Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 48(2); 2014 > Article
-
Original Article
KRAS Mutation Detection in Non-small Cell Lung Cancer Using a Peptide Nucleic Acid-Mediated Polymerase Chain Reaction Clamping Method and Comparative Validation with Next-Generation Sequencing - Boram Lee,*, Boin Lee,*, Gangmin Han, Mi Jung Kwon1, Joungho Han, Yoon-La Choi
-
Korean Journal of Pathology 2014;48(2):100-107.
DOI: https://doi.org/10.4132/KoreanJPathol.2014.48.2.100
Published online: April 28, 2014
Department of Pathology, Samsung Medical Center, Sungkyunkwan University College of Medicine, Seoul, Korea.
1Department of Pathology, Hallym University Sacred Heart Hospital, Hallym University College of Medicine, Anyang, Korea.
- Corresponding Author: Yoon-La Choi, M.D. Department of Pathology, Samsung Medical Center, Sungkyunkwan University School of Medicine, 81 Irwon-ro, Gangnam-gu, Seoul 135-710, Korea. Tel: +82-2-3410-2797, Fax: +82-2-3410-6396, ylachoi@skku.edu
- *Boram Lee and Boin Lee contributed equally to this work.
© 2014 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- KRAS is one of commonly mutated genetic "drivers" in non-small cell lung cancers (NSCLCs). Recent studies indicate that patients with KRAS-mutated tumors do not benefit from adjuvant chemotherapy, so there is now a focus on targeting KRAS-mutated NSCLCs. A feasible mutation detection method is required in order to accurately test for KRAS status.
-
Methods
- We compared direct Sanger sequencing and the peptide nucleic acid (PNA)-mediated polymerase chain reaction (PCR) clamping method in 134 NSCLCs and explored associations with clinicopathological factors. Next-generation sequencing (NGS) was used to validate the results of discordant cases. To increase the resolution of low-level somatic mutant molecules, PNA-mediated PCR clamping was used for mutant enrichment prior to NGS.
-
Results
- Twenty-one (15.7%) cases were found to have the KRAS mutations using direct sequencing, with two additional cases by the PNA-mediated PCR clamping method. The frequencies of KRAS mutant alleles were 2% and 4%, respectively, using conventional NGS, increasing up to 90% and 89%, using mutant-enriched NGS. The KRAS mutation occurs more frequently in the tumors of smokers (p=.012) and in stage IV tumors (p=.032).
-
Conclusions
- Direct sequencing can accurately detect mutations, but, it is not always possible to obtain a tumor sample with sufficient volume. The PNA-mediated PCR clamping can rapidly provide results with sufficient sensitivity.
- Patients
- One hundred and thirty-four cases of primary NSCLC were obtained from pathology files spanning from 2012 to 2013, including seventy-six biopsy samples of lung or lymph nodes, fifty-seven samples from surgical resection of lung or lymph nodes, and one pleural fluid sample. Clinical information including cancer stage and smoking history were obtained based on clinical records. This study was approved by the Institutional Review Board at the Samsung Medical Center (Seoul, Korea).
- DNA extraction
- Genomic DNA was extracted from 5-µm-thick sections of 10% neutral formalin-fixed, paraffin-embedded tumor tissue blocks using the QIAamp DNA mini kit (Qiagen, Hilden, Germany). The concentration and purity of the extracted DNA was determined by a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). The extracted DNA was stocked at -20℃ until use.
- Genomic DNA from the A549 cell lines was diluted with the DNA from the HeLa cells (New England Biolabs, Hitchin, UK) to give mutant/wild-type ratios of 0%, 0.01%, 0.05%, 0.1%, 0.5%, 1%, 10%, and 100%. The manufactured DNA targets were stored at -20℃ until use.
- Direct Sanger sequencing of KRAS
- Mutational analysis of KRAS exon 2 was performed by direct Sanger sequencing of PCR products amplified from genomic DNA. PCR was performed in a 20-µL volume containing 100 ng of template DNA, 10× PCR buffer, 0.25 mM dNTPs, 10 pmol primers, and 1.25 U Taq DNA polymerase (iNtRON, Daejeon, Korea). PCR products were electrophoresed on 2% agarose gels and purified with a QIAquick PCR purification kit (Qiagen). Bidirectional sequencing was performed using the BigDye Terminator v1.1 kit (Applied Biosystems, Foster City, CA, USA) on an ABI 3130xl genetic analyzer (Applied Biosystems). Chromatograms were manually reviewed for sequence analysis. Confirmatory re-sequencing from replicate PCR amplification reactions was performed for any sequences that were ambiguous. The results were marked as mutation positive if a mutation was detected in both the forward and reverse DNA strand.
- PNA-mediated clamping PCR of KRAS
- Assays for the detection of seven different KRAS variants were obtained through the PNAClamp KRAS, Mutation Detection kit (Panagene Inc., Daejeon, Korea). All reactions were performed in a 20-µL volume using 10 ng template DNA, a primer and PNA probe set, and SYBR Green PCR master mix. All reagents needed were included with the kit. The real-time PCR reaction of PNA-mediated clamping was performed using a CFX 96 (Bio-Rad, Hercules, CA, USA). PCR cycling conditions were a 5-minute hold at 94℃ followed by 40 cycles of 94℃ for 30 seconds, 70℃ for 20 seconds, 63℃ for 30 seconds, and 72℃ for 30 seconds. Detection of each of seven mutations in the KRAS gene was possible using one-step PNA-mediated real-time PCR clamping. In this assay, PNA probes and DNA primers were used together in the clamping reaction. Positive signals were detected by intercalation of SYBR Green fluorescent dye. The PNA probe sequence, which was complementary to wild-type DNA, suppressed amplification of the wild-type target, thereby enhancing preferential amplification of mutant sequences by competitively inhibiting DNA primer binding to wild-type DNA. PCR efficiency was determined by measuring the threshold cycle (Ct) value. Ct values for the control and mutation assays were obtained by observing the SYBR Green amplification plots. Mutation status was determined based on a Ct value difference greater than 2 between the control and sample.
- Next-generation sequencing
- Target samples were analyzed by next-generation sequencing for KRAS mutations with the Cancer Panel on a GS Junior Sequencer (Roche Diagnostics, Mannheim, Germany). For conventional 454 targeted resequencing, 30 ng of genomic DNA was used for PCR of the KRAS panel (SeaSun Biomaterials, Daejeon, Korea). Subsequent processing of the samples was performed according to the manufacturer's protocol.
- To increase the resolution of low level somatic mutant molecules within a high background of wild-type molecules, Insight Onco Panel for KRAS (SeaSun Biomaterials) was used for mutant enrichment PCR according to the manufacturer's instructions. The mutant-specific enrichment PCR was performed using 30 ng of genomic DNA and subsequent processing of the samples was performed according to the manufacturer's protocol. Nonspecific PCR products were removed by Agencourt AMPure XP beads (Beckman Coulter, Vienna, Austria) using a 1:1 DNA to bead ratio.
- Sequencing library preparation PCR was performed using 2 µL of purified PCR products from enrichment PCR amplification as a template, KRAS codon 12/13 Insight 2× Seq Lib Pep Premix (SeaSun Biomaterials) and each barcoded primer pair. The sequencing adaptor with multiplex identifier (MID) was conjugated using the manufacturer's protocol. Unwanted short fragments were removed by Agencourt AMPure XP beads (Beckman Coulter) using a 1:1 DNA to bead ratio.
- Purified amplicons were quantitated by Pico-Green (Life Technologies, Carlsbad, CA, USA) utilizing an external Infinite F200Pro fluorometer (Tecan, Grodig, Austria) with Magellan v7.0 Software (Tecan). Based on the standard concentrations, the signals were directly translated to ng/µL and the coefficient of determination (validation criteria r2>0.99) was calculated from eight DNA standards in a range from 0 to 100 ng/µL. For emulsion PCR amplification, the concentrations from the amplicons were converted in molecules/µL using the associated amplicon length. The manufactured DNA pools were stored at -20℃ until further use.
- Pyrosequencing was carried out according to the manufacturer's protocol for amplicons using the GS Junior System (Roche Diagnostics). Emulsion PCR, breaking, and bead enrichment was carried out using the GS Junior Titanium emPCR Kit Lib-L, emPCR Reagents Lib-L kit, Oil and Breaking kit, and the Bead Recovery Reagents kit according to the supplier's instructions (Roche Diagnostics). For emulsion PCR, we used a copy-per-bead ratio of 0.5. Enrichment of DNA-carrying beads was done with magnetic beads and a magnetic particle collector (Invitrogen, Carlsbad, CA, USA). To evaluate the amount of enriched beads, counting was performed using the GS Junior Bead Counter (Roche Diagnostics). Finally, we loaded 100,000-500,000 beads onto the PicoTiterPlate (Roche Diagnostics). Sequencing was carried out according to standard Roche/454 protocols using the GS Titanium Sequencing kit (Roche Diagnostics) and the GS Junior device.
- Processed and quality-filtered reads were analyzed with the GS Amplicon Variant Analyzer. Sequencing reads data were visualized using the GS Amplicon Variant Analyzer (Roche Diagnostics). The KRAS amplicons (excluding adaptors and MIDs) were used as the references to align amplicon reads, template-specific portions of the fusion primers were considered primer A and primer B, and the known mutations of the samples selected were defined as substitutions relative to the reference sequence. Correspondence of samples and MID tags was specified and, as the MID was present in primer A, a "Primer 1 MID" encoding multiplexer was used to demultiplex the reads.
- Statistical analysis
- Statistical differences between the two methods were analyzed by the McNemar test. The chi-square test or Fisher's exact test was used to compare the qualitative data. The t-test was used to compare means. Statistical calculation was performed with SPSS ver. 18 (SPSS Inc., Chicago, IL, USA). A p<.05 was considered statistically significant.
MATERIALS AND METHODS
PCR amplification for conventional NGS
Enrichment PCR for mutant-enriched NGS and sequencing library preparation
Quantification and normalization of sequencing amplicons
Ultradeep pyrosequencing
Data analysis
- The mean age of patients was 63 years. Seventy-eight of the patients were male and fifty-six were female. The majority of cases were adenocarcinomas (n=124), though there were also four cases of mucinous adenocarcinoma, three cases of squamous cell carcinoma, and one case of pleomorphic carcinoma. Thirty-four cases had stage I tumors, fifteen cases had stage II tumors, nineteen cases had stage III tumors, and sixty-six cases had stage IV tumors.
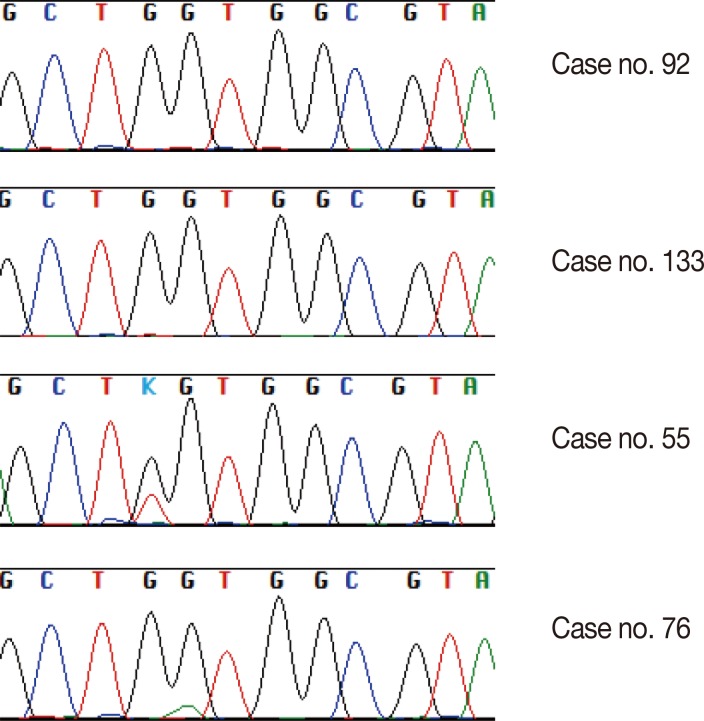
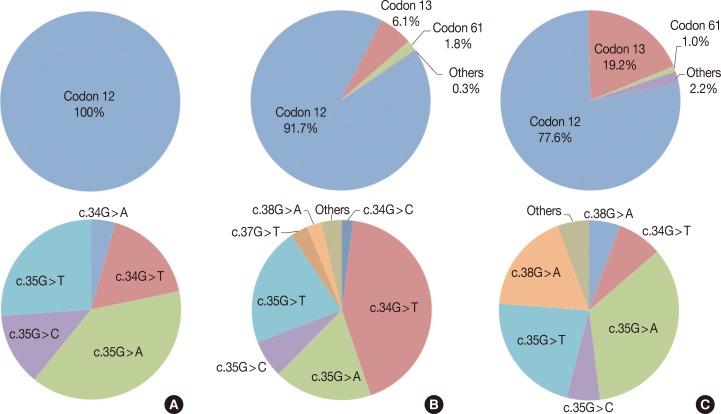
- Twenty-one (15.7%) of the 134 cases were found to have a KRAS missense mutation using direct Sanger sequencing. However, when the samples were tested using the PNA-mediated PCR clamping method, two additional cases (case nos. 92 and 133) were found to have a KRAS mutation. Chromatograms of discordant cases were reviewed, and clearly showed no abnormal peaks (Fig. 1). However, the difference between the two methods was not statistically significant (p=.5). All missense mutations were located on codon 12, and 18 mutations were located on the 35th base (Fig. 2A). This frequency was similar to that reported in the Catalogue of Somatic Mutations in Cancers (COSMIC) (http://www.sanger.ac.uk/cosmic) (Fig. 2B).
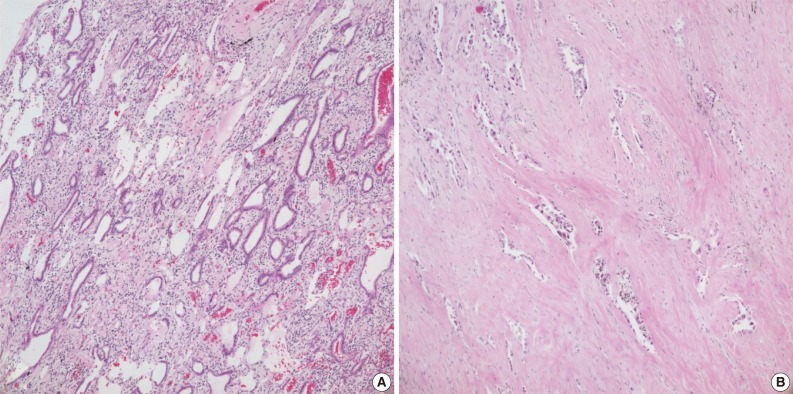
- Case no. 92 was diagnosed as adenocarcinoma, with a tumor size of 1 cm and a tumor volume of approximately 20% (Fig. 3A). The sample of case no. 133 was taken from an adenocarcinoma that had been resected following neoadjuvant chemotherapy. This tumor showed a diffuse desmoplastic reaction with scattered tumor cells, with tumor comprising only about 5% of the resected tissue (Fig. 3B).
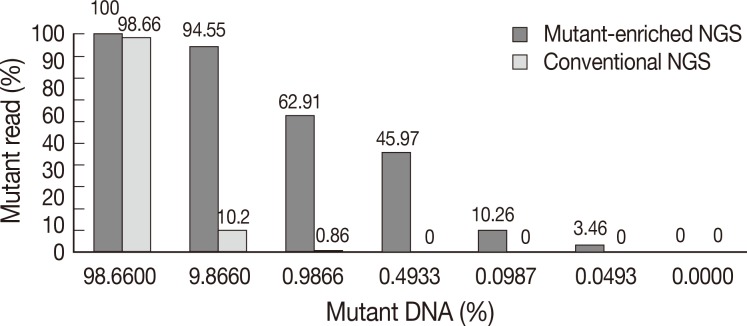
- We validated the discrepant cases by NGS with and without mutant enrichment using PNA-clamping. We compared the sensitivity of conventional NGS and mutant-enriched NGS by diluting control DNA. KRAS mutations could be detected by conventional NGS when the mutant allele comprised more than 1% of the sample. However, following enrichment, the mutation could be detected when the mutant allele comprised only 0.05% of the sample (Fig. 4).
- Using conventional NGS, the percentage of KRAS mutations detected in case no. 92 and case no. 133 was 2% and 4%, respectively. The mutant KRAS allele was increased in mutant-enriched NGS up to 90% and 89% in each case, respectively (Table 1). NGS was also performed on a wild-type case (case 2) for comparison.
- Clinicopathologic characteristics of 23 cases are summarized in Table 2. We analyzed the correlation between clinical data and KRAS mutation status (Table 3). KRAS mutations tended to be present in the tumors of smokers (p=.011). Of the 23 patients who had a KRAS mutated tumor, five (9.0%) were women and 18 (23%) were men. Cancer stage was also found to be related to KRAS mutation; stage IV tumors were more likely to have a KRAS mutation (p=.032).
RESULTS
- Of 134 cases of NSCLC, KRAS mutations were detected in 21 cases using direct sequencing and in 23 cases using a PNA-mediated PCR clamping method. Although the difference was not statistically significant, two additional cases of KRAS mutation were detected with the PNA-mediated PCR clamping method. The mutation status of the two discordant samples was confirmed using NGS. The percentage of mutant alleles in both samples was less than 5%. With such a low percentage, a mutation peak would be indistinguishable from background noise by direct sequencing.11,12 One of these cases had a low viable tumor volume due to neoadjuvant chemotherapy. The tumor volume of the other case was not as low. Several factors may have contributed to the lower mutant proportion. DNA of inflammatory cells around the tumor can dilute tumor DNA. Tumor volume may have been diminished on serial sections of paraffin block. In addition, tumor heterogeneity may be present. In conventional NGS, the percentage of mutations detected was approximately the same as the percentage of mutant alleles in the sample. This allows one to reliably determine tumor heterogeneity. A proportion of mutant alleles in a sample of less than 1% means that the mutation cannot be detected using conventional NGS. In contrast, as few as 0.05% mutant alleles can be detected after enrichment for the mutant alleles using the PNA clamping method.
- Similar to previous studies,1,16 our results show an association between KRAS mutation and smoking status. KRAS mutations were more frequent in male patients than female patients. It is possible that there is a sex bias in this relationship, as 71 of 73 smokers in the sample were men. Tumor stage was also related to KRAS mutation according to our data. However, in other reports, stage was not significantly correlated with KRAS mutation.17,18,19,20,21,22 In one meta-analysis,23 KRAS mutation was found to be a statistically significant prognostic factor in lung adenocarcinoma. There have been multiple reports recently addressing the relationship between KRAS mutation status and survival, with various outcomes (Table 4). Since the mutant-enriched sequencing method is more sensitive than direct sequencing, the KRAS mutation rate is relatively higher in reports using mutant-enriched sequencing. Racial difference and proportion of adenocarcinoma included in study also affected KRAS mutation rate. Studies of a Caucasian population or adenocarcinoma have higher KRAS mutation rates than studies of Asian populations or any histologic type of NSCLC.
- In this study, the number of cases was too small to establish statistical significance between the two methods. Survival analysis could not be performed due to the short follow-up period. Further large-scale studies may be required to assess the differences between the two methods of KRAS mutation detection in NSCLC.
- As targeted therapy for KRAS mutation continues to develop, testing in NSCLCs will become more important. Direct sequencing can accurately detect mutations when the percentage of tumor cells in the analytical sample is sufficiently high. However, it is not always possible to obtain samples with enough volume to undertake these tests. In these situations, methods other than direct sequencing are required. The PNA-mediated PCR clamping method can quickly provide results and is sufficiently sensitive in this situation.
DISCUSSION
Acknowledgments
Acknowledgments
- 1. Riely GJ, Marks J, Pao W. KRAS mutations in non-small cell lung cancer. Proc Am Thorac Soc 2009; 6: 201-205. ArticlePubMed
- 2. Forbes S, Clements J, Dawson E, et al. Cosmic 2005. Br J Cancer 2006; 94: 318-322. ArticlePubMedPMCPDF
- 3. Herbst RS, Prager D, Hermann R, et al. TRIBUTE: a phase III trial of erlotinib hydrochloride (OSI-774) combined with carboplatin and paclitaxel chemotherapy in advanced non-small-cell lung cancer. J Clin Oncol 2005; 23: 5892-5899. ArticlePubMed
- 4. Pao W, Wang TY, Riely GJ, et al. KRAS mutations and primary resistance of lung adenocarcinomas to gefitinib or erlotinib. PLoS Med 2005; 2: e17.ArticlePubMedPMC
- 5. Kim ES, Herbst RS, Wistuba II, et al. The BATTLE trial: personalizing therapy for lung cancer. Cancer Discov 2011; 1: 44-53. ArticlePubMedPMCPDF
- 6. Mok TS, Wu YL, Thongprasert S, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med 2009; 361: 947-957. ArticlePubMed
- 7. Zhou C, Wu YL, Chen G, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol 2011; 12: 735-742. ArticlePubMed
- 8. Lindeman NI, Cagle PT, Beasley MB, et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. Arch Pathol Lab Med 2013; 137: 828-860. ArticlePubMedPMC
- 9. Thiede C, Bayerdörffer E, Blasczyk R, Wittig B, Neubauer A. Simple and sensitive detection of mutations in the ras proto-oncogenes using PNA-mediated PCR clamping. Nucleic Acids Res 1996; 24: 983-984. ArticlePubMedPMC
- 10. Nielsen PE, Egholm M, Berg RH, Buchardt O. Sequence-selective recognition of DNA by strand displacement with a thymine-substituted polyamide. Science 1991; 254: 1497-1500. ArticlePubMed
- 11. Beau-Faller M, Legrain M, Voegeli AC, et al. Detection of K-Ras mutations in tumour samples of patients with non-small cell lung cancer using PNA-mediated PCR clamping. Br J Cancer 2009; 100: 985-992. ArticlePubMedPMCPDF
- 12. Kwon MJ, Lee SE, Kang SY, Choi YL. Frequency of KRAS, BRAF, and PIK3CA mutations in advanced colorectal cancers: Comparison of peptide nucleic acid-mediated PCR clamping and direct sequencing in formalin-fixed, paraffin-embedded tissue. Pathol Res Pract 2011; 207: 762-768. ArticlePubMed
- 13. Araki T, Shimizu K, Nakamura K, et al. Usefulness of peptide nucleic acid (PNA)-clamp smart amplification process version 2 (SmartAmp2) for clinical diagnosis of KRAS codon 12 mutations in lung adenocarcinoma: comparison of PNA-clamp SmartAmp2 and PCR-related methods. J Mol Diagn 2010; 12: 118-124. PubMedPMC
- 14. Jeong D, Jeong Y, Lee S, et al. Detection of BRAF(V600E) mutations in papillary thyroid carcinomas by peptide nucleic acid clamp real-time PCR: a comparison with direct sequencing. Korean J Pathol 2012; 46: 61-67. ArticlePubMedPMC
- 15. Lee HJ, Xu X, Kim H, et al. Comparison of direct sequencing, PNA clamping-real time polymerase chain reaction, and pyrosequencing methods for the detection of EGFR mutations in non-small cell lung carcinoma and the correlation with clinical responses to EGFR tyrosine kinase inhibitor treatment. Korean J Pathol 2013; 47: 52-60. ArticlePubMedPMC
- 16. Le Calvez F, Mukeria A, Hunt JD, et al. TP53 and KRAS mutation load and types in lung cancers in relation to tobacco smoke: distinct patterns in never, former, and current smokers. Cancer Res 2005; 65: 5076-5083. ArticlePubMedPDF
- 17. Cadranel J, Mauguen A, Faller M, et al. Impact of systematic EGFR and KRAS mutation evaluation on progression-free survival and overall survival in patients with advanced non-small-cell lung cancer treated by erlotinib in a French prospective cohort (ERMETIC project. part 2). J Thorac Oncol 2012; 7: 1490-1502. ArticlePubMed
- 18. Guan JL, Zhong WZ, An SJ, et al. KRAS mutation in patients with lung cancer: a predictor for poor prognosis but not for EGFR-TKIs or chemotherapy. Ann Surg Oncol 2013; 20: 1381-1388. ArticlePubMedPDF
- 19. Kim HR, Shim HS, Chung JH, et al. Distinct clinical features and outcomes in never-smokers with nonsmall cell lung cancer who harbor EGFR or KRAS mutations or ALK rearrangement. Cancer 2012; 118: 729-739. ArticlePubMed
- 20. Kosaka T, Yatabe Y, Onozato R, Kuwano H, Mitsudomi T. Prognostic implication of EGFR, KRAS, and TP53 gene mutations in a large cohort of Japanese patients with surgically treated lung adenocarcinoma. J Thorac Oncol 2009; 4: 22-29. ArticlePubMed
- 21. Ragusa M, Vannucci J, Ludovini V, et al. Impact of epidermal growth factor receptor and KRAS mutations on clinical outcome in resected non-small cell lung cancer patients. Am J Clin Oncol 2013 1 24 [Epub]. http://dx.doi.org/10.1097/COC.0b013e31827a7e7a. Article
- 22. Sonobe M, Kobayashi M, Ishikawa M, et al. Impact of KRAS and EGFR gene mutations on recurrence and survival in patients with surgically resected lung adenocarcinomas. Ann Surg Oncol 2012; 19(Suppl 3):S347-S354. ArticlePubMedPDF
- 23. Mascaux C, Iannino N, Martin B, et al. The role of RAS oncogene in survival of patients with lung cancer: a systematic review of the literature with meta-analysis. Br J Cancer 2005; 92: 131-139. ArticlePubMedPDF
- 24. Marchetti A, Milella M, Felicioni L, et al. Clinical implications of KRAS mutations in lung cancer patients treated with tyrosine kinase inhibitors: an important role for mutations in minor clones. Neoplasia 2009; 11: 1084-1092. ArticlePubMedPMC
- 25. Sun JM, Hwang DW, Ahn JS, Ahn MJ, Park K. Prognostic and predictive value of KRAS mutations in advanced non-small cell lung cancer. PLoS One 2013; 8: e64816.ArticlePubMedPMC
- 26. Johnson ML, Sima CS, Chaft J, et al. Association of KRAS and EGFR mutations with survival in patients with advanced lung adenocarcinomas. Cancer 2013; 119: 356-362. ArticlePubMedPDF
- 27. Kerner GS, Schuuring E, Sietsma J, et al. Common and rare EGFR and KRAS mutations in a Dutch non-small-cell lung cancer population and their clinical outcome. PLoS One 2013; 8: e70346.ArticlePubMedPMC
- 28. Lim EH, Zhang SL, Li JL, et al. Using whole genome amplification (WGA) of low-volume biopsies to assess the prognostic role of EGFR, KRAS, p53, and CMET mutations in advanced-stage non-small cell lung cancer (NSCLC). J Thorac Oncol 2009; 4: 12-21. ArticlePubMed
- 29. Liu HP, Isaac Wu HD, Chang JW, et al. Prognostic implications of epidermal growth factor receptor and KRAS gene mutations and epidermal growth factor receptor gene copy numbers in patients with surgically resectable non-small cell lung cancer in Taiwan. J Thorac Oncol 2010; 5: 1175-1184. ArticlePubMed
- 30. Scoccianti C, Vesin A, Martel G, et al. Prognostic value of TP53, KRAS and EGFR mutations in nonsmall cell lung cancer: the EUELC cohort. Eur Respir J 2012; 40: 177-184. ArticlePubMed
REFERENCES
| Case No. | Result | Sex | Age (yr) | Pattern | Size (cm) | T stage | N stage | M stage | Stage | Smoking | Chemotherapy |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | c.35G>C (p.G12A) | F | 61 | A&S | 1.8 | 2a | 2 | 1b | IV | Never | Yes |
| 5 | c.35G>T (p.G12V) | M | 77 | NA | 2.5 | 1b | 2 | 1b | IV | Former | Yes |
| 6 | c.35G>A (p.G12D) | F | 67 | S | 3.5 | 2a | 1 | 0 | IIA | Never | No |
| 11 | c.35G>C (p.G12A) | M | 59 | NA | 4.3 | 2a | 1 | 1b | IV | Former | No |
| 16 | c.35G>A (p.G12D) | M | 61 | S | 6 | 2b | 0 | 0 | IIA | Current | No |
| 26 | c.35G>A (p.G12D) | F | 52 | NA | 8 | 3 | 0 | 1a | IV | Never | Yes |
| 29 | c.35G>A (p.G12D) | M | 74 | A | 3.7 | 4 | 0 | 1a | IV | Current | Yes |
| 38 | c.35G>A (p.G12D) | M | 52 | A | 2.2 | 1b | 2 | 0 | IIIA | Current | No |
| 49 | c.35G>T (p.G12V) | M | 67 | A | 3.4 | 2a | 0 | 1b | IV | Former | No |
| 51 | c.34G>T (p.G12C) | M | 56 | A&S | 3 | 1b | 1 | 1b | IV | Current | No |
| 52 | c.35G>A (p.G12D) | M | 58 | P | 5.5 | 3 | 3 | 1b | IV | Current | No |
| 54 | c.34G>T (p.G12C) | M | 66 | A | 5.9 | 4 | 3 | 1b | IV | Current | No |
| 55 | c.34G>T (p.G12C) | M | 55 | S | 7.8 | 3 | 0 | 0 | IIB | Former | No |
| 61 | c.35G>T (p.G12V) | M | 70 | NA | 4.1 | 2a | 2 | 1a | IV | Current | No |
| 72 | c.35G>A (p.G12D) | M | 73 | A&P | 2.4 | 1b | 1 | 1a | IV | Former | Yes |
| 76 | c.35G>A (p.G12D) | M | 66 | A | 2.5 | 1b | 1 | 0 | IIA | Current | No |
| 80 | c.35G>A (p.G12D) | M | 73 | A&S | 4 | 4 | 3 | 1b | IV | Former | No |
| 81 | c.35G>T (p.G12V) | M | 75 | S | 3.5 | 4 | 3 | 1a | IV | Former | No |
| 92a | c.35G>T (p.G12V) | F | 69 | A | 1 | 1a | 0 | 0 | IA | Never | No |
| 96 | c.35G>C (p.G12A) | M | 55 | A | 4.3 | 2a | 0 | 1a | IV | Current | No |
| 113 | c.35G>T (p.G12V) | M | 70 | P | 1.9 | 1a | 3 | 1b | IV | Former | No |
| 133a | c.34G>T (p.G12C) | M | 55 | A | 1.8 | 2a | 2 | 0 | IIIA | Current | Yes |
| 134 | c.34G>A (p.G12S) | F | 58 | A | 6.9 | 2a | 3 | 1b | IV | Never | No |
| Country | KRAS(+) (%) | No, of cases | Method | Patients | Survival relation | |
|---|---|---|---|---|---|---|
| Guan et al. (2013) [18] | China | 5 | 1,928 | DS or high resolution melting analysis | Operable and inoperable | Yes |
| Marchetti et al. (2009) [24] | Italy | 36 | 83 | Mutant-enriched sequencing | Adenocarcinoma | Yes |
| Treated with EGFR-TKI | ||||||
| Sun et al. (2013) [25] | Korea | 8 | 484 | DS | Advanced stage | Yes |
| Johnson et al. (2013) [26] | USA | 23 | 1,036 | DS or mass-spectrometry-based genotyping | Advanced stage | Yes |
| Adenocarcinoma | ||||||
| Sonobe et al. (2012) [22] | Japan | 18 | 180 | RFLP | Resection | No |
| Adenocarcinoma | ||||||
| Cadranel et al. (2012) [17] | France | 14 | 522 | DS or nested sequencing | Advanced stage | Yes |
| Treated with EGFR-TKI | ||||||
| Kerner et al. (2013) [27] | The Netherlands | 30 | 442 | DS | Operable and inoperable | No |
| Kim et al. (2012) [19] | Korea | 4 | 229 | DS | Never-smoker | Yes |
| Ragusa et al. (2013) [21] | Italy | 17 | 230 | DS | Resection | No |
| Kosaka et al. (2009) [20] | Japan | 13 | 397 | DS | Resection | No |
| Adenocarcinoma | ||||||
| Lim et al. (2009) [28] | Singapore | 6 | 88 | Whole genome amplification and DS | Advanced stage | Yes |
| Liu et al. (2010) [29] | Taiwan | 5 | 73 | DS | Resection | No |
| Scoccianti et al. (2012) [30] | Europe | 19 | 250 | Mutant-enriched sequencing | Resection | No |
Figure & Data
References
Citations

- KRAS mutated Non‐Small Lung Carcinoma: A Real World Context from the Indian subcontinent
Ullas Batra, Shrinidhi Nathany, Mansi Sharma, Amrith BP, Joslia T. Jose, Harkirat Singh, Sakshi Mattoo, Anurag Mehta
Cancer Medicine.2023; 12(3): 2869. CrossRef - Mechanism exploration and prognosis study of Astragali Radix-Spreading hedyotis herb for the treatment of lung adenocarcinoma based on bioinformatics approaches and molecular dynamics simulation
Junfeng Guo, Yuting Zhao, Xuanyu Wu, Ganggang Li, Yuwei Zhang, Yang Song, Quanyu Du
Frontiers in Chemistry.2023;[Epub] CrossRef - PEAC: An Ultrasensitive and Cost-Effective MRD Detection System in Non-small Cell Lung Cancer Using Plasma Specimen
Jianping Xu, Yue Pu, Rui Lin, Shanshan Xiao, Yingxue Fu, Tao Wang
Frontiers in Medicine.2022;[Epub] CrossRef - Potential applications of peptide nucleic acid in biomedical domain
Kshitij RB Singh, Parikipandla Sridevi, Ravindra Pratap Singh
Engineering Reports.2020;[Epub] CrossRef - A Highly Sensitive Next-Generation Sequencing-Based Genotyping Platform for EGFR Mutations in Plasma from Non-Small Cell Lung Cancer Patients
Jung-Young Shin, Jeong-Oh Kim, Mi-Ran Lee, Seo Ree Kim, Kyongmin Sarah Beck, Jin Hyoung Kang
Cancers.2020; 12(12): 3579. CrossRef - Association with PD-L1 Expression and Clinicopathological Features in 1000 Lung Cancers: A Large Single-Institution Study of Surgically Resected Lung Cancers with a High Prevalence of EGFR Mutation
Seung Eun Lee, Yu Jin Kim, Minjung Sung, Mi-Sook Lee, Joungho Han, Hong Kwan Kim, Yoon-La Choi
International Journal of Molecular Sciences.2019; 20(19): 4794. CrossRef - Peptide Nucleic Acid Clamping and Direct Sequencing in the Detection of Oncogenic Alterations in Lung Cancer: Systematic Review and Meta-Analysis
Jae-Uk Song, Jonghoo Lee
Yonsei Medical Journal.2018; 59(2): 211. CrossRef - Comparative analysis of Adam33 mutations in murine lung cancer cell lines by droplet digital PCR, real-time PCR and Insight Onco™ NGS
Soo-Jin Kim, Eunhee Kim, Kyung-Taek Rim
Molecular & Cellular Toxicology.2018; 14(2): 221. CrossRef - KRAS mutation analysis by next‐generation sequencing in endoscopic ultrasound‐guided sampling for solid liver masses
Hyun Jong Choi, Jong Ho Moon, Hee Kyung Kim, Yun Nah Lee, Tae Hoon Lee, Sang‐Woo Cha, Young Deok Cho, Sang‐Heum Park
Journal of Gastroenterology and Hepatology.2017; 32(1): 154. CrossRef - Molecular Testing of Lung Cancers
Hyo Sup Shim, Yoon-La Choi, Lucia Kim, Sunhee Chang, Wan-Seop Kim, Mee Sook Roh, Tae-Jung Kim, Seung Yeon Ha, Jin-Haeng Chung, Se Jin Jang, Geon Kook Lee
Journal of Pathology and Translational Medicine.2017; 51(3): 242. CrossRef - Peptide nucleic acids: Advanced tools for biomedical applications
Anjali Gupta, Anuradha Mishra, Nidhi Puri
Journal of Biotechnology.2017; 259: 148. CrossRef - Application of Peptide Nucleic Acid-based Assays Toward Detection of Somatic Mosaicism
Christopher S Hong, Chunzhang Yang, Zhengping Zhuang
Molecular Therapy - Nucleic Acids.2016; 5: e314. CrossRef - Detecting Primary KIT Mutations in Presurgical Plasma of Patients with Gastrointestinal Stromal Tumor
Guhyun Kang, Byeong Seok Sohn, Jung-Soo Pyo, Jung Yeon Kim, Boram Lee, Kyoung-Mee Kim
Molecular Diagnosis & Therapy.2016; 20(4): 347. CrossRef - Transformation to Small Cell Lung Cancer of Pulmonary Adenocarcinoma: Clinicopathologic Analysis of Six Cases
Soomin Ahn, Soo Hyun Hwang, Joungho Han, Yoon-La Choi, Se-Hoon Lee, Jin Seok Ahn, Keunchil Park, Myung-Ju Ahn, Woong-Yang Park
Journal of Pathology and Translational Medicine.2016; 50(4): 258. CrossRef - Clinicopathologic characteristics of EGFR, KRAS, and ALK alterations in 6,595 lung cancers
Boram Lee, Taebum Lee, Se-Hoon Lee, Yoon-La Choi, Joungho Han
Oncotarget.2016; 7(17): 23874. CrossRef - Detection of KIT and PDGFRA mutations in the plasma of patients with gastrointestinal stromal tumor
Guhyun Kang, Byung Noe Bae, Byeong Seok Sohn, Jung-Soo Pyo, Gu Hyum Kang, Kyoung-Mee Kim
Targeted Oncology.2015; 10(4): 597. CrossRef - Low Frequency of KRAS Mutation in Pancreatic Ductal Adenocarcinomas in Korean Patients and Its Prognostic Value
Mi Jung Kwon, Jang Yong Jeon, Hye-Rim Park, Eun Sook Nam, Seong Jin Cho, Hyung Sik Shin, Ji Hyun Kwon, Joo Seop Kim, Boram Han, Dong Hoon Kim, Yoon-La Choi
Pancreas.2015; 44(3): 484. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
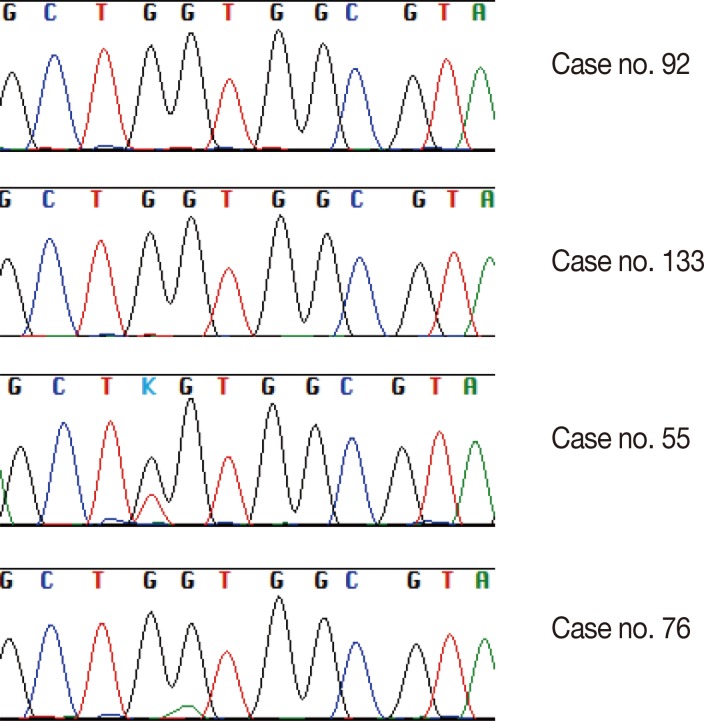
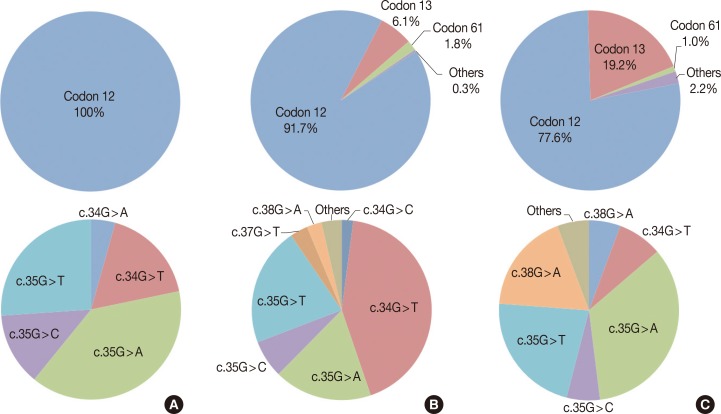
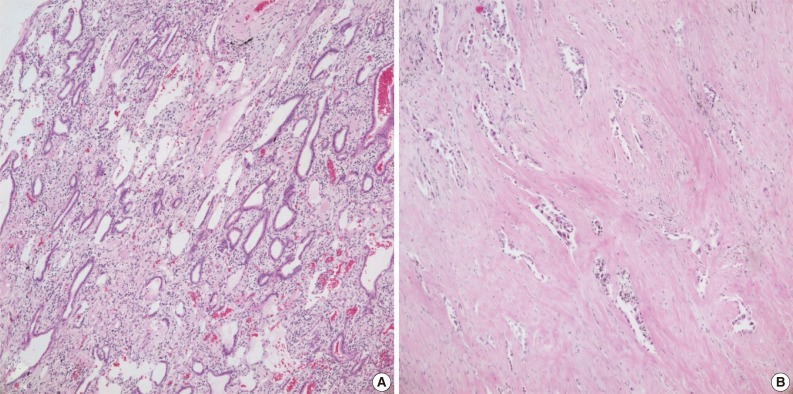
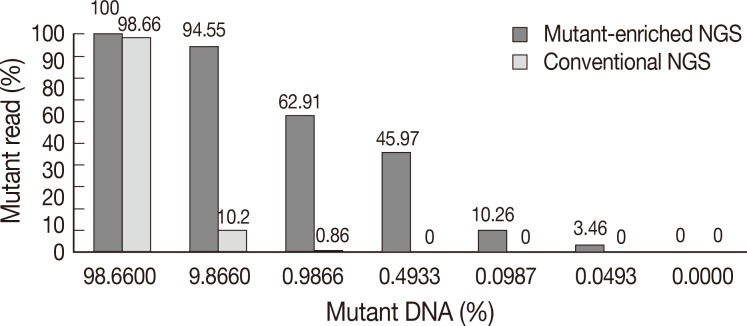
Fig. 1
Fig. 2
Fig. 3
Fig. 4
| Sample |
Depth | Mutant (%) |
Character | ||||
|---|---|---|---|---|---|---|---|
| n | Application | G12C | G12S | G12V | |||
| 1 | 2 | Insight | 9,215 | 0 | 0 | 0 | Wild |
| 2 | 2 | Normal | 17,592 | 0 | 0 | 0 | |
| 3 | 92 | Insight | 8,714 | 0 | 0 | 90.37 | G12V |
| 4 | 92 | Normal | 16,983 | 0 | 0 | 2.07 | |
| 5 | 133 | Insight | 10,016 | 89.34 | 0 | 0 | G12C |
| 6 | 133 | Normal | 17,339 | 4 | 0 | 0 | |
| 7 | A549 | Insight | 11,220 | 0 | 95.38 | 0 | G12S |
| 8 | Hela | Insight | 12,227 | 0 | 0 | 0 | Wild |
| Case No. | Result | Sex | Age (yr) | Pattern | Size (cm) | T stage | N stage | M stage | Stage | Smoking | Chemotherapy |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | c.35G>C (p.G12A) | F | 61 | A&S | 1.8 | 2a | 2 | 1b | IV | Never | Yes |
| 5 | c.35G>T (p.G12V) | M | 77 | NA | 2.5 | 1b | 2 | 1b | IV | Former | Yes |
| 6 | c.35G>A (p.G12D) | F | 67 | S | 3.5 | 2a | 1 | 0 | IIA | Never | No |
| 11 | c.35G>C (p.G12A) | M | 59 | NA | 4.3 | 2a | 1 | 1b | IV | Former | No |
| 16 | c.35G>A (p.G12D) | M | 61 | S | 6 | 2b | 0 | 0 | IIA | Current | No |
| 26 | c.35G>A (p.G12D) | F | 52 | NA | 8 | 3 | 0 | 1a | IV | Never | Yes |
| 29 | c.35G>A (p.G12D) | M | 74 | A | 3.7 | 4 | 0 | 1a | IV | Current | Yes |
| 38 | c.35G>A (p.G12D) | M | 52 | A | 2.2 | 1b | 2 | 0 | IIIA | Current | No |
| 49 | c.35G>T (p.G12V) | M | 67 | A | 3.4 | 2a | 0 | 1b | IV | Former | No |
| 51 | c.34G>T (p.G12C) | M | 56 | A&S | 3 | 1b | 1 | 1b | IV | Current | No |
| 52 | c.35G>A (p.G12D) | M | 58 | P | 5.5 | 3 | 3 | 1b | IV | Current | No |
| 54 | c.34G>T (p.G12C) | M | 66 | A | 5.9 | 4 | 3 | 1b | IV | Current | No |
| 55 | c.34G>T (p.G12C) | M | 55 | S | 7.8 | 3 | 0 | 0 | IIB | Former | No |
| 61 | c.35G>T (p.G12V) | M | 70 | NA | 4.1 | 2a | 2 | 1a | IV | Current | No |
| 72 | c.35G>A (p.G12D) | M | 73 | A&P | 2.4 | 1b | 1 | 1a | IV | Former | Yes |
| 76 | c.35G>A (p.G12D) | M | 66 | A | 2.5 | 1b | 1 | 0 | IIA | Current | No |
| 80 | c.35G>A (p.G12D) | M | 73 | A&S | 4 | 4 | 3 | 1b | IV | Former | No |
| 81 | c.35G>T (p.G12V) | M | 75 | S | 3.5 | 4 | 3 | 1a | IV | Former | No |
| 92 |
c.35G>T (p.G12V) | F | 69 | A | 1 | 1a | 0 | 0 | IA | Never | No |
| 96 | c.35G>C (p.G12A) | M | 55 | A | 4.3 | 2a | 0 | 1a | IV | Current | No |
| 113 | c.35G>T (p.G12V) | M | 70 | P | 1.9 | 1a | 3 | 1b | IV | Former | No |
| 133 |
c.34G>T (p.G12C) | M | 55 | A | 1.8 | 2a | 2 | 0 | IIIA | Current | Yes |
| 134 | c.34G>A (p.G12S) | F | 58 | A | 6.9 | 2a | 3 | 1b | IV | Never | No |
| Mutation | Wild type | p-value | ||
|---|---|---|---|---|
| Gender | Male | 18 (78) | 60 (54) | .032 |
| Female | 5 (22) | 51 (46) | ||
| Age (yr) | Mean | 63.9 | 62.6 | .517 |
| ≤ 60 | 9 (39) | 44 (40) | .964 | |
| >60 | 14 (61) | 67 (60) | ||
| Smoking | Never | 5 (22) | 56 (50) | .011 |
| Former | 8 (35) | 35 (32) | ||
| Current | 10 (43) | 20 (18) | ||
| Type | Adenocarcinoma | 22 (96) | 101 (91) | .644 |
| Mucinous adenocarcinoma | 1 (4) | 3 (3) | ||
| Squamous cell carcinoma | 0 (0) | 3 (3) | ||
| Others | 0 (0) | 4 (4) | ||
| Size (cm) | ≤ 3 | 9 (39) | 47 (42) | .766 |
| >3 | 14 (61) | 64 (58) | ||
| T stage | 0-1 | 7 (30) | 42 (38) | .921 |
| 2 | 9 (39) | 42 (38) | ||
| 3 | 3 (13) | 14 (13) | ||
| 4 | 4 (17) | 13 (12) | ||
| N stage | 0 | 7 (30) | 51 (46) | .446 |
| 1 | 5 (22) | 13 (12) | ||
| 2 | 5 (22) | 23 (21) | ||
| 3 | 6 (26) | 24 (22) | ||
| M stage | 0 | 7 (30) | 61 (55) | .032 |
| 1 | 16 (70) | 50 (45) |
| Country | KRAS(+) (%) | No, of cases | Method | Patients | Survival relation | |
|---|---|---|---|---|---|---|
| Guan et al. (2013) [18] | China | 5 | 1,928 | DS or high resolution melting analysis | Operable and inoperable | Yes |
| Marchetti et al. (2009) [24] | Italy | 36 | 83 | Mutant-enriched sequencing | Adenocarcinoma | Yes |
| Treated with EGFR-TKI | ||||||
| Sun et al. (2013) [25] | Korea | 8 | 484 | DS | Advanced stage | Yes |
| Johnson et al. (2013) [26] | USA | 23 | 1,036 | DS or mass-spectrometry-based genotyping | Advanced stage | Yes |
| Adenocarcinoma | ||||||
| Sonobe et al. (2012) [22] | Japan | 18 | 180 | RFLP | Resection | No |
| Adenocarcinoma | ||||||
| Cadranel et al. (2012) [17] | France | 14 | 522 | DS or nested sequencing | Advanced stage | Yes |
| Treated with EGFR-TKI | ||||||
| Kerner et al. (2013) [27] | The Netherlands | 30 | 442 | DS | Operable and inoperable | No |
| Kim et al. (2012) [19] | Korea | 4 | 229 | DS | Never-smoker | Yes |
| Ragusa et al. (2013) [21] | Italy | 17 | 230 | DS | Resection | No |
| Kosaka et al. (2009) [20] | Japan | 13 | 397 | DS | Resection | No |
| Adenocarcinoma | ||||||
| Lim et al. (2009) [28] | Singapore | 6 | 88 | Whole genome amplification and DS | Advanced stage | Yes |
| Liu et al. (2010) [29] | Taiwan | 5 | 73 | DS | Resection | No |
| Scoccianti et al. (2012) [30] | Europe | 19 | 250 | Mutant-enriched sequencing | Resection | No |
F, female; A, acinar; S, solid; M, male; NA, not applicable; P, papillary. In this case, mutation is not detected by direct sequencing.
Values are presented as number (%).
DS, direct sequencing; EGFR-TKI, epidermal growth factor receptor tyrosine kinase inhibitor; RFLP, restriction fragment length polymorphism.

 E-submission
E-submission






