Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 51(3); 2017 > Article
-
Review
Good Laboratory Standards for Clinical Next-Generation Sequencing Cancer Panel Tests - Jihun Kim,1,2,*, Woong-Yang Park3,*, Nayoung K. D. Kim3, Se Jin Jang1,2, Sung-Min Chun1,2, Chang-Ohk Sung1,2, Jene Choi1, Young-Hyeh Ko4, Yoon-La Choi4, Hyo Sup Shim5, Jae-Kyung Won6, The Molecular Pathology Study Group of Korean Society of Pathologists
-
Journal of Pathology and Translational Medicine 2017;51(3):191-204.
DOI: https://doi.org/10.4132/jptm.2017.03.14
Published online: May 10, 2017
1Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
2Center for Cancer Genome Discovery, Asan Institute for Life Sciences, Seoul, , Korea
3Samsung Genome Institute, Sungkyunkwan University School of Medicine, Seoul, Korea
4Department of Pathology, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea
5Department of Pathology, Yonsei University College of Medicine, Seoul, Korea
6Department of Pathology, Seoul National University College of Medicine, Seoul, Korea
- Corresponding Author Jihun Kim, MD, PhD Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, 88 Olympic-ro 43-gil, Songpa-gu, Seoul 05505, Korea Tel: +82-2-3010-4556 Fax: +82-2-472-7898 E-mail: 'jihunkim@amc.seoul.kr'
- *Jihun Kim and Woong-Yang Park contributed equally to this work as co-first authors.
© 2017 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted noncommercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
- Next-generation sequencing (NGS) has recently emerged as an essential component of personalized cancer medicine due to its high throughput and low per-base cost. However, no sufficient guidelines for implementing NGS as a clinical molecular pathology test are established in Korea. To ensure clinical grade quality without inhibiting adoption of NGS, a taskforce team assembled by the Korean Society of Pathologists developed laboratory guidelines for NGS cancer panel testing procedures and requirements for clinical implementation of NGS. This consensus standard proposal consists of two parts: laboratory guidelines and requirements for clinical NGS laboratories. The laboratory guidelines part addressed several important issues across multistep NGS cancer panel tests including choice of gene panel and platform, sample handling, nucleic acid management, sample identity tracking, library preparation, sequencing, analysis and reporting. Requirements for clinical NGS tests were summarized in terms of documentation, validation, quality management, and other required written policies. Together with appropriate pathologist training and international laboratory standards, these laboratory standards would help molecular pathology laboratories to successfully implement NGS cancer panel tests in clinic. In this way, the oncology community would be able to help patients to benefit more from personalized cancer medicine.
- Choice of genes to be tested and test platforms
- Traditionally, only a limited number of clinically validated predictive or prognostic biomarkers have been approved as clinical tests. However, accumulating genomic data obtained from multiple tumor types and expansion of genome-based clinical trials suggest that many of the future anti-cancer treatments and clinical trials will require comprehensive diagnostic panels that allow the detection of multiple mutations at the same time. Such tests might involve either probe-based capture or primer-based amplification for the enrichment of genomic regions to be tested.
- The number and scope of genes to be tested depend on the purpose of the test. If the purpose is limited to companion diagnostics for current standard care, the number of genes would be very limited. However, if there is a need for clinical trials for which NGS-based tests are required to stratify patients, a broader range of genes should be interrogated. In this context, the currently used cancer panel tests are mainly focused on clinically actionable genomic alterations at selected protein-coding regions that are defined by the availability of approved drugs and pathognomonic molecular features. However, there is an increasing need for adding genomic alterations associated with resistance to molecularly targeted therapies or predicted response to investigational drugs. Furthermore, genomic data could be used to refine pathological diagnosis. For example, mutations in IDH1 or IDH2 and co-deletion of chromosome 1p and 19q are already incorporated in recently revised pathological diagnosis of diffuse glioma [5].
- There are several NGS platforms such as Illumina, Ion Torrent, and Roche 454, and all of which have been reported to be as reliable as the current standard genotyping tools at least for the important cancer genes provided that laboratories perform adequate quality control [6-8]. Thus, pathology laboratories may choose their preferred platforms based on their individual requirements, such as expected sample status (i.e., small biopsies, resected tissue samples, or liquid biopsies, etc.), expected number of samples, and/or types of variants to be analyzed (e.g., copynumber analysis might be limited in platforms that use primer-based amplification during target enrichment). It is important for laboratories to be aware of platform characteristics and perform adequate quality controls depending on the platform characteristics.
- Specimen handling
- Adequate processing of tissue samples is essential in a reliable NGS cancer panel test. Required specimen handling procedures are nearly the same as those required for traditional single-gene tests. Briefly, the quality and the amount of neutral buffered formalin relative to the size of the specimen should be monitored. The time interval from specimen acquisition to fixation should be minimized, and optimal fixation duration should be monitored [9]. The optimal fixation duration depends on the dimension of each sample because formalin penetrates tissue at a rate of approximately 1 mm per hour. The rule of thumb for recommended fixation duration for surgically resected specimen is 24 hours [10].
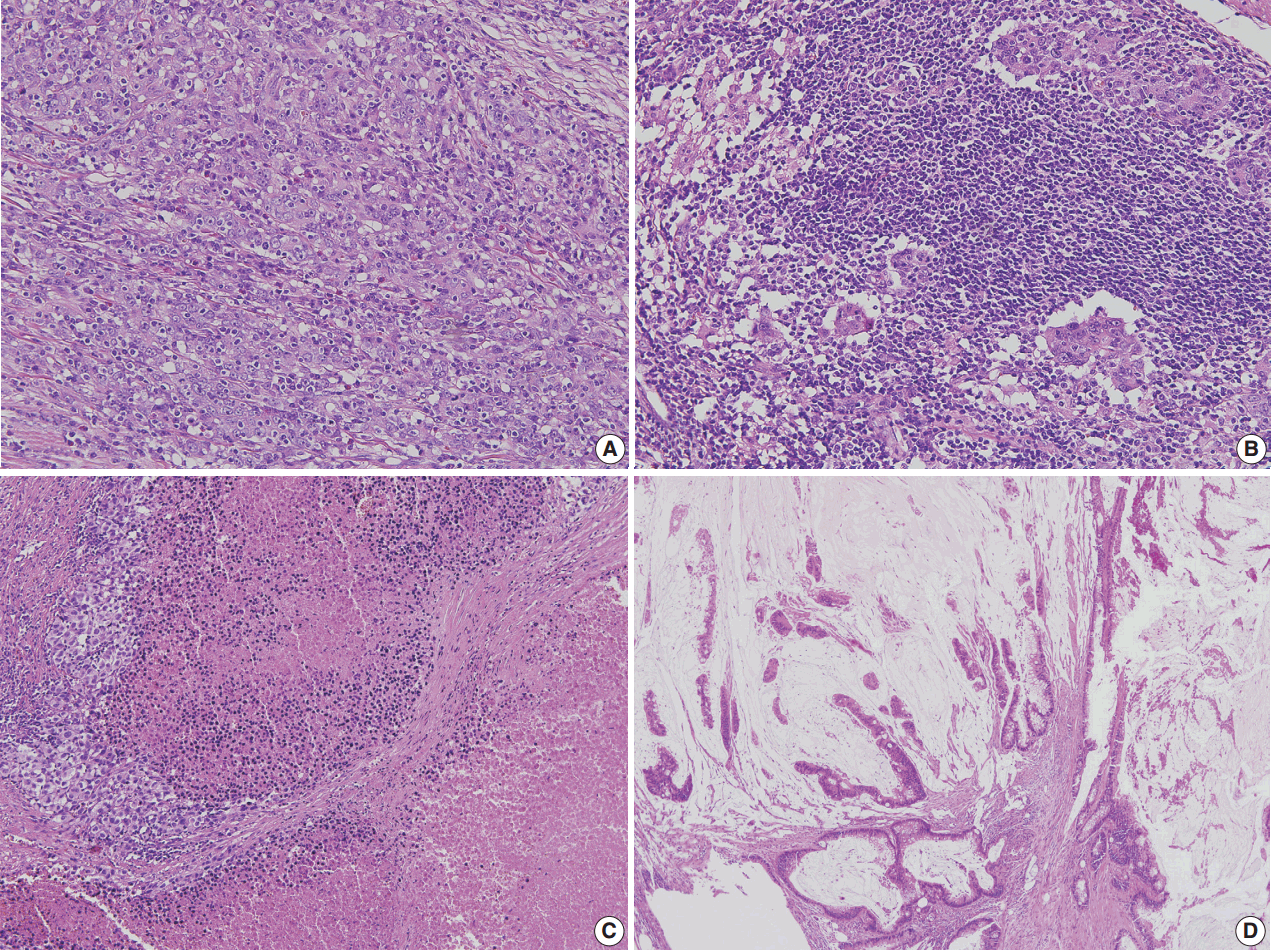
- Correct pathologic diagnosis and assessment of the fraction of neoplastic cell nuclei in tumor area by pathologists are crucial for the interpretation of NGS results. Once the diagnosis and tumor purity are established, it is strongly recommended that the pathologists circle an area from which nucleic acid is extracted so that neoplastic cells are enriched to the appropriate level of tumor purity. The minimum requirement of tumor purity can be established in the initial validation of test performances (see “Validation” section). The estimated tumor purity of a selected area should be documented. When tumor purity is assessed, the pathologists should be aware of the fact that the relative number of tumor cell nuclei reflects tumor purity rather than occupied tumor area. Thus, an area with heavy inflammatory cell infiltration should be avoided (Fig. 1A, B). Also, to ensure DNA quality, pathologists should try to avoid necrotic areas (Fig. 1C) and areas with extensive extracellular mucin (Fig. 1D).
- Nucleic acid extraction, quantification, and storage
- For clinical tests, DNA extraction kits should have a high level of performance specification to obtain DNA of sufficient quality and quantity for intended NGS tests from formalin-fixed paraffin embedded (FFPE) samples. In addition, DNA extraction procedures should have appropriate mechanisms by which sample contamination or misidentification could be avoided. Several commercially available kits that use silica-based or magnetic bead–based extraction protocols meet such requirements.
- A major cause of sequence artifacts is deamination of cytosine resulting in C to T transitions during amplification [11-14]. Formalin fixation and longer storage period contribute to this process [15]. Since those sequence artifacts are usually present at a very low frequency, such artifacts are unlikely to affect test results if sufficient amounts of unique DNA molecules are available. However, if the DNA input amount is too small or if the purpose is to detect variants with low allelic fraction, these artifacts would be a problem. In addition, amplicon-based methods are more susceptible to these artifacts than hybridization capture-based methods [11]. Thus, laboratories may choose DNA extraction protocols with enzymatic removal of uracil-containing templates when they interrogate FFPE samples with highly fragmented DNA or with low tumor purity by amplicon-based methods. In the case of decalcified specimens, DNA quality or quantity is inferior to those of the specimens without decalcification. Regarding this, it has been reported that a decalcification protocol involving ethylenediaminetetraacetic acid might be better than that involving hydrochloric acid in terms of usable nucleic acid for subsequent NGS analysis [16].
- Quantitation of extracted nucleic acids can be done by Nano-Drop, Qubit, or the Picogreen method. Among them, Nano-Drop is not recommended because it also detects nucleic acids that are not suitable for downstream analyses. Nucleic acids should be stored under highly controlled conditions in order to maintain sample identity and integrity. Extracted DNA is to be stored at –20˚C and RNA at –80˚C. Sequencing libraries and polymerase chain reaction (PCR) products may be stored in –20˚C but should be separated from pre-amplification materials to prevent them from contaminating pre-amplification materials.
- Sample identity tracking
- Like other single gene–based tests, verification of sample identity is the most basic and important aspect in clinical NGS test. The NGS cancer panel tests involve many steps, making them inherently subject to sample mix-up or swapping. Thus, test procedures should have an appropriate system to minimize such critical events. For example, panels could be designed to include a number of single nucleotide polymorphism (SNP) markers that allow molecular barcoding of patient samples [17] so that sample identity can be traced. In addition, an electronic laboratory information management system could be useful for this purpose. With any tool in place, it is important to enable end-to-end sample tracking in clinical NGS cancer panel tests.
- Library preparation
- Library preparation is the step where extracted nucleic acids are prepared for the sequencing reaction. It involves DNA fragmentation, adaptor ligation, and enrichment of the target of interest. Target enrichment can be done either by an amplicon-based approach or by hybridization-based capture. Molecular barcodes are usually introduced to enable sample identification and pooling of multiple samples in a single flow cell lane. To avoid contamination, all steps before amplification should be done in a separate space. In addition, extra-caution should be in place during sample transportation from pre-PCR area to post-PCR area. After library preparation, appropriate quality controls should be applied to determine whether the rest of the procedures should be continued. Quality controls include quantitation, fragment size analysis, and quantitative PCR using adaptor sequences for priming.
- Data analysis of sequence reads
- Bioinformatics pipelines used for the analysis of NGS data consists of multiple steps, such as de-multiplexing, read alignment, de-duplication, base calibration, variant calling, filtering, and annotation. Currently, no single “gold-standard” algorithm exists. Therefore, laboratories should choose the most suitable algorithm for the types of variants to be reported and optimize them. In the absence of a “gold standard,” it is important to validate the analytic performance of the bioinformatics pipelines [18-20]. It is also important to make sure that all versions of algorithms are traceable and properly updated (see “Validation” section).
- The initial step after sequencing includes converting the base intensities in a sequencer to digital-level nucleotide sequences, called FASTQ. Although the types of signals differ among various sequencing platforms, most have their internal software for translating base calling into the compressed and de-multiplexed FASTQ files. The nucleotides in FASTQ have corresponding base quality scores that are in a form of logarithmic scales indicating the probabilistic confidence level of the bases. Appropriate quality control should be done to confirm the general integrity of the sequencing data, such as the total number of bases, sequence contents (including GC contents), per base sequence quality, etc. Read trimming is recommended if the base quality or composition does not meet the quality control threshold that each laboratory has empirically set up or that the sequencer manufacturer has recommended.
- This step is to find where the short read sequences are located. In general, single or pair-end reads are first aligned (mapped) to a human genome reference. It is essential to include the version of the human reference sequence in the clinical report. Since poorly mapped sequences may lead to compromise the reliability of called variants, especially in solid tumors with low tumor content, sequencing reads should be filtered based on mapping quality score so that only confidently mapped reads are processed further. After the initial mapping, read duplicates should be removed, because unwanted clonal amplification of reads with sequence artifacts may lead to false-positive variant calls. Alignment is often challenging when the sequence reads come from genomic regions having large insertions or deletions, repetitive regions, pseudogenes, or homologous genes because there can be several other similar-looking genomic regions across the reference genome. Realignment using known references that have the suspected insertions or deletions may be necessary in such cases.
- Somatic variants can be identified by subtracting variants found in non-neoplastic cells from those found in cancer. If a laboratory chooses not to sequence the corresponding normal samples, it should be noted that some germline variations might exist in the result. Laboratories should consider the implementation of modular analysis pipelines, in which different algorithms or settings are used to call different types of variants: single nucleotide variations (SNVs), insertion/deletion (Indels), copy-number variations (CNVs), and translocations.
- The quality of variant calls is strongly related to the quality of alignment. The key challenge of variant calling is to distinguish real variants from sequencing errors. In general, the more times the variant is sequenced, the more reliable the variant call is. The minimum depth of coverage depends on the required sensitivity of the intended assay, the sequencing platform and the types of mutations to be detected. Although sensitivity is increased in proportion to sequencing depth, false-positive calls may also be increased especially in cases with low tumor content. There are various reasons for false-positive variant calls; they often result from PCR errors, sequencing errors, mis-mapped reads on repetitive sequences or homologous regions and so forth. Because each variant calling algorithm uses different strategies to filter out false calls, different algorithms sometimes generate discrepant results. Thus, laboratories should find optimal parameter settings during assay development and validation to minimize algorithm-dependent result variability.
- Reliable identification of Indels is particularly challenging because sequence reads containing this type of variants are not often accurately mapped. Thus, the sensitivity and specificity for this type of variants is often reduced. With regard to this, laboratories should identify clinically important Indels and validate assay performances to establish reasonable sensitivity and positive predictive value for the identification of those Indels. Due to the high rate of false-positives, manual review by using visualization tools and comparison with the same regions in other samples on the same run is recommended for all Indel calls.
- The reliable identification of CNV is quite difficult in NGS cancer panel sequencing because of the uneven target coverage related to hybridization capturing steps, the absence of matched normal data, or the lack of coverage uniformity [21]. Although algorithms for detecting CNV in targeted NGS tests are improving [22], the inherent limitations in cancer panel sequencing of clinical FFPE samples require robust validation of test performance. This type of validation can be done by testing characterized cancer cell lines or clinical samples with known CNV profiles (see “Validation” section) [23], although there is no agreement upon the minimal number of samples for appropriate validation.
- Translocation can be identified based on the DNA level, using discordant or split sequencing reads. However, the inherent limitations of short-reads in terms of alignment can result in many false positive calls. Therefore, the test performance should be appropriately validated using reference materials with known translocations across targeted genomic regions that have been confirmed by the current gold standard. Furthermore, it is highly recommended that all translocation calls be manually reviewed by using genome visualization tools such as the Integrative Genomics Viewer. Finally, it should be noted that a translocation can be missed if the breakpoint of fusion (may be somewhere in the introns not covered by the panel) is not included in targeted genomic regions, even though protein-coding regions of the translocation partners are included in the panel.
- Variant annotation determines if a sequence variant is real and provides predicted resulting amino acid changes. To identify true somatic variants, false variants should be properly filtered. Important sources of false variants include cytosine deamination, amplification errors, and sequencing errors. Cytosine deaminations are introduced ex vivo; these variants are not copied to the opposite strand, meaning that the artifacts are only present on one strand. To facilitate the detection of cytosine deamination artifacts, laboratories may use techniques such as molecular inversion probe and HaloPlex and Duplex sequencing, to enrich and sequence both the sense and antisense DNA strands [24]. Amplification errors can be introduced due to DNA polymerase errors during amplification steps of library enrichment. These errors might be minimized by the application of unique barcodes to individual DNA molecules during library enrichment. If the same variant is detected in multiple unique molecules, the variant might be real because it is unlikely that individual molecules acquire the same polymerase error during amplification. Sequencing errors are highly dependent on the sequencing platform and sequencing chemistry. The ability to call SNVs and Indels is known to be similarly accurate for data generated on the PGM and Illumina platform, provided that there is sufficient coverage [24].
- Depending on the types of variants (e.g., missense, nonsense, etc.) and the types of genes (e.g., hotspots in oncogenes vs randomly distributed mutations in tumor suppressor genes), interpretation of detected variants can be simple or quite complicated. Basically, the ACMG strongly recommends that interpretation is performed by trained staffs such as clinical molecular geneticists or molecular pathologists [4]. A multi-disciplinary sequencing data analysis team with various scientific backgrounds including clinical oncology, genomics, bioinformatics, and pathology, is recommended for accurate interpretation.
- For variants that are not hotspot mutations, germline variants should be excluded first. Since most cancer panel tests do not analyze matched non-neoplastic tissue, laboratories should prepare mechanisms to filter out potential germline variants based on the genetic polymorphism data on the population to which the tested patient belongs. In most instances, laboratories use public databases on germline polymorphism such as the 1000 Genomes Project, ExAC, or dbSNP (Table 1), but ideally, these data should be derived from the same ethnic group as the tested patient.
- Since the clinical and biological significance of cancer-related genomic variants are increasingly characterized, many variants detected in most cancer panel tests have related information in public databases (see “Reporting” section for details). However, previously uncharacterized variants may also be detected considering the characteristics of NGS tests. Potential biological significance, or pathogenicity, could be inferred from the archived genotype-phenotype correlation data such as ClinVar, Human Gene Mutation Database, and Leiden Open Variation Database (Table 1). In addition, the in silico prediction of functional impacts is available in dbNSFP or Ensembl Variant Effector (Table 1). Since no single database is perfect, it is essential to refer to multiple resources for appropriate interpretation. Finally, knowledge about variants will be continuously improved with the accumulation of each lab’s experiences and feedbacks from clinicians.
- Reporting
- Reporting of NGS cancer panel test results should follow the general professional organizations’ recommendations and guidelines [25]. There are two major essential parts of a report: proper patient identification and detected actionable variants. The patient identification part is the same as that in the current standard single gene-based tests, but the detected variant part is more complex and sophisticated in clinical NGS cancer panel tests because typical NGS cancer panel tests detect multiple variants at the same time. The clinical NGS report should include the essential information listed in Table 2, and the most pertinent information, such as actionable variants and a critical summary of those variants, should be placed in a clearly visible section on the first page. Detected sequence variants should be annotated in concordance with the Human Genome Variation Society mutation nomenclature and the version of the human reference sequence to which sequence reads were aligned should also be included in the report. Variants, which are not clinically actionable but are potentially useful for future practice, might follow the actionable variants. Then, detailed information about the detected variants (see the “Interpretation” section) and essential technical information, such as genes or genomic regions included in the panel and key quality control metrics, may be listed. Inclusion of granular details of technical information is not recommended, but a description of how clinicians can obtain the details may be included [26].
- Since typical cancer panel results include several cancer-related genomic variants with different levels of clinical or biological evidences, variants should be classified and reported according to the level of evidences. Several leading institutions use their own variant classification system and each institution may discuss with clinicians about which variant classification system should be used. However, a recently published related guideline is a good example [27]: (1) Tier 1, strong clinical significance such as Food and Drug Administration (FDA)–approved therapies, professional guidelines, prognostic and diagnostic markers based on well-powered studies with expert consensus; (2) Tier 2, potential clinical significance such as FDA-approved therapies for different tumor types, investigational therapies, prognostic and diagnostic markers based on small published studies with some consensus and preclinical trials or case reports without consensus; (3) Tier 3, unknown clinical significance; and (4) Tier 4, benign or likely benign. Tier 1 to 3 variants should be included in the clinical NGS report in a decreasing order of clinical significance and it is not recommended to include Tier 4 variants in clinical NGS report.
- Interpretation of detected variants in terms of their clinical impact and pathogenicity is a daunting task. As mentioned in the “Interpretation of computational output” section, many information sources such as public databases, published guidelines, and computational prediction algorithms should be integrated for proper interpretation. In addition to the previously mentioned public databases on germline polymorphism, in silico prediction of the functional impacts, genotype-phenotype relationship, and several cancer-specific mutation databases are available online (Table 1). These resources are very useful in the interpretation of variants in the context of cancer, but it is important to make sure that the database is properly curated, referenced and updated in a regular basis. Regarding this, FDA guidance suggested that appropriate databases should implement decision matrices with published details of each variant’s interpretation and have documented standard operating procedures (SOPs) for the curation and update of this information [28].
- When assigning variants to each tier, it is essential that they should be interpreted in the context of both variant details (e.g., BRAF V600E) and patient tumor type (e.g., malignant melanoma vs colorectal cancer). This is because different variants in a single gene may have different clinical impacts and the clinical significance of a particular variant may vary depending on the tumor type. For tier 1 variants, a clear statement on their clinical relevance and any appropriate action should be included. Also, it might be helpful to clarify the absence of other variants that are important in the patient tumor type. For tier 2 variants, advising the ordering physician about available clinical trials may be useful. In addition, tier information is subject to change as new evidences and therapeutic options are continuously emerging.
- In cases where only limited interpretation can be made and full quality control standards could not be met, it is essential that pathologists make a professional judgment on whether the result should be reported. Also, any limitation of the analysis should be clarified in the report. For example, negative results in samples with low neoplastic cell content at near or below the established sample rejection criteria should be accompanied by a note on potential false negatives. Furthermore, clinical NGS laboratories should have clearly defined protocols about when additional confirmatory tests should be advised, as well as performance validation data in cases where those confirmatory tests are not necessary.
LABORATORY GUIDELINES FOR NEXT-GENERATION SEQUENCING CANCER PANEL TESTS
Sample transportation, receipt, and storage
Morphological assessment and designation of ideal tumor area
Sequencing read
Alignment
Variant calling: single nucleotide variation, insertion/deletion, copy-number variation, and translocation
SNV/Indel
Copy-number variation
Translocation
Variant annotation and filtering
Interpretation of computational output
General format
Presentation of detected variants and clinical translation
- Wet Bench Analytic Process
- The detailed documentation of SOPs is critical for quality assurance of a complex, multi-step wet bench process. All SOPs of each step of the wet bench process must be documented so that each step can be traced. This includes documentation of all methods, reagents, instruments and controls (if applicable). Most of the documentations should be similar to those of current standard single gene testing, but those specific to NGS testing include detailed information regarding captured regions, such as genomic coordinates of captured probes and lists of genes and target enrichment protocols. Clinical laboratories that process different types of samples, such as FFPE samples or blood, should establish SOPs for each validated sample type. Metrics for quality control to assess run status must also be documented. Examples include mean target coverage, percentage of reads that map to target regions, and the fraction of bases meeting specified quality and coverage thresholds. Laboratories must define and document acceptance or rejection criteria for each step of the wet bench process, such as DNA extraction, library preparation, and sequencing.
- Before testing patient samples, clinical NGS laboratories must establish the analytical validity of the intended tests. If the intended test is approved by the Ministry of Food and Drug Safety (MFDS), laboratories can verify the performance specifications established by the manufacturer. If the intended test is not approved by the MFDS, i.e., a laboratory-developed test, laboratories must establish performance characteristics such as accuracy, precision, sensitivity, specificity, and reportable range.
- Because NGS cancer panel tests involve complex, multistep processes, each step needs to be empirically optimized to determine optimal assay conditions. Once those optimizations are done, an analytic validation should be performed for a whole test in a “beginning to end” fashion. Test performance should be separately validated for different types of samples, such as FFPE or blood.
- For “beginning to end” validation, a number of samples should be analyzed to assess the test performance. There is no general agreement on how many samples are required for this type of validation. With regard to this issue, the CAP concluded that adding a minimum number of samples for validation is premature given the ongoing evolution of NGS technologies and the diversity of diagnostic applications [3]. Literature review revealed that the sample number for validation varied with a range of 20–80 samples [29-32]. The minimum number of samples needed for an appropriate validation might be affected by the size of the test, i.e., a larger gene panel will require more extensive validation, and by the range of samples to be tested, i.e., FFPE samples with variable tumor content. Here, we suggest the general principles of validation in terms of several important analytic performance parameters.
- Considering the inherent characteristics of NGS cancer panel tests, it is important to evaluate as many different genomic regions as possible because sequence context can influence sequencing results. In addition, laboratories should determine analytic performances for all variant types relevant to the intended test (e.g., SNV, Indel, CNV, and translocations). Since NGS-based tests interrogate multiple variants at the same time, the validation of test performances involves two parts: method-specific (detection of as many variants as possible in a single sample) and analyte-specific (detection of a certain genomic variant in multiple samples). Laboratories can establish the test performances by combining those two approaches.
- Accuracy validation is a “method-specific” way of validation. For this type of validation, laboratories might use well-characterized cancer cell line samples to interrogate whether the intended test accurately detects all known variants within the genomic regions covered by the panel. In this case, reference cell line samples should be periodically monitored for identity and passage number to prevent a significant genetic drift. In addition, laboratories might use well-characterized HapMap samples (NA12878 and NA19240), variants of which are readily available on the web [33,34]. Details of available reference materials are summarized in Table 3. Evaluation of these reference samples might be quite useful to validate the accuracy across a wide range of genomic regions [23]. During this type of validation, an adequate depth of coverage threshold [35] necessary to accurately call all expected variants can be empirically established by plotting false-positives and false-negatives as a function of coverage.
- Analytic sensitivity can be evaluated by comparing NGS test results with the current gold standard test results for known genetic variants in reference materials. To this end, laboratories may use historical controls such as accumulated clinical samples with well-characterized genomic variants by gold standard methods, provided that the gold standard tests were done in an appropriately accredited environment.
- Analytic specificity is theoretically calculated by determining the fraction of test negatives (wild type sequence calls by the intended test) per true-negatives (samples that are known to have wild type sequences by the gold standard method). However, this concept often does not work well for NGS-based tests because too many potential variants are included in typical cancer panels. For most laboratories, it might be reasonable to leverage specificity by determining the average number of false-positive calls for the regions tested in a number of well-characterized clinical samples.
- The limit of detection can be determined by cell line “mixing” experiments, in which a cell line with known genetic variants is serially diluted with another cell line without such variants to have different mixture ratios, which are then tested to determine whether the intended test accurately detects all specified variants under a certain coverage threshold. Alternatively, laboratories may determine the limit of detection by using the HapMap samples (NA12878 and NA19240) mentioned earlier by mixing them in various ratios and interrogating variants that differ between the two different HapMap samples.
- Precision (repeatability for intra-run and reproducibility for inter-run variability) can be determined by repeating the same samples in the same run (repeatability) or by repeating the same sample in a different run (on a different day by a different operator). Ideally, testing at least three samples is recommended to adequately establish precision adequately [3].
- Any changes to a clinical NGS test, such as changes in instruments, specimen types, reagents, or software, require that performance specifications be re-established or be shown to be unchanged. For example, inclusion of new genes to an existing gene panel requires revalidation to make sure that new sequence variations are reliably detected without compromising the quality of the original assay. The extent of re-validation depends on the predicted influence on the test performance related to the change. For example, if only the bioinformatics pipeline is updated, it may not be necessary to re-validate all steps before data analysis.
- Clinical NGS laboratories should establish and follow a quality management plan. This plan should be integrated within the institution’s overall quality assurance program. Components of the NGS quality management program are not much different from the traditional single gene-based tests. Once laboratories establish an initial validation of test performances, laboratories must perform internal quality controls daily and external quality controls periodically. Each component of the quality assurance program specific to NGS cancer panel tests is discussed in detail below.
- Recently, guidelines for quality control and recommendations for the use of NGS in different applications have been published [3,4,25,36-42] and are summarized in Table 4. This guidance will work as a checklist for each component of the intended NGS test and related quality control metrics that require reviewing in order to feel confident about the quality of results. It is impractical to include multiple positive controls with different variant types during each run due to unacceptable cost and time. Instead, a single characterized external control with known variants in each run may be sufficient to demonstrate that the procedure is successful [36]. Typically, this can be done by preparing a number of DNA aliquots from a large FFPE sample block whose genome has already been well-characterized genomically and include those aliquots in each run of patient samples.
- The Clinical Laboratory Improvement Amendment (CLIA) requires participation in a proficiency testing (PT) program twice a year for most clinical tests. For any clinical tests without such PT requirements, laboratories must verify test performance twice a year [24]. Since PT programs specific to NGS cancer panel tests do not currently exist, laboratories may verify their test performance by using alternate assessment (AA), where deidentified patient samples are exchanged and re-tested between different laboratories until formal PT programs are set [24]. A Korean guideline published by the MFDS follows the same principle and requires documentation of all procedures related to PT or AA [43]. There is no general agreement upon the number of reference materials per one round of PT or AA but analysis of two samples has been recommended [36].
- As NGS cancer panel tests with appropriate quality controls are reported to meet a clinical grade performance [23,44-47], routine confirmatory testing is not recommended. However, laboratories should have a policy that clearly documents both any indication for confirmatory testing as well as any performance validation data upon which they decided that such confirmatory testing is not necessary. In addition, the CAP is flexible regarding the methods used for any needed confirmatory testing.
- Keeping comprehensive laboratory records is essential in monitoring complex, multi-step NGS cancer panel tests. In this regards, such records should be maintained in such a way that all detailed information about test procedures including reagents, sequencing runs, wet lab, and bioinformatics procedures and responsible technicians is traceable. While all details need not be included in the clinical report, laboratories should maintain a database from which detailed information regarding the analysis of individual specimen can be obtained.
- Laboratories must be prepared for upgrades to make sure that they are not using obsolete methods. A policy for the upgrade of instruments, sequencing chemistries, and reagents or kits, as well as subsequent post-upgrade validation of test performances should be in place. The policy may include specified intervals for upgrade and required validation processes, depending on the type or extent of upgrade.
- Bioinformatics process
- Laboratories should document all bioinformatics processes, including all data files, variant caller’s parameters, and versions of the bioinformatics algorithms. Sources and versions of all bioinformatics algorithms should be documented and updated properly. Quality control information on bioinformatics analysis, such as the cut-off of read depth, base quality score, and mapping quality, should also be documented.
- General validation principles were already discussed in the Validation proportion of the “Wet Bench Analytic Process” section. Briefly, for bioinformatics pipelines, laboratories should iteratively find parameters for optimal performance of computational algorithms before applying it to the lab process. Once the pipeline is initially validated, variations between sequencing runs should be monitored daily. Principles of the quality management were already discussed in the “Wet Bench Analytic Process” section. In short, laboratories should monitor any deviation from established performance characteristics in terms of quality metrics and analysis results. For any deviation, laboratories should document the investigational measures and corrective actions made to resolve the deviation. Essential quality metrics for bioinformatics performance verification include depth of coverage, uniformity of coverage and base call quality scores. In addition, GC bias, proportion of reads that map to nontargeted regions, and percentage of duplicated reads could also be used to monitor performances of sequencing reaction and subsequent bioinformatics analyses (Table 5).
- The bioinformatics pipeline should be revalidated upon any changes in operating systems, software, or overall pipelines, which may otherwise affect its analytic performance. Since a huge amount of data files are generated from the bioinformatics pipeline, it is impractical to store all sorts of files considering the significant cost. Instead, the CAP NGS workgroup recommended that some important file formats, such as FASTQ, BAM, and VCF, should be stored for quality controls or investigational use [3]. There is no general agreement on the required storage period, but it is important for laboratories to set their own storage policies in accordance with local or national requirements (if any) and inform clients of those policies.
REQUIREMENTS FOR CLINICAL NEXT-GENERATION SEQUENCING LABORATORIES
Documentation
Validation
Quality management
Internal quality assurance program
External quality assurance program: proficiency testin and alternate assessment
Policies: confirmatory testing, laboratory records, upgrades
Documentation
Validation and quality management
Policies: upgrades, storage, and data management
- Although the advancement of NGS technologies unraveled the genomic landscape of human malignancies and opened an era of genome-guided anti-cancer therapies, the complexity of NGS technologies made it especially difficult to set up regulatory or professional standards for assuring analytical validity of test results. In this report, the workgroup addressed general guidelines related to the test procedures and proposed requirements for clinical NGS cancer panel tests such as test validation, quality controls, reference materials and participation in PT or AA. Due to space limitations and the clinical implementation of NGS cancer panel tests being in its infancy, granular details were not covered in this review and those details should be addressed in further publications.
- As clinical experiences accumulate and technologies evolve, more refined recommendations or guidelines could be published in the near future. In addition, good clinical decision support systems or knowledge bases need to be developed to help patients in terms of benefits from personalized FDA-approved or investigational therapies and prognostic information. To realize personalized cancer medicine in the end, it is important for medical communities to share information on genotype (oncogenic variants in patients’ samples)–phenotype (drug response or prognosis) relationships, and all these efforts start with accurate genomic profiling of cancer tissue. To ensure reliability of NGS-based genomic profiling, a multi-disciplinary team approach, involving clinicians, pathologists, laboratory scientists, bioinformaticians, manufacturers, professional organizations, and government agencies, is essential.
CONCLUSION
-
Conflicts of Interest
No potential conflict of interest relevant to this article was reported.
Notes
Acknowledgments
| Subject | Database/Algorithm | URL |
|---|---|---|
| Germline polymorphism | dbSNP, dbVar | https://www.ncbi.nlm.nih.gov/projects/SNP/ |
| 1000 Genomes Project | http://browser.1000genomes.org/ | |
| ExAC | http://exac.broadinstitute.org | |
| Cancer-specific somatic variants | COSMIC | http://cancer.sanger.ac.uk/cosmic |
| cBioPortal | http://www.cbioportal.org/ | |
| My Cancer Genome | https://www.mycancergenome.org/ | |
| CIViC | https://civic.genome.wustl.edu/#/home | |
| Personalized Cancer Therapy, MD Anderson Cancer Center | https://pct.mdanderson.org/#/ | |
| Genotype-phenotype association, not limited to cancer | ClinVar | https://www.ncbi.nlm.nih.gov/clinvar/ |
| Human Gene Mutation Database | http://www.hgmd.cf.ac.uk/ac/index.php | |
| Leiden Open Variation Database | http://www.lovd.nl/3.0/home | |
| In silico functional prediction | dbNSFP (pre-computed in silico functional prediction and annotation of non-synonymous SNVs) | http://sites.google.com/site/jpopgen/dbNSFP |
| Ensembl Variant Effector Predictor | http://www.ensembl.org/info/docs/tools/vep/index.html |
This list is not comprehensive and only provides some examples. All websites were last accessed on January 3, 2017.
ExAC, Exome Aggregation Consortium; COSMIC, Catalog of Somatic Mutations in Cancer; CIViC, Clinical Interpretations of Variants in Cancer; NSFP, nonsynonymous functional prediction; SNV, single nucleotide variation.
Adapted by permission from Macmillan Publishers Ltd: [Nature Biotechnology] Gargis et al. 2012;30:1033-6, [36] copyright (2012).
NGS, next-generation sequencing.
Adapted by permission from Macmillan Publishers Ltd: [Nature Biotechnology] Gargis et al. 2012;30:1033-6, [36] copyright (2012).
NGS, next-generation sequencing.
- 1. Bentley DR, Balasubramanian S, Swerdlow HP, et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 2008; 456: 53–9. ArticlePubMedPMC
- 2. Metzker ML. Sequencing technologies: the next generation. Nat Rev Genet 2010; 11: 31–46. ArticlePubMed
- 3. Aziz N, Zhao Q, Bry L, et al. College of American Pathologists' laboratory standards for next-generation sequencing clinical tests. Arch Pathol Lab Med 2015; 139: 481–93. ArticlePubMed
- 4. Rehm HL, Bale SJ, Bayrak-Toydemir P, et al. ACMG clinical laboratory standards for next-generation sequencing. Genet Med 2013; 15: 733–47. ArticlePubMedPMC
- 5. Louis DN, Ohgaki H, Wiestler OD, Cavenee WK. WHO classification of tumours of the central nervous system. Revised 4th ed. Lyon: IARC Press, 2016.
- 6. Burghel GJ, Hurst CD, Watson CM, et al. Towards a next-generation sequencing diagnostic service for tumour genotyping: a comparison of panels and platforms. Biomed Res Int 2015; 2015: 478017.ArticlePubMedPMCPDF
- 7. Hinrichs JW, van Blokland WT, Moons MJ, et al. Comparison of next-generation sequencing and mutation-specific platforms in clinical practice. Am J Clin Pathol 2015; 143: 573–8. ArticlePubMed
- 8. McCourt CM, McArt DG, Mills K, et al. Validation of next generation sequencing technologies in comparison to current diagnostic gold standards for BRAF, EGFR and KRAS mutational analysis. PLoS One 2013; 8: e69604. Article
- 9. Hicks DG, Boyce BF. The challenge and importance of standardizing pre-analytical variables in surgical pathology specimens for clinical care and translational research. Biotech Histochem 2012; 87: 14–7. ArticlePubMed
- 10. Howat WJ, Wilson BA. Tissue fixation and the effect of molecular fixatives on downstream staining procedures. Methods 2014; 70: 12–9. ArticlePubMedPMC
- 11. Do H, Dobrovic A. Sequence artifacts in DNA from formalin-fixed tissues: causes and strategies for minimization. Clin Chem 2015; 61: 64–71. ArticlePubMed
- 12. Do H, Wong SQ, Li J, Dobrovic A. Reducing sequence artifacts in amplicon-based massively parallel sequencing of formalin-fixed paraffin-embedded DNA by enzymatic depletion of uracil-containing templates. Clin Chem 2013; 59: 1376–83. ArticlePubMed
- 13. Lin MT, Mosier SL, Thiess M, et al. Clinical validation of KRAS, BRAF, and EGFR mutation detection using next-generation sequencing. Am J Clin Pathol 2014; 141: 856–66. ArticlePubMedPMC
- 14. Serizawa M, Yokota T, Hosokawa A, et al. The efficacy of uracil DNA glycosylase pretreatment in amplicon-based massively parallel sequencing with DNA extracted from archived formalin-fixed paraffin-embedded esophageal cancer tissues. Cancer Genet 2015; 208: 415–27. ArticlePubMed
- 15. Marchetti A, Felicioni L, Buttitta F. Assessing EGFR mutations. N Engl J Med 2006; 354: 526–8. ArticlePubMed
- 16. Choi SE, Hong SW, Yoon SO. Proposal of an appropriate decalcification method of bone marrow biopsy specimens in the era of expanding genetic molecular study. J Pathol Transl Med 2015; 49: 236–42. ArticlePubMedPMCPDF
- 17. Pengelly RJ, Gibson J, Andreoletti G, Collins A, Mattocks CJ, Ennis S. A SNP profiling panel for sample tracking in whole-exome sequencing studies. Genome Med 2013; 5: 89.ArticlePubMedPMC
- 18. Brownstein CA, Beggs AH, Homer N, et al. An international effort towards developing standards for best practices in analysis, interpretation and reporting of clinical genome sequencing results in the CLARITY Challenge. Genome Biol 2014; 15: R53.ArticlePubMedPMC
- 19. Cornish A, Guda C. A comparison of variant calling pipelines using genome in a bottle as a reference. Biomed Res Int 2015; 2015: 456479.ArticlePubMedPMCPDF
- 20. Hwang S, Kim E, Lee I, Marcotte EM. Systematic comparison of variant calling pipelines using gold standard personal exome variants. Sci Rep 2015; 5: 17875.ArticlePubMedPMCPDF
- 21. Teo SM, Pawitan Y, Ku CS, Chia KS, Salim A. Statistical challenges associated with detecting copy number variations with next-generation sequencing. Bioinformatics 2012; 28: 2711–8. ArticlePubMed
- 22. Onsongo G, Baughn LB, Bower M, et al. CNV-RF is a random forest-based copy number variation detection method using next-generation sequencing. J Mol Diagn 2016; 18: 872–81. ArticlePubMed
- 23. Frampton GM, Fichtenholtz A, Otto GA, et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol 2013; 31: 1023–31. ArticlePubMedPMC
- 24. Deans ZC, Costa JL, Cree I, et al. Integration of next-generation sequencing in clinical diagnostic molecular pathology laboratories for analysis of solid tumours; an expert opinion on behalf of IQN Path ASBL. Virchows Arch 2017; 470: 5–20. ArticlePubMedPDF
- 25. ISO 15189: 2012 Medical laboratories: requirements for quality and competence [Internet]. Geneva: International Organization for Standardization, c2012-2017 [cited 2017 Jan 31]. Available from: https://www.iso.org/obp/ui/#iso:std:iso:15189:ed-3:v2:en.
- 26. Nomenclature for the description of sequence variants [Internet]. Melbourne: Human Genome Variation Society, c2010-2017 [cited 2017 Jan 31]. Available from: http://www.hgvs.org/mutnomen/.
- 27. Li MM, Datto M, Duncavage EJ, et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: a joint consensus recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J Mol Diagn 2017; 19: 4–23. ArticlePubMedPMC
- 28. Use of standards in FDA regulatory oversight of next generation sequencing (NGS)-based in vitro diagnostics (IVDs) used for diagnosing germline diseases [Internet]. Rockville: US Department of Health and Services, Food and Drug Administration, c2016-2017 [cited 2017 Jan 31]. Available from: http://www.fda.gov/downloads/MedicalDevices/DeviceRegulationandGuidance/GuidanceDocuments/UCM509838.pdf.
- 29. Chin EL, da Silva C, Hegde M. Assessment of clinical analytical sensitivity and specificity of next-generation sequencing for detection of simple and complex mutations. BMC Genet 2013; 14: 6.ArticlePubMedPMC
- 30. Cottrell CE, Al-Kateb H, Bredemeyer AJ, et al. Validation of a next-generation sequencing assay for clinical molecular oncology. J Mol Diagn 2014; 16: 89–105. ArticlePubMedPMC
- 31. Hagemann IS, Cottrell CE, Lockwood CM. Design of targeted, capture-based, next generation sequencing tests for precision cancer therapy. Cancer Genet 2013; 206: 420–31. ArticlePubMed
- 32. Williams ES, Hegde M. Implementing genomic medicine in pathology. Adv Anat Pathol 2013; 20: 238–44. ArticlePubMed
- 33. Zook JM, Chapman B, Wang J, et al. Integrating human sequence data sets provides a resource of benchmark SNP and indel genotype calls. Nat Biotechnol 2014; 32: 246–51. ArticlePubMed
- 34. National Center for Biotechnology Information. GeT-RM Homo sapiens [Internet]. Bethesda: US National Library of Medicine, c2017 [cited 2017 Jan 31]. Available from: https://www.ncbi.nlm.nih.gov/variation/tools/get-rm/.
- 35. Ajay SS, Parker SC, Abaan HO, Fajardo KV, Margulies EH. Accurate and comprehensive sequencing of personal genomes. Genome Res 2011; 21: 1498–505. ArticlePubMedPMC
- 36. Gargis AS, Kalman L, Berry MW, et al. Assuring the quality of next-generation sequencing in clinical laboratory practice. Nat Biotechnol 2012; 30: 1033–6. ArticlePubMed
- 37. Gargis AS, Kalman L, Bick DP, et al. Good laboratory practice for clinical next-generation sequencing informatics pipelines. Nat Biotechnol 2015; 33: 689–93. ArticlePubMedPMC
- 38. Association for Molecular Pathology. Clinical practice guidelines [Internet]. Bethesda: Association for Molecular Pathology, c2009-2017 [cited 2017 Jan 31]. Available from: https://www.amp.org/committees/clinical_practice/AMPclinicalpracticeguidelines.cfm.
- 39. ESMO clinical practice guidelines [Internet]. Viganello-Lugano: European Society for Medical Oncology, c2017 [cited 2017 Jan 31]. Available from: http://www.esmo.org/Guidelines.
- 40. Cree IA, Deans Z, Ligtenberg MJ, et al. Guidance for laboratories performing molecular pathology for cancer patients. J Clin Pathol 2014; 67: 923–31. ArticlePubMedPMC
- 41. Luthra R, Chen H, Roy-Chowdhuri S, Singh RR. Next-generation sequencing in clinical molecular diagnostics of cancer: advantages and challenges. Cancers (Basel) 2015; 7: 2023–36. ArticlePubMedPMC
- 42. Schrijver I, Aziz N, Farkas DH, et al. Opportunities and challenges associated with clinical diagnostic genome sequencing: a report of the Association for Molecular Pathology. J Mol Diagn 2012; 14: 525–40. ArticlePubMed
- 43. Next generation sequence analysis (NGS). Clinical laboratory certification guidelines [Internet]. Cheongju: Ministry of Food and Drug Safety, c2016 [cited 2017 Jan 31]. Available from: http://www.mfds.go.kr/index.do?mid=1161&seq=11001&cmd=v.
- 44. Cheng DT, Mitchell TN, Zehir A, et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J Mol Diagn 2015; 17: 251–64. ArticlePubMedPMC
- 45. Fisher KE, Zhang L, Wang J, et al. Clinical validation and implementation of a targeted next-generation sequencing assay to detect somatic variants in non-small cell lung, melanoma, and gastrointestinal malignancies. J Mol Diagn 2016; 18: 299–315. ArticlePubMedPMC
- 46. Pritchard CC, Salipante SJ, Koehler K, et al. Validation and implementation of targeted capture and sequencing for the detection of actionable mutation, copy number variation, and gene rearrangement in clinical cancer specimens. J Mol Diagn 2014; 16: 56–67. ArticlePubMedPMC
- 47. Sikkema-Raddatz B, Johansson LF, de Boer EN, et al. Targeted next-generation sequencing can replace Sanger sequencing in clinical diagnostics. Hum Mutat 2013; 34: 1035–42. ArticlePubMed
References
Figure & Data
References
Citations

- Validation and Clinical Application of ONCOaccuPanel for Targeted Next-Generation Sequencing of Solid Tumors
Moonsik Kim, Changseon Lee, Juyeon Hong, Juhee Kim, Ji Yun Jeong, Nora Jee-Young Park, Ji-Eun Kim, Ji Young Park
Cancer Research and Treatment.2023; 55(2): 429. CrossRef - Establishing molecular pathology curriculum for pathology trainees and continued medical education: a collaborative work from the Molecular Pathology Study Group of the Korean Society of Pathologists
Jiwon Koh, Ha Young Park, Jeong Mo Bae, Jun Kang, Uiju Cho, Seung Eun Lee, Haeyoun Kang, Min Eui Hong, Jae Kyung Won, Youn-La Choi, Wan-Seop Kim, Ahwon Lee
Journal of Pathology and Translational Medicine.2023; 57(5): 265. CrossRef - Clinical applications of next-generation sequencing in the diagnosis of genetic disorders in Korea: a narrative review
Jihoon G. Yoon, Man Jin Kim, Yong Jin Kwon, Jong-Hee Chae
Journal of the Korean Medical Association.2023; 66(10): 613. CrossRef - Obtaining spatially resolved tumor purity maps using deep multiple instance learning in a pan-cancer study
Mustafa Umit Oner, Jianbin Chen, Egor Revkov, Anne James, Seow Ye Heng, Arife Neslihan Kaya, Jacob Josiah Santiago Alvarez, Angela Takano, Xin Min Cheng, Tony Kiat Hon Lim, Daniel Shao Weng Tan, Weiwei Zhai, Anders Jacobsen Skanderup, Wing-Kin Sung, Hwee
Patterns.2022; 3(2): 100399. CrossRef - Update on Molecular Diagnosis in Extranodal NK/T-Cell Lymphoma and Its Role in the Era of Personalized Medicine
Ka-Hei (Murphy) Sun, Yin-Ting (Heylie) Wong, Ka-Man (Carmen) Cheung, Carmen (Michelle) Yuen, Yun-Tat (Ted) Chan, Wing-Yan (Jennifer) Lai, Chun (David) Chao, Wing-Sum (Katie) Fan, Yuen-Kiu (Karen) Chow, Man-Fai Law, Ho-Chi (Tommy) Tam
Diagnostics.2022; 12(2): 409. CrossRef - Defining Novel DNA Virus-Tumor Associations and Genomic Correlates Using Prospective Clinical Tumor/Normal Matched Sequencing Data
Chad M. Vanderbilt, Anita S. Bowman, Sumit Middha, Kseniya Petrova-Drus, Yi-Wei Tang, Xin Chen, Youxiang Wang, Jason Chang, Natasha Rekhtman, Klaus J. Busam, Sounak Gupta, Meera Hameed, Maria E. Arcila, Marc Ladanyi, Michael F. Berger, Snjezana Dogan, Ahm
The Journal of Molecular Diagnostics.2022; 24(5): 515. CrossRef - Performance Evaluation of Three DNA Sample Tracking Tools in a Whole Exome Sequencing Workflow
Gertjan Wils, Céline Helsmoortel, Pieter-Jan Volders, Inge Vereecke, Mauro Milazzo, Jo Vandesompele, Frauke Coppieters, Kim De Leeneer, Steve Lefever
Molecular Diagnosis & Therapy.2022; 26(4): 411. CrossRef - Clinical Quality Considerations when Using Next-Generation Sequencing (NGS) in Clinical Drug Development
Timothé Ménard, Alaina Barros, Christopher Ganter
Therapeutic Innovation & Regulatory Science.2021; 55(5): 1066. CrossRef - Fast Healthcare Interoperability Resources (FHIR)–Based Quality Information Exchange for Clinical Next-Generation Sequencing Genomic Testing: Implementation Study
Donghyeong Seong, Sungwon Jung, Sungchul Bae, Jongsuk Chung, Dae-Soon Son, Byoung-Kee Yi
Journal of Medical Internet Research.2021; 23(4): e26261. CrossRef - Status of Next-Generation Sequencing-Based Genetic Diagnosis in Hematologic Malignancies in Korea (2017-2018)
JinJu Kim, Ja Young Lee, Jungwon Huh, Myung-Hyun Nam, Myungshin Kim, Young-Uk Cho, Sun-Young Kong, Seung-Tae Lee, In-Suk Kim
Laboratory Medicine Online.2021; 11(1): 25. CrossRef - MSI-Testung
Josef Rüschoff, Gustavo Baretton, Hendrik Bläker, Wolfgang Dietmaier, Manfred Dietel, Arndt Hartmann, Lars-Christian Horn, Korinna Jöhrens, Thomas Kirchner, Ruth Knüchel, Doris Mayr, Sabine Merkelbach-Bruse, Hans-Ulrich Schildhaus, Peter Schirmacher, Mark
Der Pathologe.2021; 42(4): 414. CrossRef - Molecular biomarker testing for non–small cell lung cancer: consensus statement of the Korean Cardiopulmonary Pathology Study Group
Sunhee Chang, Hyo Sup Shim, Tae Jung Kim, Yoon-La Choi, Wan Seop Kim, Dong Hoon Shin, Lucia Kim, Heae Surng Park, Geon Kook Lee, Chang Hun Lee
Journal of Pathology and Translational Medicine.2021; 55(3): 181. CrossRef - MSI testing
Josef Rüschoff, Gustavo Baretton, Hendrik Bläker, Wolfgang Dietmaier, Manfred Dietel, Arndt Hartmann, Lars-Christian Horn, Korinna Jöhrens, Thomas Kirchner, Ruth Knüchel, Doris Mayr, Sabine Merkelbach-Bruse, Hans-Ulrich Schildhaus, Peter Schirmacher, Mark
Der Pathologe.2021; 42(S1): 110. CrossRef - 16S rDNA microbiome composition pattern analysis as a diagnostic biomarker for biliary tract cancer
Huisong Lee, Hyeon Kook Lee, Seog Ki Min, Won Hee Lee
World Journal of Surgical Oncology.2020;[Epub] CrossRef - Risk Stratification Using a Novel Genetic Classifier IncludingPLEKHS1Promoter Mutations for Differentiated Thyroid Cancer with Distant Metastasis
Chan Kwon Jung, Seung-Hyun Jung, Sora Jeon, Young Mun Jeong, Yourha Kim, Sohee Lee, Ja-Seong Bae, Yeun-Jun Chung
Thyroid.2020; 30(11): 1589. CrossRef - Biomarker testing for advanced lung cancer by next-generation sequencing; a valid method to achieve a comprehensive glimpse at mutational landscape
Anurag Mehta, Smreti Vasudevan, Sanjeev Kumar Sharma, Manoj Panigrahi, Moushumi Suryavanshi, Mumtaz Saifi, Ullas Batra
Applied Cancer Research.2020;[Epub] CrossRef - Application Areas of Traditional Molecular Genetic Methods and NGS in relation to Hereditary Urological Cancer Diagnosis
Dmitry S. Mikhaylenko, Alexander S. Tanas, Dmitry V. Zaletaev, Marina V. Nemtsova
Journal of Oncology.2020; 2020: 1. CrossRef - Assembling and Validating Bioinformatic Pipelines for Next-Generation Sequencing Clinical Assays
Jeffrey A SoRelle, Megan Wachsmann, Brandi L. Cantarel
Archives of Pathology & Laboratory Medicine.2020; 144(9): 1118. CrossRef - Standard operating procedure for somatic variant refinement of sequencing data with paired tumor and normal samples
Erica K. Barnell, Peter Ronning, Katie M. Campbell, Kilannin Krysiak, Benjamin J. Ainscough, Lana M. Sheta, Shahil P. Pema, Alina D. Schmidt, Megan Richters, Kelsy C. Cotto, Arpad M. Danos, Cody Ramirez, Zachary L. Skidmore, Nicholas C. Spies, Jasreet Hun
Genetics in Medicine.2019; 21(4): 972. CrossRef - A DNA pool of FLT3-ITD positive DNA samples can be used efficiently for analytical evaluation of NGS-based FLT3-ITD quantitation - Testing several different ITD sequences and rates, simultaneously
Zoltán A. Mezei, Dávid Tornai, Róza Földesi, László Madar, Andrea Sümegi, Mária Papp, Péter Antal-Szalmás
Journal of Biotechnology.2019; 303: 25. CrossRef - Pharmacogenomic Testing: Clinical Evidence and Implementation Challenges
Hippman, Nislow
Journal of Personalized Medicine.2019; 9(3): 40. CrossRef - Cancer Panel Assay for Precision Oncology Clinic: Results from a 1-Year Study
Dohee Kwon, Binnari Kim, Hyeong Chan Shin, Eun Ji Kim, Sang Yun Ha, Kee-Taek Jang, Seung Tae Kim, Jeeyun Lee, Won Ki Kang, Joon Oh Park, Kyoung-Mee Kim
Translational Oncology.2019; 12(11): 1488. CrossRef - Analytical Evaluation of an NGS Testing Method for Routine Molecular Diagnostics on Melanoma Formalin-Fixed, Paraffin-Embedded Tumor-Derived DNA
Mancini, Simi, Salvianti, Castiglione, Sonnati, Pinzani
Diagnostics.2019; 9(3): 117. CrossRef - Benchmark Database for Process Optimization and Quality Control of Clinical Cancer Panel Sequencing
Donghyeong Seong, Jongsuk Chung, Ki-Wook Lee, Sook-Young Kim, Byung-Suk Kim, Jung-Keun Song, Sungwon Jung, Taeseob Lee, Donghyun Park, Byoung-Kee Yi, Woong-Yang Park, Dae-Soon Son
Biotechnology and Bioprocess Engineering.2019; 24(5): 793. CrossRef - Use of the Ion PGM and the GeneReader NGS Systems in Daily Routine Practice for Advanced Lung Adenocarcinoma Patients: A Practical Point of View Reporting a Comparative Study and Assessment of 90 Patients
Simon Heeke, Véronique Hofman, Elodie Long-Mira, Virginie Lespinet, Salomé Lalvée, Olivier Bordone, Camille Ribeyre, Virginie Tanga, Jonathan Benzaquen, Sylvie Leroy, Charlotte Cohen, Jérôme Mouroux, Charles Marquette, Marius Ilié, Paul Hofman
Cancers.2018; 10(4): 88. CrossRef - Use of the Ion AmpliSeq Cancer Hotspot Panel in clinical molecular pathology laboratories for analysis of solid tumours: With emphasis on validation with relevant single molecular pathology tests and the Oncomine Focus Assay
Ahwon Lee, Sung-Hak Lee, Chan Kwon Jung, Gyungsin Park, Kyo Young Lee, Hyun Joo Choi, Ki Ouk Min, Tae Jung Kim, Eun Jung Lee, Youn Soo Lee
Pathology - Research and Practice.2018; 214(5): 713. CrossRef - Recent Advancement of the Molecular Diagnosis in Pediatric Brain Tumor
Jeong-Mo Bae, Jae-Kyung Won, Sung-Hye Park
Journal of Korean Neurosurgical Society.2018; 61(3): 376. CrossRef - The long tail of molecular alterations in non-small cell lung cancer: a single-institution experience of next-generation sequencing in clinical molecular diagnostics
Caterina Fumagalli, Davide Vacirca, Alessandra Rappa, Antonio Passaro, Juliana Guarize, Paola Rafaniello Raviele, Filippo de Marinis, Lorenzo Spaggiari, Chiara Casadio, Giuseppe Viale, Massimo Barberis, Elena Guerini-Rocco
Journal of Clinical Pathology.2018; 71(9): 767. CrossRef - Clinical laboratory utilization management and improved healthcare performance
Christopher Naugler, Deirdre L. Church
Critical Reviews in Clinical Laboratory Sciences.2018; 55(8): 535. CrossRef - Development of HLA-A, -B and -DR Typing Method Using Next-Generation Sequencing
Dong Hee Seo, Jeong Min Lee, Mi Ok Park, Hyun Ju Lee, Seo Yoon Moon, Mijin Oh, So Young Kim, Sang-Heon Lee, Ki-Eun Hyeong, Hae-Jin Hu, Dae-Yeon Cho
The Korean Journal of Blood Transfusion.2018; 29(3): 310. CrossRef - Value-based genomics
Jun Gong, Kathy Pan, Marwan Fakih, Sumanta Pal, Ravi Salgia
Oncotarget.2018; 9(21): 15792. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure

 E-submission
E-submission