HPV 16 was the most prevalent at 28.1%, followed by HPV 58 at 17.6%, and HPV 52 at 10.4%. These results are similar to those of other studies examining the prevalence of HPV genotypes in Korea. The prevalence of HPV 58 has been demonstrated to be very different across global populations: 2.7% of Africans, 1.2% of Europeans, 0% of North Americans, 2% of East Asians, 4% of Japanese, and 23.8% of Hong Kong Chinese.5
- A large number of studies have related the inactivation of tumor suppressor genes to the development of cervical neoplasia. When tumors are HPV positive, the expression of HPV E6 and E7 proteins increases, which inactivates the tumor suppressor gene p53.10,11 In this study, p53 expression was positive in 23 (37.1%) cases and negative in 39 cases (62.9%) of cervical tissues infected with HPV 52 and 58 genotypes. While not statistically significant, the loss of p53 function in HPV-infected cervical neoplasia may be related to a loss of p53 proteins in cells. Among the 62 cases of HPV 52 and 58 infected cervical tissues, p16 expression was positive in 41 (66.1%) cases and negative in 21 (33.9%) cases. It is possible that the inactivation of Rb is caused by the HPV E7 protein, which may then lead to increased expression of p16 through a negative feedback mechanism. Keating et al.18 found that low-risk HPV genotypes reduce p16 expression, while high-risk HPV genotypes show different p16 levels according to the grading of lesions. For high-risk HPV genotypes, they reported an increase in p16 expression with cervical cancer progression.19,20 This study also reported that, p16 expression tended to increase with more advanced histologic grades of CIN, although this result was not statistically significant. Considering the high expression of p16 in 10 (58.8%) out of 17 cases of normal tissues infected with HPV 52 and 58 genotypes, we recommend that more studies should be conducted in this area, as p16 levels can serve as an indicator of infection by high-risk HPV genotypes. Song et al.21 reported significant relationships between HPV and p16, between HPV and p53, and between p16 and p53 in HPV-infected cervical neoplasia. In this study, p53 protein expression was generally positive when p16 was absent in cervical neoplasia infected with HPV 52 and 58 genotypes, but the results were not significant. While this may be interpreted as a loss of p53 due to HPV E6 and an increase in p16 from Rb inactivation due to HPV E7, some follow-up is needed due to the fact that this study was limited to 45 cases of cervical neoplasia infected with HPV 52 and 58 genotypes.
- Currently, there are only two vaccines available for the prevention of cervical cancer from HPV infection. Gardasil and Cervarix are at least 90% effective in preventing cervical cancer in women uninfected by HPV, but cross-protection has surfaced as a topic of interest with the growing concern over HPV types not covered by existing vaccines.15,16 According to studies conducted after the World Health Organization established a definition of "cross-protection" in 2007, bivalent vaccines are 31.6% effective against 6-month persistent infections of the HPV 52 genotype, while quadrivalent vaccines are 25% effective against 6-month persistent infections of the HPV 58 genotype.15,16,22 As demonstrated in this study, HPV genotypes other than HPV 16 and 18 also have a high prevalence, and so appropriate preventive measures must be put in place. More vaccines should be developed for the various HPV genotypes to prevent persistent infections and to reduce the initial infection rate from the current levels of 27.4% (17/62) in HPV 52 and 58 infected normal cervical tissues and 32.3% (20/62) in CIN I.
- This study found a high prevalence of HPV 52 and 58 genotypes, in addition to HPV 16, among high-risk strains in cervical neoplasia in Korean women. Overexpression of p16 and loss of the p53 protein have been demonstrated in a high percentage of cervical neoplasia infected with HPV 52 and 58 genotypes. Although these observations were not statistically significant, our results provide a valuable basis for additional studies involving a higher number of cases. To understand the mechanism of cervical cancer development caused by HPV 52 and 58 infections, more research is required on the relationships between cell cycle regulator p16, tumor suppressor p53 and other proteins.
DISCUSSION
Acknowledgments
Acknowledgment
- 1. Woodman CB, Collins SI, Young LS. The natural history of cervical HPV infection: unresolved issues. Nat Rev Cancer 2007; 7: 11-22. ArticlePubMedPDF
- 2. Shin HR, Jung KW, Won YJ, et al. National cancer incidence for the year 2002 in Korea. Cancer Res Treat 2007; 39: 139-149. ArticlePubMedPMCPDF
- 3. Muñoz N, Bosch FX, de Sanjosé S, et al. Epidemiologic classification of human papillomavirus types associated with cervical cancer. N Engl J Med 2003; 348: 518-527. ArticlePubMed
- 4. Lee JH, Lee KH, Lee IH, et al. Clinical efficacy of HPV DNA chip test in human papillomavirus detection of uterine cervix. Korean J Obstet Gynecol 2006; 49: 2128-2136.
- 5. Clifford GM, Smith JS, Plummer M, Muñoz N, Franceschi S. Human papillomavirus types in invasive cervical cancer worldwide: a meta-analysis. Br J Cancer 2003; 88: 63-73. ArticlePubMedPMCPDF
- 6. Huang S, Afonina I, Miller BA, Beckmann AM. Human papillomavirus types 52 and 58 are prevalent in cervical cancers from Chinese women. Int J Cancer 1997; 70: 408-411. ArticlePubMedPDF
- 7. Chansaenroj J, Junyangdikul P, Chinchai T, et al. Large scale study of HPV genotypes in cervical cancer and different cytological cervical specimens in Thailand. J Med Virol 2014; 86: 601-607. ArticlePubMed
- 8. Song JS, Kim EJ, Choi J, Gong G, Sung CO. Significance of HPV-58 infection in women who are HPV-positive, cytology-negative and living in a country with a high prevalence of HPV-58 infection. PLoS One 2013; 8: e58678.ArticlePubMedPMC
- 9. Choi IH, Jin SY, Lee DW, Kim DW, Jeen YM. Cytomorphologic features according to HPV DNA type in histologically proven cases of the uterine cervix. Korean J Pathol 2011; 45: 612-620.
- 10. Finlay CA, Hinds PW, Tan TH, Eliyahu D, Oren M, Levine AJ. Activating mutations for transformation by p53 produce a gene product that forms an hsc70-p53 complex with an altered half-life. Mol Cell Biol 1988; 8: 531-539. PubMedPMC
- 11. Levine AJ, Momand J, Finlay CA. The p53 tumour suppressor gene. Nature 1991; 351: 453-456. PubMed
- 12. Xiao S, Li D, Corson JM, Vijg J, Fletcher JA. Codeletion of p15 and p16 genes in primary non-small cell lung carcinoma. Cancer Res 1995; 55: 2968-2971. PubMed
- 13. Yoshida S, Todoroki T, Ichikawa Y, et al. Mutations of p16Ink4/CDKN2 and p15Ink4B/MTS2 genes in biliary tract cancers. Cancer Res 1995; 55: 2756-2760. PubMed
- 14. Lee S, Kim H, Kim H, Kim C, Kim I. The utility of p16INK4a and Ki-67 as a conjunctive tool in uterine cervical lesions. Korean J Pathol 2012; 46: 253-260. PubMedPMC
- 15. Lowy DR, Schiller JT. Prophylactic human papillomavirus vaccines. J Clin Invest 2006; 116: 1167-1173. PubMedPMC
- 16. Pollack AE, Balkin M, Edouard L, Cutts F, Broutet N. WHO/UNFPA Working Group on Sexual and Reproductive Health and HPV Vaccines. Ensuring access to HPV vaccines through integrated services: a reproductive health perspective. Bull World Health Organ 2007; 85: 57-63. PubMedPMC
- 17. Paavonen J, Jenkins D, Bosch FX, et al. Efficacy of a prophylactic adjuvanted bivalent L1 virus-like-particle vaccine against infection with human papillomavirus types 16 and 18 in young women: an interim analysis of a phase III double-blind, randomised controlled trial. Lancet 2007; 369: 2161-2170. PubMed
- 18. Keating JT, Cviko A, Riethdorf S, et al. Ki-67, cyclin E, and p16INK4 are complimentary surrogate biomarkers for human papillomavirus-related cervical neoplasia. Am J Surg Pathol 2001; 25: 884-891. PubMed
- 19. Klaes R, Friedrich T, Spitkovsky D, et al. Overexpression of p16 (INK4A) as a specific marker for dysplastic and neoplastic epithelial cells of the cervix uteri. Int J Cancer 2001; 92: 276-284. PubMed
- 20. Klaes R, Benner A, Friedrich T, et al. p16INK4a immunohistochemistry improves interobserver agreement in the diagnosis of cervical intraepithelial neoplasia. Am J Surg Pathol 2002; 26: 1389-1399. PubMed
- 21. Song SH, Park HM, Eom DW, et al. The expression of p16 (INK4a) and Ki-67 in relation to high-risk human papilloma viral load and residual disease after conization with positive margins. Int J Gynecol Cancer 2007; 17: 858-867. PubMed
- 22. Ault KA. Future II Study Group. Effect of prophylactic human papillomavirus L1 virus-like-particle vaccine on risk of cervical intraepithelial neoplasia grade 2, grade 3, and adenocarcinoma in situ: a combined analysis of four randomised clinical trials. Lancet 2007; 369: 1861-1868. PubMed
REFERENCES
| HPV genotypes |
Histology |
||||
|---|---|---|---|---|---|
| WNL | CIN I | CIN II/III | CIS | Total | |
| HPV 52 (single) | 8 | 6 | 3 | 2 | 19 |
| HPV 58 (single) | 3 | 8 | 10 | 2 | 23 |
| HPV 52 (multiple) | 1 | 2 | 0 | 1 | 4 |
| HPV 58 (multiple) | 5 | 4 | 4 | 3 | 16 |
| Total | 17 | 20 | 17 | 8 | 62 |
Figure & Data
References
Citations

- Screening for High-Risk Human Papillomavirus Reveals HPV52 and HPV58 among Pediatric and Adult Patient Saliva Samples
Hunter Hinton, Lorena Herrera, Sofia Valenzuela, Katherine M. Howard, Karl Kingsley
Dentistry Journal.2024; 12(3): 56. CrossRef - Usefulness Analysis of Urine Samples for Early Screening of Human Papilloma Virus Infection
Yoon Sung Choi, Hyunwoo Jin, Kyung Eun Lee
Journal of Cancer Prevention.2019; 24(4): 240. CrossRef - Relationship between Expression of P16 and Ki-67 and Persistent Infection of HPV in Cervical Carcinoma Patients
群欢 黄
Advances in Clinical Medicine.2018; 08(08): 776. CrossRef - Analysis of Sequence Variation and Risk Association of Human Papillomavirus 52 Variants Circulating in Korea
Youn Jin Choi, Eun Young Ki, Chuqing Zhang, Wendy C. S. Ho, Sung-Jong Lee, Min Jin Jeong, Paul K. S. Chan, Jong Sup Park, Xuefeng Liu
PLOS ONE.2016; 11(12): e0168178. CrossRef
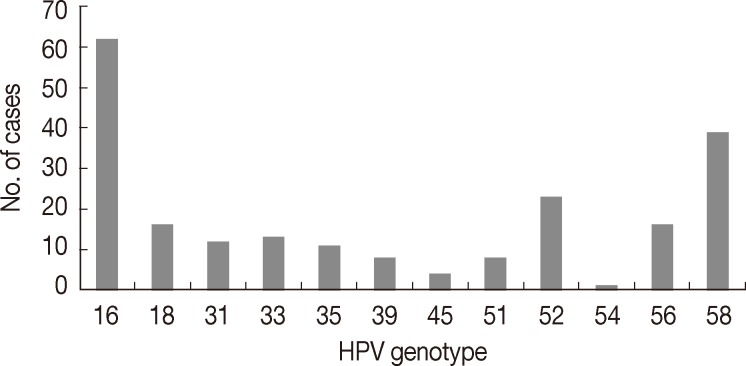
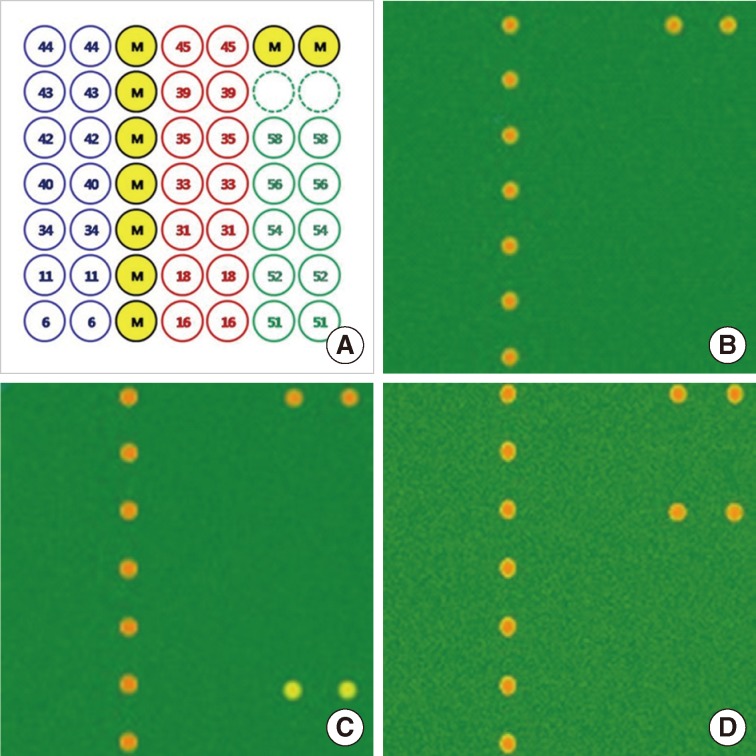
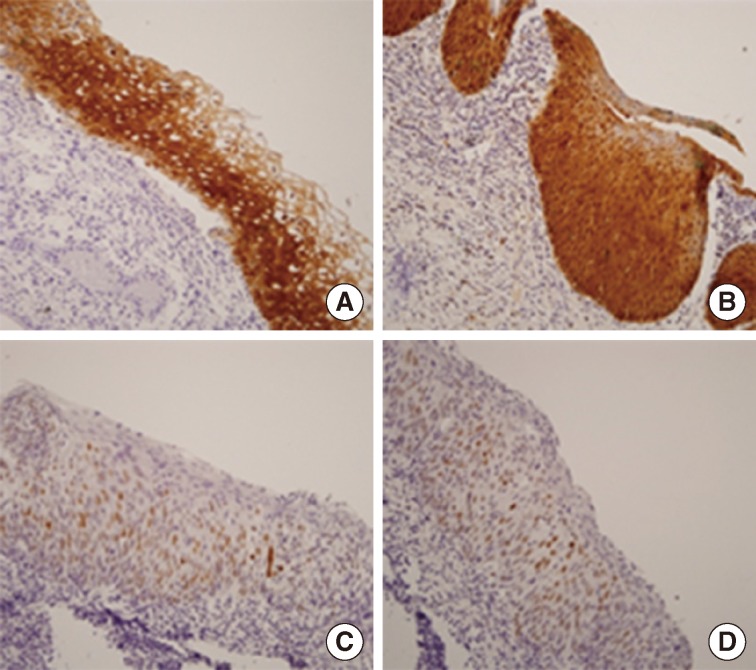
Fig. 1
Fig. 2
Fig. 3
| HPV genotypes | Histology |
||||
|---|---|---|---|---|---|
| WNL | CIN I | CIN II/III | CIS | Total | |
| HPV 52 (single) | 8 | 6 | 3 | 2 | 19 |
| HPV 58 (single) | 3 | 8 | 10 | 2 | 23 |
| HPV 52 (multiple) | 1 | 2 | 0 | 1 | 4 |
| HPV 58 (multiple) | 5 | 4 | 4 | 3 | 16 |
| Total | 17 | 20 | 17 | 8 | 62 |
| Histology | p16 expression |
p-value | p53 expression |
p-value | ||
|---|---|---|---|---|---|---|
| Negative | Positive | Negative | Positive | |||
| WNL | 7 (41.2) | 10 (58.8) | .47 | 11 (64.7) | 6 (35.3) | .80 |
| CIN I | 8 (40.0) | 12 (60.0) | 11 (55.0) | 9 (45.0) | ||
| CIN II/III | 5 (29.4) | 12 (70.6) | 12 (70.6) | 5 (29.4) | ||
| CIS | 1 (12.5) | 7 (87.5) | 5 (62.5) | 3 (37.5) | ||
| Total | 21 (33.9) | 41 (66.1) | 39 (62.9) | 23 (37.1) | ||
| Expression | p53 protein |
p-value | |
|---|---|---|---|
| Negative | Positive | ||
| p16 protein | .42 | ||
| Negative | 7 (25.0) | 7 (41.2) | |
| Positive | 21 (75.0) | 10 (58.8) | |
HPV, human papillomavirus; WNL, within normal limits; CIN, cervical intraepithelial neoplasia; CIS, squamous cell carcinoma
Values are presented as number (%). HPV, human papillomavirus; WNL, within normal limits; CIN, cervical intraepithelial neoplasia; CIS, squamous cell carcinoma
Values are presented as number (%). HPV, human papillomavirus.

 E-submission
E-submission
 PubReader
PubReader Cite this Article
Cite this Article




