Difference of Genome-Wide Copy Number Alterations between High-Grade Squamous Intraepithelial Lesions and Squamous Cell Carcinomas of the Uterine Cervix
Article information
Abstract
Background
About 10% of high-grade squamous intraepithelial lesions (HSILs) progress to invasive carcinomas within 2-10 years. By delineating the events that occur in the early stage of the invasion, the pathogenesis of cervical cancer could be better understood. This will also propose the possible methods for inhibiting the tumor invasion and improving the survival of patients.
Methods
We compared the genomic profiles between the HSIL and the invasive squamous cell carcinoma (SCC) using an array comparative genomic hybridization. Using recurrently altered genes, we performed a principal component analysis to see variation of samples in both groups. To find possibly affected pathways by altered genes, we analyzed genomic profiles with the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway database and GOEAST software.
Results
We found 11q12.3 and 2p24.1 regions have recurrent copy number gains in both groups. 16p12-13 and 20q11-13 regions showed an increased copy number only in cases of HSIL. 1q25.3 and 3q23-29 regions showed copy number gains only in cases of SCC. Altered genes in the SCC group were related to the mitogen-activated protein kinase signaling pathway and the RNA transport. Altered genes in the HSIL group were related to the ubiquitin mediated proteolysis and cell adhesion molecules.
Conclusions
Our results showed not only that gains in 11q12.3 and 2p24.1 were early events occurring in the premalignant lesions and then maintained in cases of SCC but also that gains in 1q25.3 and 3q23-29 were late events occurring after invasion in those of SCC.
Uterine cervical cancer is one of the most common malignancies that occur in women. With the widespread use of a screening method, the mortality of cervical cancer has been greatly decreased and the early detection of it has become possible. Nonetheless, the cervical cancer remains one of the major health problems in women.
There are several histological types of cervical cancer. The two most common types of cervical cancer include squamous cell carcinoma (SCC) and adenocarcinoma. SCC accounts for approximately 80% of total cases of cervical cancer.1 The most well-known cause of cervical cancer is human papillomavirus (HPV). Over 100 types of HPV have been introduced to date.2 High-risk HPV is known as the most important factor in cervical carcinogenesis.1 It is also well known that high-grade squamous intraepithelial lesion (HSIL) predate the onset of invasive cervical cancer. In only a small portion of women with high-risk HPV, however, cervical cancer occurs. Little is known about the genomic alterations during transition from HSIL to invasive cancer. In an effort to clarify the pathogenesis of cervical cancer, studies have been conducted focusing other causative events than HPV infections.3,4 It is known that HPV stimulates the cell cycle by interfering with the function of Rb and p53 in host cells.1 In particular, its two viral proteins, E6 and E7, play a key role in the pathogenesis of uterine cervical cancer. These viral proteins have oncogenic effects, of which the centrosome duplication and genome instability can result in the alteration of copy number in the genome of host cells.
The alteration of copy number is one of mechanisms by which oncogenes are activated. As a good example, two oncogenes such as N-myc and erbB2/neu are activated by the increased copy number. The clinical significance of the activation of these oncogenes has been verified in cases of neuroblastoma and breast cancer in the corresponding order.5,6 In addition, the role of the alteration of copy number in carcinogenesis has been reported to be of clinical significance in other several human malignancies including lung cancer, prostate cancer, and colorectal cancer.7-9
To date, a comparative genomic hybridization (CGH) has been used to identify genome-wide changes in the copy number in cases of cervical cancer. Due to a lower resolution of metaphase chromosome CGH, it gives the information about the cytoband level. In addition, it is difficult to detect the alterations of the narrow region. Since an array CGH using BAC/PAC and oligonucleotide probes was first introduced, the resolution of CGH has been improved to such a sufficient extent that even the changes limited to specific genes or exons could be detected.10
Up to present, there are several published studies about the alteration of copy number.3,11-15 In most of these studies, the fluorescent in situ hybridization (FISH) or CGH have been used to detect any changes in cases of cervical cancer.
Due to the limitation of useful research modalities, however, most of the studies have reported that the alterations occurred in both very wide and various regions. It is therefore difficult to delineate their genetic contribution to the pathogenesis of cervical cancer. It has been reported that deletions at 3p and amplifications of 3q are associated with HPV-16.1 It has been reported that there are deletions at 2p, 2q34-q36, 3p14, 4 and 6p and gains at 1, 3q, 5ptel, 5p15.2 and 5p13 in cases of cervical cancer.13-16 According to some reports, a certain type of the alteration of copy number is detected in the specific stages of cervical cancer. Gain of chromosome arm 3q is consistently found in cases of advanced-stage cervical carcinomas.17 Losses of 3p, 11q, 6q and 10q and gain of 3q are frequently detected in the primary cervical tumor according to lymph node status.18 Gain of 3q and loss of 11q are early events in the progression of cervical cancer.19 Some of alterations have been reported to be of clinical significance. 9p deletions are more frequent in cases of carcinoma with a lymph node metastasis. In addition, losses of chromosome 11p and 18q are associated with a poor prognosis in cases of cervical carcinoma without a lymph node metastasis.20 It has been reported that gain of chromosome 3q is associated with the transition from severe dysplasia to invasive carcinoma of the uterine cervix.21 But there were no studies focusing on the genomic difference between HSIL and invasive SCC (stage IIb) of uterine cervix using a high-resolution array CGH. The morbidity and mortality of the uterine cervical cancer are closely related to the invasion and metastasis of cancer cells. The difference between HSIL and invasive SCC should be clearly delineated, which will provide clues about the pathogenesis of invasion in cases of cervical cancer.
Given the above background, we conducted this study to compare the genomic profiles and the differences in the genetic pathway and gene ontology between HSIL and SCC using an array-based CGH by which changes in gene level can be detected.
MATERIALS AND METHODS
Primary tumor tissues
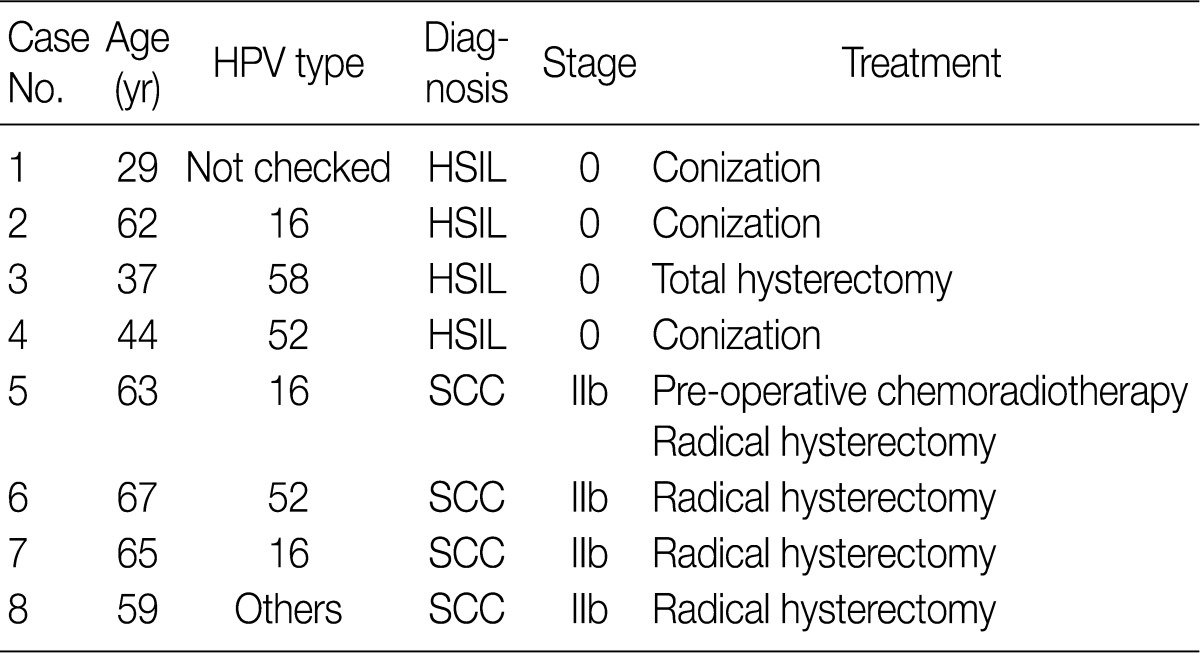
In eight cases of uterine cervix tissues, the tissue samples were fixed in a formalin solution and then prepared into paraffin-embedded sections (FFPE). The protocol of the current study was approved by the Institutional Review Board (IRB) (CUMC10U917) of the Catholic University of Korea, College of Medicine. These eight cases consisted of four cases of HSIL and another four cases of invasive SCC (Table 1).
Genomic DNA extraction
Unstained tissue sections were cut from the paraffin blocks and then deparaffinized with xylene twice. Normal areas of cervical tissues were removed from the sections using knife blades. The remaining tissues were collected in 1.5 mL microtubes and then digested for 72 hours in the dry chamber, where the temperature was set at 58℃, using Proteinase K (Invitrogen, Carlsbad, CA, USA) in ATL buffer (Qiagen, Valencia, CA, USA) with overlaid mineral oil (Sigma-Aldrich, St. Louis, MO, USA) that was applied to hinder the evaporation and concentration of the buffer. After the tissue digestion, genomic DNAs were extracted using DNeasy blood and tissue kit (Qiagen) according to the manufacturer's protocol. The concentration of the DNA was measured using NanoDrop (Thermo Scientific, Wilmington, DE, USA) and the integrity of the DNA was checked with a 1% agarose gel electrophoresis.
DNA fragmentation
Usually the genomic DNA extracted from FFPE tissue is partially degraded and chemically modified.22 The FFPE DNA produces a high-level of noise with standard array CGH protocol. To minimize the level of noise in an array CGH data, random fragmentation method using Shrimp Recombinant DNase (Affymetrix, Santa Clara, CA, USA) was exploited.22 Briefly, a 2.5 µg of extracted DNA and a 1 µg of reference DNA (Promega, Madison, WI, USA) were digested with the Shrimp Recombinant DNase (Affymetrix) to get DNA fragments of 200 to 400 bp in size.
Array CGH
Digested fragments of extracted DNA and reference DNA were labeled with Cyanine-5 and Cyanine-3 dUTP, respectively, using the Bioprime array CGH genomic labeling system (Invitrogen). After labeling reaction, the amount of labeled product and the efficiency of labeling were measured using Nanodrop (Thermo Scientific). Each labeled DNA was co-hybridized at an amount of 4 µg, identical to that of labeled reference DNA, to the 4×44K human aCGH array platform (Agilent Technologies, Santa Clara, CA, USA). Hybridization was performed at 65℃, for which the array chamber was rotated at 20 rpm for 24 hours. Then the arrays were washed according to the manufacturer's instructions. Microarray slides were scanned on a GenePix 4000B microarray scanner (Molecular Devices, Sunnyvale, CA, USA). The scanned images were analyzed using Feature Extraction software (Agilent Technologies).
Microarray data analysis
To define segments with an altered copy number, we used the aberration detection method 2 (ADM-2) algorithm with a threshold of 6 using DNA analytics software (Agilent Technologies).23 To minimize false positives, we excluded aberrations spanning less than three consecutive probes from further analysis. To focus on recurrent changes in each group, we used aberrations found in at least two cases (50%) of each group. Besides, we used R 2.13.1 software (R Foundation for Statistical Computing, Vienna, Austria) and Matlab 7.12 software (Mathworks, Natick, MA, USA) for statistical calculations and graphical presentations.
Real-time polymerase chain reaction (PCR) of syndecan1 (SDC1) gene
Of the samples that showed an increased copy number of SDC1 on an array CGH, three cases were selected to confirm the copy number gain of SDC1 in the cervical lesion. A 20 ng of genomic DNA extracted from the cervical lesion was used for each reaction. SYBR Premix Ex Taq (Takara Bio Inc., Shiga, Japan) was used to amplify target (SDC1) and reference (beta actin, ACTB) genes using a LightCycler 480 II (Roche, Rotkreuz, Switzerland). We designed two pairs of primers for SDC1 to detect copy number of two regions (exon 2 and exon 3) of SDC1 and one pair for ACTB as reference. The following oligonucleotides were used as the primer: SDC1-2F (exon 2) 5'-GTTTTTGCAACGGCTAAGGA-3'; SDC1-2R (exon 2) 5'-TCTACTGCCGGATTCCTCTC-3'; SDC1-3F (exon 3) 5'-CGGCCAAGCCTTTACTCATA-3'; SDC1-3R (exon 3) 5'-AGCCGGAGAAGTTGTCAGAG-3'; ACTB-F 5'-AGAAAATCTGGCACCACACC-3'; ACTB-R 5'-AACGGCAGAAGAGAGAACCA-3'.
After the pre-incubation at 95℃ for 5 minutes, 45 cycles of the PCR (95℃ for 30 seconds, 60℃ for 30 seconds, and 72℃ for 30 seconds) were performed. Then, the melting curves were analyzed to confirm specific PCRs; all samples were performed in triplicate. We used pooled genomic DNA extracted from the stromal tissues in the same cases as a calibrator. We used the E-method of LightCycler 480 II (Roche) to calculate template concentration, where we normalized the copy number of the target and reference using the calibrator.
Pathway analysis
To find possibly affected pathways consisting of altered genes in cases of cervical cancer, we searched the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway database (http://www.genome.jp/kegg/pathway.html).
Gene ontology (GO) analysis
To analyze a set of genes with commonly altered copy numbers based on a gene ontology, we used Gene Ontology enrichment analysis software toolkit (GOEAST).24 The list of gene symbols showing a recurrent aberration of the copy number was submitted to the GOEAST. Of the GO term IDs that the GOEAST identified from the list, those with p-value of <0.05 were considered to be statistically significant.
RESULTS
Distribution of the alteration of copy number
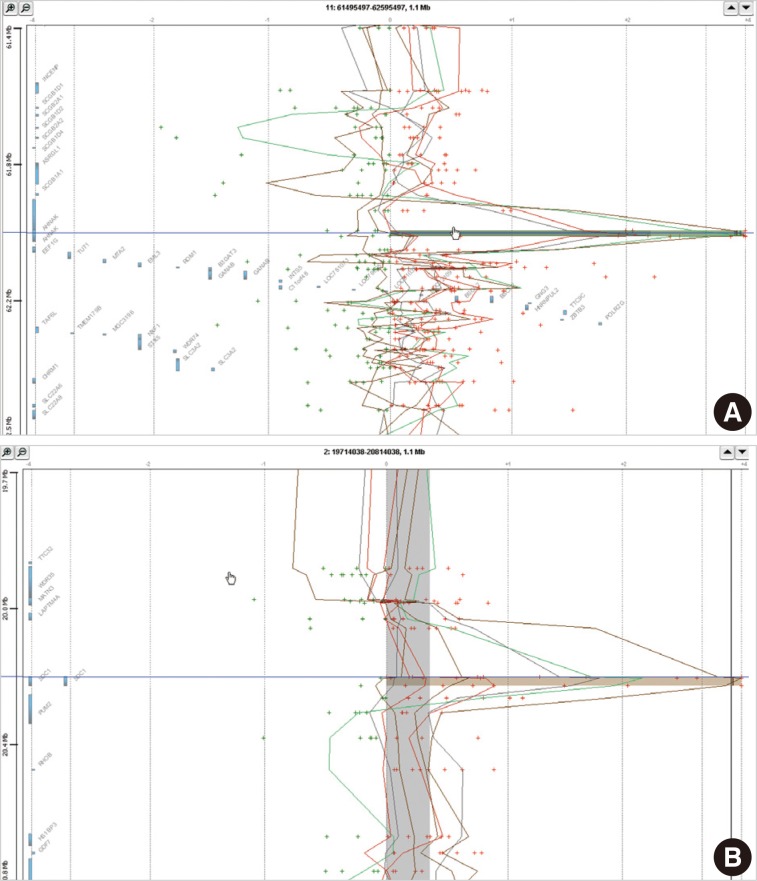
In eight cases of cervical cancer, we found 18,472 microarray probes showing the alteration of copy number. To find recurrent aberrations in each group, we collected aberrations detected in at least two cases of each group (Table 2). No regions showed a decreased copy number in two or more cases in each group. 11q12.3 (AHNAK, TRIM29) and 2p24.1 (SDC1) were found to show a copy number gain in both the HSIL group and the SCC group (Fig. 1). The recurrent aberrations of the 16p12-13 and 20q11-13 regions were found only in the HSIL group and those of the 1q25.3 (IVNS1SBP) and 3q23-29 regions were found only in the SCC group.
Principal component analysis (PCA)
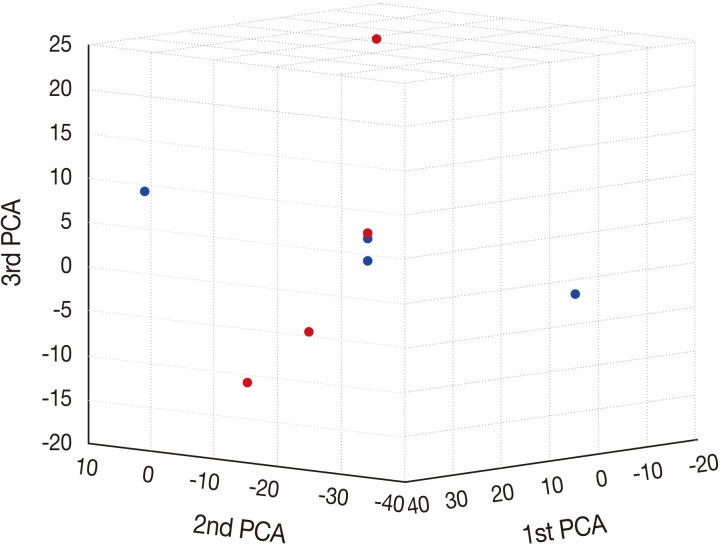
To characterize the aberration profiles of the tissue samples of HSIL and SCC, we performed the PCA using all aberration information (Fig. 2). There was a variability in the first, second and third factors forming the principal components, approximately showing an orthogonal linear relationship between HSIL and SCC. This indicates that there is a distinct difference in the pattern of the variation between HSIL and SCC based on their genomic profile.
Analysis of the KEGG pathway database
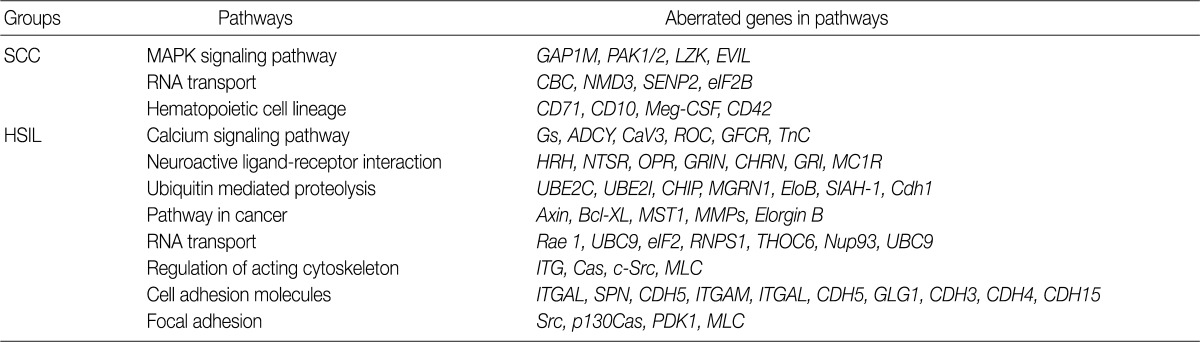
To find possibly affected pathways consisting of altered genes in cases of cervical cancer, we searched the KEGG pathway database (Table 3). The mitogen-activated protein kinase signaling pathway is found in genes with an aberration of the copy number in cases of SCC. In cases of HSIL with an alteration of the copy number, pathways related to ubiquitin-mediated proteolysis and cell adhesion molecules were found. RNA transport pathway was found in both groups.
GO analysis
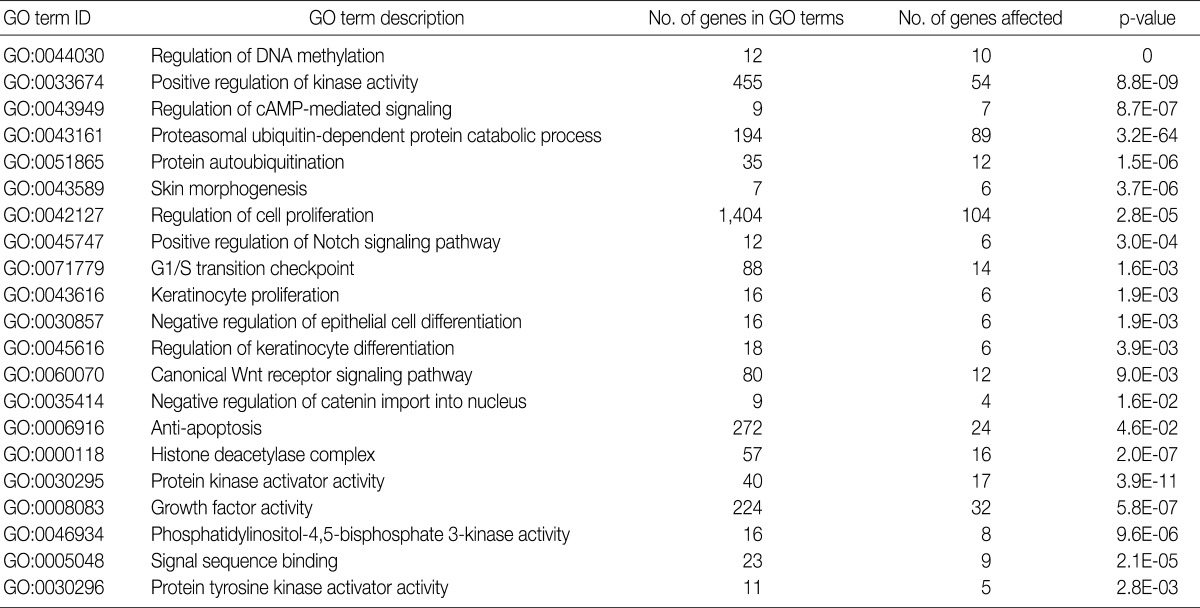
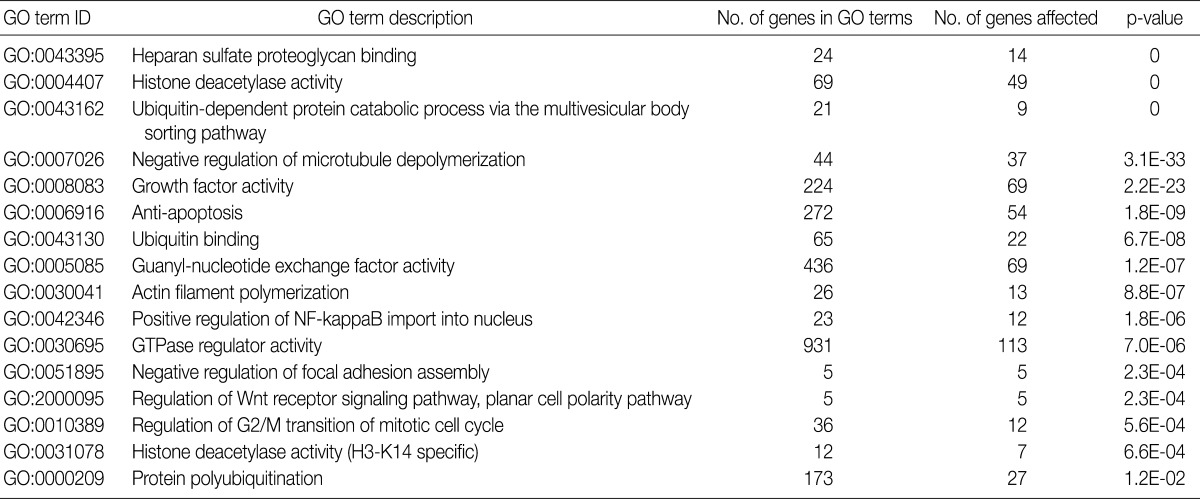
Using the GOEAST, we found 757 significant GO term IDs from the list of genes with an alteration of the copy number only in the SCC group. We also found 407 GO term IDs from the list of genes with an alteration of the copy number in both groups. Of the GO term IDs that had been found, representative cancer-related IDs are represented in Tables 4 and 5.
The alteration of copy number of SDC1 gene
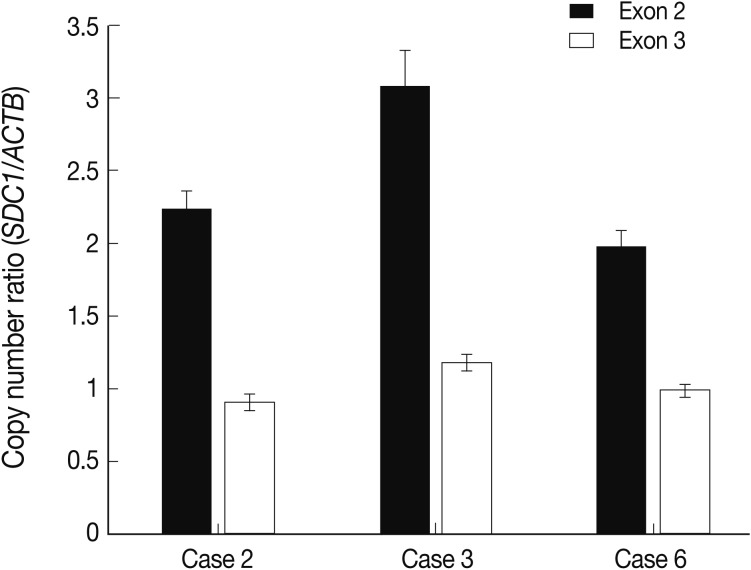
Of the genes with an aberration of the copy number, we were interested in SDC1 whose gene product is involved in the cell binding, cytoskeletal organization and might play a role in the tumor invasion and metastasis of the tumor. To confirm the dose of an alteration of the copy number in SDC1 gene, we performed the real-time PCR. There three cases of an increased copy number in the region of exon 2 but not in that of exon 3 (Fig. 3).
DISCUSSION
In the current study, we screened whole genomes with any recurrent alterations of the copy number in cases of cervical SCC. We also analyzed a set of genes with an alteration of the copy number using bioinformatics database and tools. Our results were partly in agreement with previous reports about cervical cancer. Of the alterations of the copy number that have been reported up to present, gains of 1q and 3q are frequently mentioned on a variety of reports about cervical cancer.3,11,12 Our results showed that the gain of 3q was a pivotal aberration during the transition from HSIL to invasive SCC.21 Using an array CGH, we narrowed down the region to 3q23-29 and thereby detected another recurrent aberration of IVNS1ABP during the above transition. By contrast, a loss of the copy number has not been frequently reported on the articles. We could not also identify any recurrent losses of the copy number in the current study. This suggests that a gain of the copy number rather than a loss of it might be more relevant in the pathogenesis of cervical cancer.
Advanced cancers are known to have chromosomal instabilities. Due to the instability, cancer genomes harbor many non-specific patterns of the alteration of copy number. By focusing on relatively earlier stage of SCC (International Federation of Genycology and Obstetrics [FIGO] stage IIb) and comparing its findings with those of premalignant lesions (HSIL), we were successful in not only demonstrating that there was a difference in the alteration between SCC and HSIL but also minimizing nonspecific patterns of the alterations due to the chromosomal instability.
In cancer cells cultured in in vitro conditions, there is a tendency to randomly accumulate the alteration of copy number due to chromosomal instabilities.25 Cell lines, derived from malignant cells and grown in culture condition, harbor many nonspecific patterns of the alteration of copy number. This is seen after the collection of tissue samples from patients. It is therefore probable that reports about the alterations of copy number using cell lines may not be applicable to cancer cells in an in vivo condition. To minimize this bias of cell line study, we used primary malignant and premalignant tissues to investigate the alteration of copy number.
Of the alteration of copy number that had been reported from cervical cancer, some had a prognostic value. Oncostatin M receptor (OSMR) frequently shows a copy number gain in cervical cancer, which is associated with adverse clinical outcomes.26 Our results showed, however, that there was no recurrent aberration of the copy number in the 5p13.1 region where OSMR is located. This discrepancy can be explained by OSMR gain is a late event in cervical carcinogenesis as the authors26 mentioned. In their report, more than half of the samples they studied were of FIGO stage IIIb or higher.
Another variant of cervical cancer, cervical adenocarcinoma shows different patterns of the alteration of copy number from SCC. It has been reported that cervical adenocarcinomas have frequent gains at 3q, 17q, 1p, 1q, and 11q.27 According to these authors,27 the gain was more commonly seen than the loss of copy number. This was also shown in the current study. In addition, 3q28-29 gain was found in both the above authors' reports and our findings.
It is not always observed that the alteration of copy number results in that of gene expression. In about half of total cases, the alteration of the copy number leads to that of the gene expression. According to the combination study about the dosage and expression of genes, EPHB2, MMP1, AKT1, ABCC3 genes are both amplified and over-expressed in cases of cervical cancer.28
To analyze the possible net effect of genomic alterations, we searched for the KEGG pathway database with lists of genes with an alteration of the copy number. In cases of HSIL, genes with an alteration of the copy number are related to both the pathways of ubiquitin-mediated proteolysis and adhesion related pathways (Table 3). HPV E6 induces the rapid degradation of p53 via ubiquitin-dependent proteolysis.1 In cases of HSIL, the recurrent copy number gain of genes related to ubiquitin-mediated proteolysis might explain the possible underlying mechanisms by which the rapid degradation of p53 in HPV-infected cells occurs before entry into the invasive stage of SCC.
There are no reports explaining the reason why there was no persistent presence of the alteration of copy number in cases of HSIL. This might be due to chromosomal instability in malignant cells.
We have previously reported an increased expression of SDC1 in cases of cervical cancer and its association with a prolonged survival.29 In the previous report, we performed an interphase FISH to investigate the alteration of copy number of SDC1 but in vain. The detection limit of FISH is generally set at 10 Mbp. Conventional types of the FISH analysis could not therefore detect any changes with a span of <10 Mbp. If the alteration of copy number of SDC1 gene is partly present, as we shown in the current study, it might not be detected on the FISH analysis. This explains the discrepancy between our previous report and the current study.
Limitations of the current study are due to a relatively smaller number of samples. In the current study, we used only four samples of HSIL and another four samples of invasive SCC. Due to a smaller number of samples, the nature of genome-wide alteration of copy number may not be fully explained in cases of HSIL and those of invasive SCC. Nevertheless, we applied stringent methods to minimize false positives and get pertinent findings in a total of eight cases of samples. In addition, we have intentionally excluded any alterations with a span of <three consecutive probes on the genome from the data analysis to resolve two problems. The one problem arises from the FFPE DNA samples. Modified genomic DNA from the FFPE tissues can over-represent or under-represent the original copy number of genomic DNA on certain probes. The other problem arises from the probes on microarrays. The probes are well designed and their applicability has been validated by a wide range of researchers, although one or two misbehaving probes may show erroneous results. By ruling out the results obtained from one or two consecutive probes, we resolved both problems mentioned above. It is unavoidable, however, that this strategy increases false negatives. This might be partly because we found no recurrent loss of the copy number in the current study.
In the current study, we used HSIL and SCC tissues obtained from different patients. If we had compared the tissue samples of HSIL and SCC obtained from the same patient, we could have better shown the direct relationship between the two lesions. In the methods of the current study, we had required a significant amount of genomic DNA with a good quality. But we could not obtained a sufficient amount of genomic DNA from the same patient for an array CGH, the real-time PCR and other necessary procedures for quality control. We collected tissue samples of HSIL and SCC from the different patients, but tried to find recurrent changes in both groups. To our knowledge, an analysis of the recurrent alterations will reduce a bias due to the difference between the individual cases and it will also provide the information about the role of the alteration of copy number that can be generally accepted as the pathogenesis of cervical cancer.
To summarize, our results showed that the recurrent alteration of copy number occurred in cases of HSIL and those of SCC based on the genomic profiles of both disease entities. The alteration of copy number in AHNAK, TRIM29, and SDC1 genes and that in IVNS1ABP gene were found in premalignant lesions and invasive SCC, respectively.
Acknowledgments
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (2010-0028032).
Notes
No potential conflict of interest relevant to this article was reported.