Search
- Page Path
- HOME > Search
Original Articles
- Correlation between HER2 gene copy number and immunohistochemistry categories in HER2-negative breast cancer: diagnostic utility for differentiating HER2-null, ultralow, and low tumors
- Min Chong Kim, Young Kyung Bae
- J Pathol Transl Med. 2026;60(2):193-201. Published online February 25, 2026
- DOI: https://doi.org/10.4132/jptm.2025.11.07
- 1,373 View
- 166 Download
-
 Abstract
Abstract
 PDF
PDF - Background
The recent recognition of human epidermal growth factor receptor 2 (HER2)–low and HER2-ultralow breast cancers (BCs) has expanded the therapeutic relevance of HER2 testing in the antibody-drug conjugate era. However, the biological continuum of HER2 expression measured by immunohistochemistry (IHC) and its relationship with the HER2 gene copy number remain unclear. Methods: We retrospectively analyzed 135 HER2-negative invasive BCs and reclassified them as HER2-null (IHC 0), HER2-ultralow (0+), or HER2-low (1+ or 2+ without amplification). HER2 gene copy number was determined using silver-enhanced in situ hybridization. Statistical analyses were performed to compare HER2 copy number among IHC categories and evaluate the discriminatory value of HER2 copy number for distinguishing IHC subgroups. Results: The mean HER2 copy number increased stepwise across IHC categories: 1.95 ± 0.54 (null), 2.03 ± 0.43 (ultralow), 2.25 ± 0.65 (low, 1+), and 3.29 ± 1.05 (low, 2+). Significant differences were observed between the ultralow and low groups (p = .003) and between the null and low groups (p < .001), but not between the null and ultralow groups or between the ultralow and 1+ groups. Conclusions: HER2 gene copy number was positively correlated with protein expression as reflected by IHC categories. Although HER2 gene copy number was statistically higher in HER2-low than in HER2-null tumors, the substantial overlap in copy number ranges likely limits its utility in distinguishing HER2-low from HER2- null BCs.
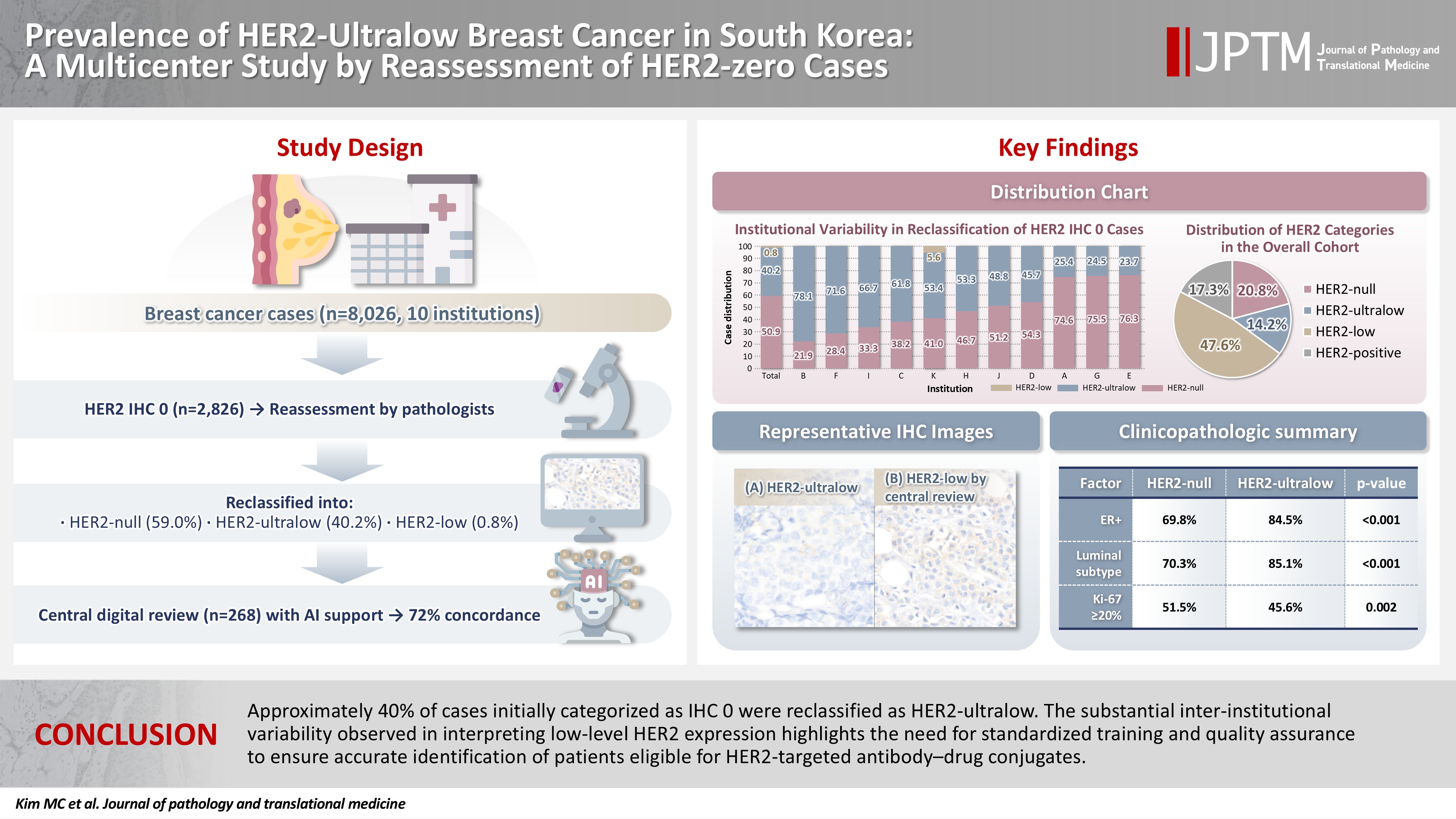
- Prevalence of HER2-ultralow breast cancer in South Korea: a multicenter study by reassessment of HER2-zero cases
- Min Chong Kim, Eun Yoon Cho, Hee Jin Lee, Ji Shin Lee, Jee Yeon Kim, Wan Seop Kim, Chungyeul Kim, Sun-Young Jun, Hye Jeong Choi, So Mang Lee, Ahrong Kim, Ji-Young Kim, Jeong Yun Shim, Gyungyub Gong, Young Kyung Bae
- J Pathol Transl Med. 2026;60(2):184-192. Published online February 23, 2026
- DOI: https://doi.org/10.4132/jptm.2025.10.22
- 1,795 View
- 170 Download
-
 Abstract
Abstract
 PDF
PDF Supplementary Material
Supplementary Material - Background
This study aimed to determine the prevalence of human epidermal growth factor receptor 2 (HER2)–ultralow breast cancer among cases initially classified as HER2 immunohistochemistry (IHC) 0 and assess interobserver variability in interpreting low-level HER2 expression. Methods: In this multicenter retrospective study, all invasive breast cancer cases diagnosed between January and December 2022 across 10 Korean institutions were retrieved. Institutional pathologists reexamined HER2 IHC slides originally reported as IHC 0 according to the 2018 American Society of Clinical Oncology/College of American Pathologists guidelines and reclassified them as HER2-null (0), HER2-ultralow (0+), or HER2-low (1+). Slides from 10% of HER2-null and HER2-ultralow cases were digitized for central review and independently assessed by two pathologists, with discrepancies resolved by consensus. Results: Among 8,026 cases, 2,836 cases (35.5%) were initially reported as IHC 0. Upon re-review, 1,673 (59.0%), 1,139 (40.2%), and 24 (0.8%) cases were reclassified as HER2-null, HER2-ultralow, and HER2-low, respectively. The prevalence of HER2-ultralow breast cancer varied considerably across institutions (23.7%–78.1%). Central review of 268 digitized cases showed concordance in 193 cases (72.0%). Among the 75 discordant cases, 54 tumors (72.0%) were upgraded from HER2-null to HER2-ultralow, and 18 (24.0%) tumors were upgraded from HER2-ultralow to HER2-low. Furthermore, two tumors (2.7%) were downgraded from HER2-ultralow to HER2-null. Conclusions: Approximately 40% of cases initially categorized as IHC 0 were reclassified as HER2-ultralow. The substantial inter-institutional variability observed in interpreting low-level HER2 expression highlights the need for standardized training and quality assurance to ensure accurate identification of patients eligible for HER2-targeted antibody–drug conjugates.
- CD9 Expression in Colorectal Carcinomas and Its Prognostic Significance
- Kyung-Ju Kim, Hee Jung Kwon, Min Chong Kim, Young Kyung Bae
- J Pathol Transl Med. 2016;50(6):459-468. Published online October 25, 2016
- DOI: https://doi.org/10.4132/jptm.2016.10.02
- 11,472 View
- 160 Download
- 19 Web of Science
- 17 Crossref
-
 Abstract
Abstract
 PDF
PDF - Background
CD9, a member of the tetraspanin superfamily, is a tumor suppressor in many malignancies. The aim of this study was to evaluate the immunohistochemical expression of CD9 in colorectal carcinomas (CRCs) and determine clinicopathological and prognostic significance of its expression.
Methods
The CD9 expression status of 305 CRCs was evaluated using a semi-quantitative scoring system in tumor cells (T-CD9) and immune cells (I-CD9) by classifying the results as high and low expression.
Results
High T-CD9 (T-CD9 [+]) expression was detected in 175 samples (57.6%) and high I-CD9 (I-CD9 [+]) expression was detected in 265 samples (86.9%). Using Kaplan- Meier survival analysis, the T-CD9 (+) group showed a tendency for better disease-free survival (DFS) (p = .057). In left-sided tumors, DFS was significantly longer in the T-CD9 (+) group (p = .021) but no statistical significance was observed with right-sided tumors (p = .453). I-CD9 (+) CRCs significantly correlated with well/moderately differentiation (p = .014). In Kaplan-Meier analysis, the I-CD9 (+) group had a tendency towards worse DFS compared to the I-CD9 (–) group (p = .156). In combined survival analysis of T-CD9 and I-CD9, we found that the longest DFS was among patients in the T-CD9 (+)/I-CD9 (–) group, whereas the T-CD9 (–)/I-CD9 (+) group showed the shortest DFS (p = .054).
Conclusions
High expression of T-CD9 was associated with a favorable DFS, especially in left-sided CRCs. Combined evaluation of T-CD9 and I-CD9 is required to determine the comprehensive prognostic effect of CD9 in CRCs. -
Citations
Citations to this article as recorded by- Prognostic Significance of Nuclear Factor Kappa B and CD9/Motility-related Protein-1 in Stage II-III Rectal Cancer Patients Treated with Postoperative Chemoradiotherapy
Serkan KAPLAN, Oğuz Galip YILDIZ, Işın SOYUER, Serdar SOYUER
Journal of Eurasian Medical Science.2026; : 42. CrossRef - Predicting patient outcomes with gene-expression biomarkers from colorectal cancer organoids and cell lines
Alexandra Razumovskaya, Mariia Silkina, Andrey Poloznikov, Timur Kulagin, Maria Raigorodskaya, Nina Gorban, Anna Kudryavtseva, Maria Fedorova, Boris Alekseev, Alexander Tonevitsky, Sergey Nikulin
Frontiers in Molecular Biosciences.2025;[Epub] CrossRef - Intestinal estrogen receptor beta modulates the murine colon tumor immune microenvironment
Madeleine Birgersson, Matilda Holm, Carlos J. Gallardo-Dodd, Baizhen Chen, Lina Stepanauskaitė, Linnea Hases, Claudia Kutter, Amena Archer, Cecilia Williams
Cancer Letters.2025; 622: 217661. CrossRef - Establishment of Coala: a novel 3D and 2D cancer cell line derived from colorectal cancer liver metastasis
Sandra Mersakova, Juraj Marcinek, Dusan Brany, Olga Chodelkova, Martin Vorcak, Martin Cermak, Martina Poturnajova, Zuzana Kozovska, Henrieta Skovierova, Slavomira Novakova, Barbora Mitruskova, Dusan Loderer, Marian Grendar, Veronika Holubekova, Andrea Hor
Human Cell.2025;[Epub] CrossRef - CD9, a tetraspanin in cancer: biology and therapeutic promise in acute leukemia
Océane Guého, Elie Cousin, Jérémie Rouger-Gaudichon, Anne-Gaëlle Rio, Sébastien Corre, Virginie Gandemer, Frédéric Mazurier
Oncogenesis.2025;[Epub] CrossRef - Proteomic analysis of plasma exosomes in patients with metastatic colorectal cancer
Zhaoyue Zhong, Jiayin Ji, Hongxia Li, Ling Kang, Haipeng Zhu
Clinical Proteomics.2024;[Epub] CrossRef - Prognostic value and multifaceted roles of tetraspanin CD9 in cancer
Róbert Ondruššek, Barbora Kvokačková, Karolína Kryštofová, Světlana Brychtová, Karel Souček, Jan Bouchal
Frontiers in Oncology.2023;[Epub] CrossRef - Proteomic Signature of Extracellular Vesicles Associated with Colorectal Cancer
Natalia Soloveva, Svetlana Novikova, Tatiana Farafonova, Olga Tikhonova, Victor Zgoda
Molecules.2023; 28(10): 4227. CrossRef - Anti-Human CD9 Fab Fragment Antibody Blocks the Extracellular Vesicle-Mediated Increase in Malignancy of Colon Cancer Cells
Mark F. Santos, Germana Rappa, Simona Fontana, Jana Karbanová, Feryal Aalam, Derek Tai, Zhiyin Li, Marzia Pucci, Riccardo Alessandro, Chikao Morimoto, Denis Corbeil, Aurelio Lorico
Cells.2022; 11(16): 2474. CrossRef - The Study of the Extracellular Matrix in Chronic Inflammation: A Way to Prevent Cancer Initiation?
Asia Marangio, Andrea Biccari, Edoardo D’Angelo, Francesca Sensi, Gaya Spolverato, Salvatore Pucciarelli, Marco Agostini
Cancers.2022; 14(23): 5903. CrossRef - In vivo expansion of a CD9+ decidual-like NK cell subset following autologous hematopoietic stem cell transplantation
Ane Orrantia, Enrique Vázquez-De Luis, Gabirel Astarloa-Pando, Iñigo Terrén, Ainhoa Amarilla-Irusta, Diego Polanco-Alonso, Carmen González, Alasne Uranga, Tomás Carrascosa, Juan J. Mateos-Mazón, Juan C. García-Ruiz, Sergio Callejas, Ana Quintas, Ana Dopaz
iScience.2022; 25(10): 105235. CrossRef - Inhibition of cancer-cell migration by tetraspanin CD9-binding peptide
Thanawat Suwatthanarak, Masayoshi Tanaka, Yoshitaka Miyamoto, Kenji Miyado, Mina Okochi
Chemical Communications.2021; 57(40): 4906. CrossRef - High expression of tumor susceptibility gene 101 (TSG101) is associated with more aggressive behavior in colorectal carcinoma
Elmira Gheytanchi, Leili Saeednejad Zanjani, Roya Ghods, Maryam Abolhasani, Marzieh Shahin, Somayeh Vafaei, Marzieh Naseri, Fahimeh Fattahi, Zahra Madjd
Journal of Cancer Research and Clinical Oncology.2021; 147(6): 1631. CrossRef - Increased CD9 expression predicts favorable prognosis in human cancers: a systematic review and meta-analysis
Hyun Min Koh, Bo Gun Jang, Dong Hui Lee, Chang Lim Hyun
Cancer Cell International.2021;[Epub] CrossRef - Prognostic Value of CD9 in Solid Tumor: A Systematic Review and Meta-Analysis
Ping Zeng, Meng Si, Rui-xia Sun, Xu Cheng, Xiao-yang Li, Min-bin Chen
Frontiers in Oncology.2021;[Epub] CrossRef - Matrix Effect in the Isolation of Breast Cancer-Derived Nanovesicles by Immunomagnetic Separation and Electrochemical Immunosensing—A Comparative Study
Silio Lima Moura, Mercè Martì, María Isabel Pividori
Sensors.2020; 20(4): 965. CrossRef - CD9 expression indicates a poor outcome in acute lymphoblastic leukemia
Peiqi Liang, Miao Miao, Zhuogang Liu, Hongtao Wang, Wei Jiang, Shiyu Ma, Chuan Li, Rong Hu
Cancer Biomarkers.2018; 21(4): 781. CrossRef
- Prognostic Significance of Nuclear Factor Kappa B and CD9/Motility-related Protein-1 in Stage II-III Rectal Cancer Patients Treated with Postoperative Chemoradiotherapy

 E-submission
E-submission


 First
First Prev
Prev



